Analysis of gene-derived SNP marker polymorphism in US wheat (Triticum aestivum L.) cultivars (original) (raw)
Abstract
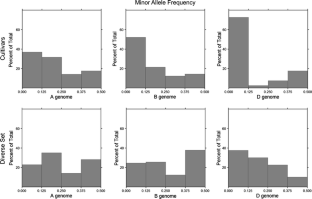
In this study, we developed 359 detection primers for single nucleotide polymorphisms (SNPs) previously discovered within intron sequences of wheat genes and used them to evaluate SNP polymorphism in common wheat (Triticum aestivum L.). These SNPs showed an average polymorphism information content (PIC) of 0.18 among 20 US elite wheat cultivars, representing seven market classes. This value increased to 0.23 when SNPs were pre-selected for polymorphisms among a diverse set of 13 hexaploid wheat accessions (excluding synthetic wheats) used in the wheat SNP discovery project (http://wheat.pw.usda.gov/SNP). PIC values for SNP markers in the D genome were approximately half of those for the A and B genomes. D genome SNPs also showed a larger PIC reduction relative to the other genomes (P < 0.05) when US cultivars were compared with the more diverse set of 13 wheat accessions. Within those accessions, D genome SNPs show a higher proportion of alleles with low minor allele frequencies (<0.125) than found in the other two genomes. These data suggest that the reduction of PIC values in the D genome was caused by differential loss of low frequency alleles during the population size bottleneck that accompanied the development of modern commercial cultivars. Additional SNP discovery efforts targeted to the D genome in elite wheat germplasm will likely be required to offset the lower diversity of this genome. With increasing SNP discovery projects and the development of high-throughput SNP assay technologies, it is anticipated that SNP markers will play an increasingly important role in wheat genetics and breeding applications.
Access this article
Subscribe and save
- Starting from 10 chapters or articles per month
- Access and download chapters and articles from more than 300k books and 2,500 journals
- Cancel anytime View plans
Buy Now
Price excludes VAT (USA)
Tax calculation will be finalised during checkout.
Instant access to the full article PDF.
Fig. 1
Similar content being viewed by others
Abbreviations
EST:
Expressed sequence tag
FP:
Fluorescence polarization
HRS:
Hard red spring
HWS:
Hard white spring
HRW:
Hard red winter
HWW:
Hard White Winter
PIC:
Polymorphism information content
SSR:
Simple sequence repeat
SNP:
Single nucleotide polymorphism
SRW:
Soft red winter
SWS:
Soft white spring
SWW:
Soft white winter
References
- Batley J, Barker G, O’Sullivan J, Edwards KJ, Edwards D (2003) Mining for single nucleotide polymorphisms and insertion/deletions in maize expressed sequence tag data. Plant Physiol 132:84–91. doi:10.1104/pp.102.019422
Article PubMed CAS Google Scholar - Blake NK, Sherman JD, Dvorak J, Talbert LE (2004) Genome-specific primer sets for starch biosynthesis genes in wheat. Theor Appl Genet 109:1295–1302. doi:10.1007/s00122-004-1743-4
Article PubMed CAS Google Scholar - Brookes A (1999) The essence of SNPs. Gene 234:177–186. doi:10.1016/S0378-1119(99)00219-X
Article PubMed CAS Google Scholar - Brumfield RT, Beerli P, Nickerson DA, Edwards SV (2003) The utility of single nucleotide polymorphisms in inferences of population history. Trends Ecol Evol 18:249–256. doi:10.1016/S0169-5347(03)00018-1
Article Google Scholar - Caldwell KS, Dvorak J, Lagudah ES, Akhunov E, Luo MC, Wolters P et al (2004) Sequence polymorphism in polyploid wheat and their D-genome diploid ancestor. Genetics 167:941–947. doi:10.1534/genetics.103.016303
Article PubMed CAS Google Scholar - Cardon LR, Abecasis GR (2003) Using haplotype blocks to map human complex trait loci. Trends Genet 19:135–140. doi:10.1016/S0168-9525(03)00022-2
Article PubMed CAS Google Scholar - Chao S, Zhang W, Dubcovsky J, Sorrells M (2007) Evaluation of genetic diversity and genome-wide linkage disequilibrium among US wheat (Triticum aestivum L.) germplasm representing different market classes. Crop Sci 47:1018–1030. doi:10.2135/cropsci2006.06.0434
Article CAS Google Scholar - Chen X, Levine L, Kwok P-Y (1999) Fluorescence polarization in homogeneous nucleic acid analysis. Genome Res 9:492–498
PubMed CAS Google Scholar - Ching A, Caldwell KS, Jung M, Dolan M, Smith OS, Tingey S et al (2002) SNP frequency, haplotype structure and linkage disequilibrium in elite maize inbred lines. BMC Genet 3:19. doi:10.1186/1471-2156-3-19
Article PubMed Google Scholar - Cho RJ, Mindrinos M, Richards DR, Sapolsky RJ, Anderson M, Drenkard E et al (1999) Genome-wide mapping with biallelic markers in Arabidopsis thaliana. Nat Genet 23:203–207. doi:10.1038/13833
Article PubMed CAS Google Scholar - Choi I-Y, Hyten DL, Matukumalli LK, Song Q, Chaky JM, Quigley CV et al (2007) A soybean transcript map: gene distribution, haplotype and single-nucleotide polymorphism analysis. Genetics 176:685–696. doi:10.1534/genetics.107.070821
Article PubMed CAS Google Scholar - Dubcovsky J, Dvorak J (2007) Genome plasticity a key factor in the success of polyploid wheat under domestication. Science 316:1862–1866. doi:10.1126/science.1143986
Article PubMed CAS Google Scholar - Dvorak J, Akhunov ED (2005) Tempos of gene locus deletions and duplications and their relationship to recombination rate during diploid and polyploid evolution in the aegilops-triticum alliance. Genetics 171:323–332. doi:10.1534/genetics.105.041632
Article PubMed CAS Google Scholar - Dvorak J, Luo MC, Yang ZL (1998) Genetic evidence on the origin of Triticum aestivum L. In: Damania AB, Valkoun J, Willcox G, Qualset CO (eds) The origins of agriculture and crop domestication. ICARDA, Aleppo, Syria, pp 235–251
Google Scholar - Feltus FA, Wan J, Schulze SR, Estill JC, Jiang N, Paterson AH (2004) An SNP resource for rice genetics and breeding based on subspecies Indica and Japonica genome alignments. Genome Res 14:1812–1819. doi:10.1101/gr.2479404
Article PubMed CAS Google Scholar - Hayes P, Szucs P (2006) Disequilibrium and association in barley: Thinking outside the glass. Proc Natl Acad Sci USA 103:18385–18386. doi:10.1073/pnas.0609405103
Article PubMed CAS Google Scholar - Kruglyak L (1997) The use of a genetic map of biallelic markers in linkage studies. Nat Genet 17:21–24. doi:10.1038/ng0997-21
Article PubMed CAS Google Scholar - Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129. doi:10.1093/bioinformatics/bti282
Article PubMed CAS Google Scholar - Mantel NA (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
PubMed CAS Google Scholar - Nasu S, Suzuki J, Ohta R, Hasegawa K, Yui R, Kitazawa N et al (2002) Search for and analysis of single nucleotide polymorphisms (SNPs) in rice (Oryza sativa, Oryza rufipogon) and establishment of SNP markers. DNA Res 9:163–171. doi:10.1093/dnares/9.5.163
Article PubMed CAS Google Scholar - Rafalski A (2002) Applications of single nucleotide polymorphism in crop genetics. Curr Opin Plant Biol 5:94–100. doi:10.1016/S1369-5266(02)00240-6
Article PubMed CAS Google Scholar - Ravel C, Praud S, Murigneux A, Canaguier A, Sapet F, Samson D et al (2006) Single-nucleotide polymorphism frequency in a set of selected lines of bread wheat (Triticum aestivum L.). Genome 49:1131–1139. doi:10.1139/G06-067
Article PubMed CAS Google Scholar - Rostoks N, Mudie S, Cardle L, Russell J, Ramsay L, Booth A et al (2005) Genome-wide SNP discovery and linkage analysis in barley based on genes responsive to abiotic stress. Mol Genet Genomics 274:515–527. doi:10.1007/s00438-005-0046-z
Article PubMed CAS Google Scholar - Rostoks N, Ramsay L, MacKenzie K, Cardle L, Bhat PR, Roose ML et al (2006) Recent history of artificial outcrossing facilitates whole-genome association mapping in elite inbred crop varieties. Proc Natl Acad Sci USA 103:18656–18661. doi:10.1073/pnas.0606133103
Article PubMed CAS Google Scholar - Russell J, Booth A, Fuller J, Harrower B, Hedley P, Machray G et al (2004) A comparison of sequence-based polymorphism and haplotype content in transcribed and anonymous regions of the barley genome. Genome 47:389–398
Article PubMed CAS Google Scholar - Schneider K, Kulosa D, Soerensen TR, Möhring S, Heine M, Durstewitz G, Polley A, Weber E, Jamsari, Lein J, Hohmann U, Tahiro E, Weisshaar B, Schulz B, Koch G, Jung C, Ganal M (2007) Analysis of DNA polymorphisms in sugar beet (Beta vulgaris L.) and development of an SNP-based map of expressed genes. Theor Appl Genet doi: 10.1007/s00122-007-0591-4
- Sears ER (1954) The aneuploids of common wheat. Mo Agric Exp Stn Res Bull 572:1–59
Google Scholar - Somers DJ, Kirkpatrick R, Moniwa M, Walsh A (2003) Mining single-nucleotide polymorphisms from hexaploid wheat ESTs. Genome 49:431–437. doi:10.1139/g03-027
Article Google Scholar - Talbert LE, Smith LY, Blake NK (1998) More than one origin of hexaploid wheat is indicated by sequence comparison of low-copy DNA. Genome 41:402–407. doi:10.1139/gen-41-3-402
Article CAS Google Scholar - Thuillet AC, Bru D, David J, Roumet P, Santoni S, Sourdille P et al (2002) Direct estimation of mutation rate for 10 microsatellite loci in durum wheat, Triticum turgidum (L.) Thell. ssp. durum desf. Mol Biol Evol 19:122–125
PubMed CAS Google Scholar - Vroh Bi I, McMullen MD, Sanchez-Villeda H, Schroeder S, Gardiner J, Polacco M et al (2006) Single nucleotide polymorphism and insertion-deletion for genetic markers and anchoring the maize fingerprint contig physical map. Crop Sci 46:12–21. doi:10.2135/cropsci2004.0706
Article CAS Google Scholar - Weir BS (1996) Genetic data analysis II. Sinauer Associate, Inc., Sunderlands
Google Scholar - Zhu YL, Song QJ, Hyten DL, van Tassell CP, Matukumalli LK, Grimm DR et al (2003) Single-nucleotide polymorphisms in soybean. Genetics 163:1123–1134
PubMed CAS Google Scholar
Acknowledgements
This research was supported in part by the funds from the U.S. Department of Agriculture, Cooperative State Research, Education and Extension Service, Coordinated Agricultural Project grant number 2006-55606-16629 and NSF Grant No. DBI-0321757. We thank Dr. Jan Dvorak for facilitating early access to the SNP data generated by the NSF project and for his useful suggestions and ideas and to Iago Lowe for his thorough revision of the manuscript.
Author information
Authors and Affiliations
- USDA-ARS Biosciences Research Lab, 1605 Albrecht Blvd., Fargo, ND, 58105-5674, USA
Shiaoman Chao - Department of Plant Sciences, University of California, Davis, Davis, CA, 95616, USA
Wenjun Zhang, Yaqin Ma, Ming-Cheng Luo & Jorge Dubcovsky - Department of Plant Pathology, Kansas State University, Manhattan, KS, 66506, USA
Eduard Akhunov - Department of Plant and Soil Sciences, Montana State University, Bozeman, MT, 59717, USA
Jamie Sherman
Authors
- Shiaoman Chao
- Wenjun Zhang
- Eduard Akhunov
- Jamie Sherman
- Yaqin Ma
- Ming-Cheng Luo
- Jorge Dubcovsky
Corresponding author
Correspondence toShiaoman Chao.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Chao, S., Zhang, W., Akhunov, E. et al. Analysis of gene-derived SNP marker polymorphism in US wheat (Triticum aestivum L.) cultivars.Mol Breeding 23, 23–33 (2009). https://doi.org/10.1007/s11032-008-9210-6
- Received: 17 January 2008
- Accepted: 09 July 2008
- Published: 29 July 2008
- Issue date: January 2009
- DOI: https://doi.org/10.1007/s11032-008-9210-6