A robust DNA mechanical device controlled by hybridization topology (original) (raw)
- Letter
- Published: 03 January 2002
Nature volume 415, pages 62–65 (2002) Cite this article
- 5916 Accesses
- 767 Citations
- 15 Altmetric
- Metrics details
Abstract
Controlled mechanical movement in molecular-scale devices has been realized in a variety of systems—catenanes and rotaxanes1,2,3, chiroptical molecular switches4, molecular ratchets5 and DNA6—by exploiting conformational changes triggered by changes in redox potential or temperature, reversible binding of small molecules or ions, or irradiation. The incorporation of such devices into arrays7,8 could in principle lead to complex structural states suitable for nanorobotic applications, provided that individual devices can be addressed separately. But because the triggers commonly used tend to act equally on all the devices that are present, they will need to be localized very tightly. This could be readily achieved with devices that are controlled individually by separate and device-specific reagents. A trigger mechanism that allows such specific control is the reversible binding of DNA strands, thereby ‘fuelling’ conformational changes in a DNA machine9. Here we improve upon the initial prototype system that uses this mechanism but generates by-products9, by demonstrating a robust sequence-dependent rotary DNA device operating in a four-step cycle. We show that DNA strands control and fuel our device cycle by inducing the interconversion between two robust topological motifs, paranemic crossover (PX) DNA10,11 and its topoisomer JX2 DNA, in which one strand end is rotated relative to the other by 180°. We expect that a wide range of analogous yet distinct rotary devices can be created by changing the control strands and the device sequences to which they bind.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Schematic drawings of the device.
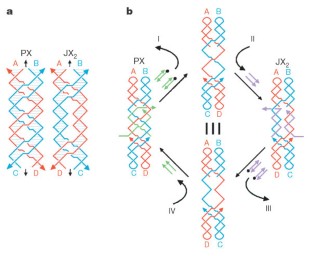
The alternative text for this image may have been generated using AI.
Figure 2: Gel evidence for the operation of the device.
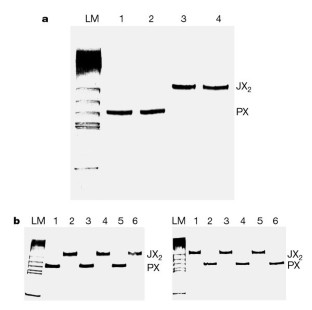
The alternative text for this image may have been generated using AI.
Figure 3: A highly simplified representation of the system used.
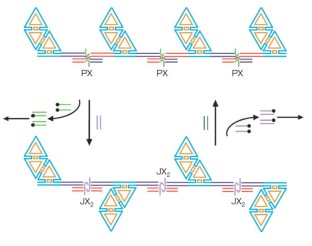
The alternative text for this image may have been generated using AI.
Figure 4: Evidence for the operation of the device, obtained from atomic force microscopy (AFM).
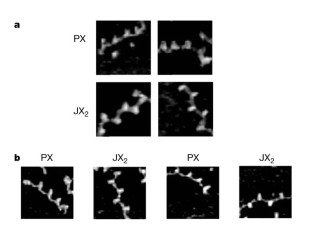
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
References
- Pease, A. R. et al. Switching devices based on interlocked molecules. Acc. Chem. Res. 34, 433–444 (2001).
Article CAS Google Scholar - Jimenez, M. C., Dietrich-Buchecker, C. & Sauvage, J.-P. Towards synthetic molecular muscles: contraction and stretching of a linear-rotaxane dimer. Angew. Chem. Int. Edn Engl. 39, 3284–3287 (2000).
Article CAS Google Scholar - Brouwer, A. M. et al. Photoinduction of fast, reversible translational motion in a hydrogen-bonded molecular shuttle. Science 291, 2124–2128 (2001).
Article ADS CAS Google Scholar - Koumura, N., Zijlstra, R. W. J., van Delden, R. A., Harada, N. & Feringa, B. L. Light-driven monodirectional molecular rotor. Nature 401, 152–155 (1999).
Article ADS CAS Google Scholar - Kelly, T. R., De Silva, H. & Silva, R. A. Unidirectional rotary motion in a molecular system. Nature 401, 150–152 (1999).
Article ADS CAS Google Scholar - Mao, C., Sun, W., Shen, Z. & Seeman, N. C. A DNA nanomechanical device based on the B-Z transition. Nature 397, 144–146 (1999).
Article ADS CAS Google Scholar - Winfree, E., Liu, F., Wenzler, L. A. & Seeman, N. C. Design and self-assembly of two-dimensional DNA crystals. Nature 394, 539–544 (1998).
Article ADS CAS Google Scholar - LaBean, T. H. et al. The construction, analysis, ligation and self-assembly of DNA triple crossover complexes. J. Am. Chem. Soc. 122, 1848–1860 (2000).
Article CAS Google Scholar - Yurke, B., Turberfield, A. J., Mills, A. P. Jr, Simmel, F. C. & Neumann, J. L. A DNA-fuelled molecular machine made of DNA. Nature 406, 605–608 (2000).
Article ADS CAS Google Scholar - Seeman, N. C. DNA nicks and nodes and nanotechnology. Nano Lett. 1, 22–26 (2001).
Article ADS CAS Google Scholar - Shen, Z. DNA Polycrossover Molecules and their Applications in Homology Recognition. Thesis, New York Univ. (1999).
Google Scholar - Fu, T.-J. & Seeman, N. C. DNA double crossover structures. Biochemistry 32, 3211–3220 (1993).
Article CAS Google Scholar - Seeman, N. C. De novo design of sequences for nucleic acid structure engineering. J. Biomol. Struct. Dyn. 8, 573–581 (1990).
Article CAS Google Scholar - Caruthers, M. H. Gene synthesis machines: DNA chemistry and its uses. Science 230, 281–285 (1985).
Article ADS CAS Google Scholar - Sun, W., Mao, C., Iwasaki, H., Kemper, B. & Seeman, N. C. No braiding of Holliday junctions in positively supercoiled DNA molecules. J. Mol. Biol. 294, 683–699 (1999).
Article CAS Google Scholar - Yang, X., Wenzler, L. A., Qi, J., Li, X. & Seeman, N. C. Ligation of DNA triangles containing double crossover molecules. J. Am. Chem. Soc. 120, 9779–9786 (1998).
Article CAS Google Scholar
Acknowledgements
We thank R. Sha for discussions. This work was supported by the Office of Naval Research, the National Institute of General Medical Sciences, the National Science Foundation/DARPA, the Information Directorate of the Air Force Research Laboratory located at Rome, New York, and the National Science Foundation.
Author information
Authors and Affiliations
- Department of Chemistry, New York University, New York, 10003, New York, USA
Hao Yan, Xiaoping Zhang, Zhiyong Shen & Nadrian C. Seeman
Authors
- Hao Yan
- Xiaoping Zhang
- Zhiyong Shen
- Nadrian C. Seeman
Corresponding author
Correspondence toNadrian C. Seeman.
Supplementary information
Rights and permissions
About this article
Cite this article
Yan, H., Zhang, X., Shen, Z. et al. A robust DNA mechanical device controlled by hybridization topology.Nature 415, 62–65 (2002). https://doi.org/10.1038/415062a
- Received: 26 April 2001
- Accepted: 08 November 2001
- Issue date: 03 January 2002
- DOI: https://doi.org/10.1038/415062a
This article is cited by
The biological applications of DNA nanomaterials: current challenges and future directions
- Wenjuan Ma
- Yuxi Zhan
- Yunfeng Lin
Signal Transduction and Targeted Therapy (2021)
Self-assembled Möbius strips with controlled helicity
- Guanghui Ouyang
- Lukang Ji
- Minghua Liu
Nature Communications (2020)
Framework Nucleic Acids for Cell Imaging and Therapy
- Zhilei Ge
- Qian Li
- Chunhai Fan
Chemical Research in Chinese Universities (2020)