Resilient circadian oscillator revealed in individual cyanobacteria (original) (raw)
- Letter
- Published: 01 July 2004
Nature volume 430, pages 81–85 (2004) Cite this article
- 3617 Accesses
- 240 Citations
- 12 Altmetric
- Metrics details
Abstract
Circadian oscillators, which provide internal daily periodicity, are found in a variety of living organisms, including mammals, insects, plants, fungi and cyanobacteria1. Remarkably, these biochemical oscillators are resilient to external and internal modifications, such as temperature and cell division cycles. They have to be ‘fluctuation (noise) resistant’2 because relative fluctuations in the number of messenger RNA and protein molecules forming the intracellular oscillators are likely to be large. In multicellular organisms, the strong temporal stability of circadian clocks, despite molecular fluctuations, can easily be explained by intercellular interactions3,4,5. Here we study circadian rhythms and their stability in unicellular cyanobacteria Synechoccocus elongatus. Low-light-level microscopy has allowed us to measure gene expression under circadian control in single bacteria, showing that the circadian clock is indeed a property of individual cells. Our measurements show that the oscillators have a strong temporal stability with a correlation time of several months. In contrast to many circadian clocks in multicellular organisms, this stability seems to be ensured by the intracellular biochemical network, because the interactions between oscillators seem to be negligible.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Circadian oscillation of bioluminescence in individual bacteria.
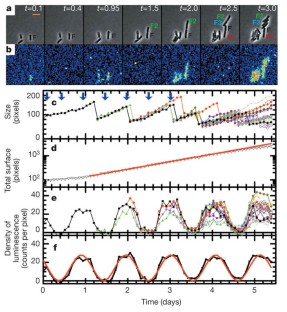
The alternative text for this image may have been generated using AI.
Figure 2: Growing micro-colonies of cyanobacteria, oscillating with different phases.
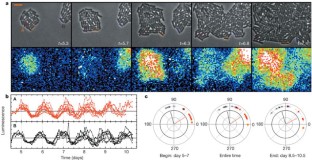
The alternative text for this image may have been generated using AI.
Figure 3: Temporal evolution of individual oscillators' phase is independent of close proximity.
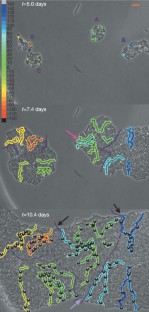
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
References
- Young, M. W. & Kay, S. A. Time zones: a comparative genetics of circadian clocks. Nature Rev. Genet. 2, 702–715 (2001)
Article CAS Google Scholar - Barkai, N. & Leibler, S. Circadian clocks limited by noise. Nature 403, 267–268 (2000)
Article ADS CAS Google Scholar - Pikovsky, A., Rosenblum, M. & Kurths, J. Synchronization: A Universal Concept in Nonlinear Sciences (Cambridge Univ. Press, Cambridge, 2001)
Book Google Scholar - Liu, C., Weaver, D. R., Strogatz, S. H. & Reppert, S. M. Cellular construction of a circadian clock: period determination in the suprachiasmatic nuclei. Cell 91, 855–860 (1997)
Article CAS Google Scholar - Yamaguchi, S. et al. Synchronization of cellular clocks in the suprachiasmatic nucleus. Science 302, 1408–1412 (2003)
Article ADS CAS Google Scholar - Kondo, T. et al. Circadian rhythms in prokaryotes: luciferase as a reporter of circadian gene expression in cyanobacteria. Proc. Natl Acad. Sci. USA 90, 5672–5676 (1993)
Article ADS CAS Google Scholar - Liu, Y. et al. Circadian orchestration of gene expression in cyanobacteria. Genes Dev. 9, 1469–1478 (1995)
Article CAS Google Scholar - Katayama, M., Tsinoremas, N. F., Kondo, T. & Golden, S. S. cpmA, a gene involved in an output pathway of the cyanobacterial circadian system. J. Bacteriol. 181, 3516–3524 (1999)
CAS PubMed PubMed Central Google Scholar - Tsinoremas, N. F., Schaefer, M. R. & Golden, S. S. Blue and red light reversibly control psbA expression in the cyanobacterium Synechococcus sp. strain PCC 7942. J. Biol. Chem. 269, 16143–16147 (1994)
CAS PubMed Google Scholar - Kondo, T. et al. Circadian rhythms in rapidly dividing cyanobacteria. Science 275, 224–227 (1997)
Article CAS Google Scholar - Mori, T. & Johnson, C. H. Independence of circadian timing from cell division in cyanobacteria. J. Bacteriol. 183, 2439–2444 (2001)
Article CAS Google Scholar - Mori, T., Binder, B. & Johnson, C. H. Circadian gating of cell division in cyanobacteria growing with average doubling times of less than 24 hours. Proc. Natl Acad. Sci. USA 93, 10183–10188 (1996)
Article ADS CAS Google Scholar - Gaspard, P. The correlation time of mesoscopic chemical clocks. J. Chem. Phys. 117, 8905–8916 (2002)
Article ADS CAS Google Scholar - Elowitz, M. B., Levine, A. J., Siggia, E. D. & Swain, P. S. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002)
Article ADS CAS Google Scholar - Kiss, I. Z., Zhai, Y. & Hudson, J. L. Emerging coherence in a population of chemical oscillators. Science 296, 1676–1678 (2002)
Article ADS CAS Google Scholar - Levine, J. D., Funes, P., Dowse, H. B. & Hall, J. C. Resetting the circadian clock by social experience in Drosophila melanogaster. Science 298, 2010–2012 (2002)
Article ADS CAS Google Scholar - Winfree, A. T. The Geometry of Biological Time, 2nd edn (Springer, New York, 2001)
Book Google Scholar - Goldbeter, A. Computational approaches to cellular rhythms. Nature 420, 238–245 (2002)
Article ADS CAS Google Scholar - Gonze, D., Halloy, J. & Goldbeter, A. Robustness of circadian rhythms with respect to molecular noise. Proc. Natl Acad. Sci. USA 99, 673–678 (2002)
Article ADS CAS Google Scholar
Acknowledgements
We thank S. Golden for the AMC412 strain and for advice, D. Peoples for advice and technical assistance, B. Houchmandzadeh, J. Paulsson, J. Vilar, C. Weitz and M. Young for discussions, and N. Questembert-Balaban, E. Kussell and M. Vallade for comments on the manuscript. This work was supported partially by Princeton University through the Lewis Thomas Fellowship (I.M.), the National Institutes of Health, the Howard Hughes Medical Institute and the Centre National de Recherche Scientifique through an ATIP and an AC ‘Dynamique et réactivité des assemblages biologiques’.
Author information
Author notes
- Weihong Hsing
Present address: Johnson & Johnson PRD, PO Box 300, Raritan, New Jersey, 08869, USA
Authors and Affiliations
- Laboratoire de Spectrométrie Physique, Université Joseph Fourier – Grenoble I, BP87, 38402, St-Martin d'Hères Cédex, France
Irina Mihalcescu - Department of Molecular Biology, Princeton University, Princeton, New Jersey, 08544, USA
Weihong Hsing - Laboratory of Living Matter and Centre for Studies in Physics and Biology, The Rockefeller University, 1230 York Avenue, New York, 10021, USA
Stanislas Leibler
Authors
- Irina Mihalcescu
- Weihong Hsing
- Stanislas Leibler
Corresponding author
Correspondence toIrina Mihalcescu.
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Discussion (download PDF )
This has four sections containing discussions and Supplementary Figures S1 to S7. 1. Circadian oscillations of individual bacteria; 2. Estimation of the oscillator noise; 3. Extraction of the detection noise; 4. Detection limit of the coupling strength between oscillators. (PDF 1599 kb)
Rights and permissions
About this article
Cite this article
Mihalcescu, I., Hsing, W. & Leibler, S. Resilient circadian oscillator revealed in individual cyanobacteria.Nature 430, 81–85 (2004). https://doi.org/10.1038/nature02533
- Received: 23 November 2003
- Accepted: 30 March 2004
- Issue date: 01 July 2004
- DOI: https://doi.org/10.1038/nature02533