Robustness–epistasis link shapes the fitness landscape of a randomly drifting protein (original) (raw)
- Letter
- Published: 19 November 2006
Nature volume 444, pages 929–932 (2006)Cite this article
- 6593 Accesses
- 434 Citations
- 26 Altmetric
- Metrics details
Abstract
The distribution of fitness effects of protein mutations is still unknown1,2. Of particular interest is whether accumulating deleterious mutations interact, and how the resulting epistatic effects shape the protein’s fitness landscape. Here we apply a model system in which bacterial fitness correlates with the enzymatic activity of TEM-1 β-lactamase (antibiotic degradation). Subjecting TEM-1 to random mutational drift and purifying selection (to purge deleterious mutations) produced changes in its fitness landscape indicative of negative epistasis; that is, the combined deleterious effects of mutations were, on average, larger than expected from the multiplication of their individual effects. As observed in computational systems3,4,5, negative epistasis was tightly associated with higher tolerance to mutations (robustness). Thus, under a low selection pressure, a large fraction of mutations was initially tolerated (high robustness), but as mutations accumulated, their fitness toll increased, resulting in the observed negative epistasis. These findings, supported by FoldX stability computations of the mutational effects6, prompt a new model in which the mutational robustness (or neutrality) observed in proteins, and other biological systems, is due primarily to a stability margin, or threshold, that buffers the deleterious physico-chemical effects of mutations on fitness. Threshold robustness is inherently epistatic—once the stability threshold is exhausted, the deleterious effects of mutations become fully pronounced, thereby making proteins far less robust than generally assumed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Negative epistasis underlines the random drift of TEM-1.
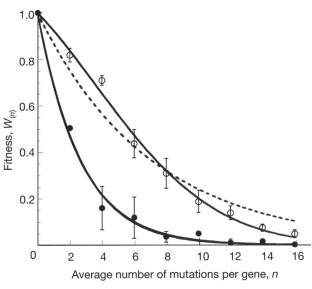
The alternative text for this image may have been generated using AI.
Figure 2: The correlation between mutational robustness and negative epistasis.
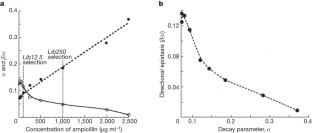
The alternative text for this image may have been generated using AI.
Figure 3: The physico-chemical and fitness changes accompanying random drifts.
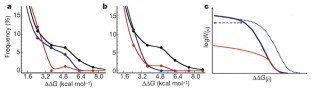
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
References
- Pal, C., Papp, B. & Lercher, M. J. An integrated view of protein evolution. Nature Rev. Genet. 7, 337–348 (2006)
Article CAS Google Scholar - DePristo, M. A., Weinreich, D. M. & Hartl, D. L. Missense meanderings in sequence space: a biophysical view of protein evolution. Nature Rev. Genet. 6, 678–687 (2005)
Article CAS Google Scholar - Wilke, C. O. & Adami, C. Interaction between directional epistasis and average mutational effects. Proc. R. Soc. Lond. B 268, 1469–1474 (2001)
Article CAS Google Scholar - Azevedo, R. B., Lohaus, R., Srinivasan, S., Dang, K. K. & Burch, C. L. Sexual reproduction selects for robustness and negative epistasis in artificial gene networks. Nature 440, 87–90 (2006)
Article ADS CAS Google Scholar - Elena, S. F., Carrasco, P., Daros, J. A. & Sanjuan, R. Mechanisms of genetic robustness in RNA viruses. EMBO Rep. 7, 168–173 (2006)
Article CAS Google Scholar - Guerois, R., Nielsen, J. E. & Serrano, L. Predicting changes in the stability of proteins and protein complexes: a study of more than 1000 mutations. J. Mol. Biol. 320, 369–387 (2002)
Article CAS Google Scholar - Wolf, J. B., Brodie, E. D. & Wade, M. J. Epistasis and the evolutionary process (Oxford Univ. Press, Oxford, 2000)
Google Scholar - Horovitz, A. Double-mutant cycles: a powerful tool for analyzing protein structure and function. Fold. Des. 1, R121–R126 (1996)
Article CAS Google Scholar - Charlesworth, B. Mutation–selection balance and the evolutionary advantage of sex and recombination. Genet. Res. 55, 199–221 (1990)
Article CAS Google Scholar - Bloom, J. D. et al. Thermodynamic prediction of protein neutrality. Proc. Natl Acad. Sci. USA 102, 606–611 (2005)
Article ADS CAS Google Scholar - Wagner, G. P., Laubichler, M. D. & Bagheri-Chaichian, H. Genetic measurement of theory of epistatic effects. Genetica 102–103, 569–580 (1998)
Article Google Scholar - Shafikhani, S., Siegel, R. A., Ferrari, E. & Schellenberger, V. Generation of large libraries of random mutants in Bacillus subtilis by PCR-based plasmid multimerization. Biotechniques 23, 304–310 (1997)
Article CAS Google Scholar - Daugherty, P. S., Chen, G., Iverson, B. L. & Georgiou, G. Quantitative analysis of the effect of the mutation frequency on the affinity maturation of single chain Fv antibodies. Proc. Natl Acad. Sci. USA 97, 2029–2034 (2000)
Article ADS CAS Google Scholar - Guo, H. H., Choe, J. & Loeb, L. A. Protein tolerance to random amino acid change. Proc. Natl Acad. Sci. USA 101, 9205–9210 (2004)
Article ADS CAS Google Scholar - Bowie, J. U., Reidhaar-Olson, J. F., Lim, W. A. & Sauer, R. T. Deciphering the message in protein sequences: tolerance to amino acid substitutions. Science 247, 1306–1310 (1990)
Article ADS CAS Google Scholar - Sauer, U. H., San, D. P. & Matthews, B. W. Tolerance of T4 lysozyme to proline substitutions within the long interdomain α-helix illustrates the adaptability of proteins to potentially destabilizing lesions. J. Biol. Chem. 267, 2393–2399 (1992)
CAS PubMed Google Scholar - Huang, W., Petrosino, J., Hirsch, M., Shenkin, P. S. & Palzkill, T. Amino acid sequence determinants of β-lactamase structure and activity. J. Mol. Biol. 258, 688–703 (1996)
Article CAS Google Scholar - Wagner, A. Distributed robustness versus redundancy as causes of mutational robustness. BioEssays 27, 176–188 (2005)
Article CAS Google Scholar - Bloom, J. D., Labthavikul, S. T., Otey, C. R. & Arnold, F. A. Protein stability promotes evolvability. Proc. Natl Acad. Sci. USA 103, 5869–5874 (2006)
Article ADS CAS Google Scholar - Kondrashov, A. S. Deleterious mutations and the evolution of sexual reproduction. Nature 336, 435–440 (1988)
Article ADS CAS Google Scholar - Lynch, M., Burger, R., Butcher, D. & Gabriel, W. The mutational meltdown in asexual populations. J. Hered. 84, 339–344 (1993)
Article CAS Google Scholar - Kiel, C. & Serrano, L. The ubiquitin domain superfold: structure-based sequence alignments and characterization of binding epitopes. J. Mol. Biol. 355, 821–844 (2006)
Article CAS Google Scholar
Acknowledgements
We thank F. Kondrashov for insights and inspiration, L. Serrano, U. Alon and G. Schreiber for contributions, and our Department, Institute and the Feinberg Graduate School, for uncompromising support. Author Contributions S.B. designed, performed and analysed the experiments. M.S. and R.B. provided technical assistance. N.T. performed the FoldX computations and helped devise the model. D.S.T. designed and supervised the experiments, and devised the model. S.B. and D.S.T. wrote the manuscript.
Author information
Authors and Affiliations
- Department of Biological Chemistry, Weizmann Institute of Science, Rehovot, 76100, Israel
Shimon Bershtein, Michal Segal, Roy Bekerman, Nobuhiko Tokuriki & Dan S. Tawfik
Authors
- Shimon Bershtein
- Michal Segal
- Roy Bekerman
- Nobuhiko Tokuriki
- Dan S. Tawfik
Corresponding author
Correspondence toDan S. Tawfik.
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Bershtein, S., Segal, M., Bekerman, R. et al. Robustness–epistasis link shapes the fitness landscape of a randomly drifting protein.Nature 444, 929–932 (2006). https://doi.org/10.1038/nature05385
- Received: 01 September 2006
- Accepted: 27 October 2006
- Published: 19 November 2006
- Issue date: 14 December 2006
- DOI: https://doi.org/10.1038/nature05385