Spontaneous network formation among cooperative RNA replicators (original) (raw)
- Article
- Published: 17 October 2012
- Michael L. Manapat2,
- Irene A. Chen3 nAff5,
- Ramon Xulvi-Brunet3,
- Eric J. Hayden4 &
- …
- Niles Lehman1
Nature volume 491, pages 72–77 (2012)Cite this article
- 12k Accesses
- 363 Citations
- 136 Altmetric
- Metrics details
Subjects
This article has been updated
Abstract
The origins of life on Earth required the establishment of self-replicating chemical systems capable of maintaining and evolving biological information. In an RNA world, single self-replicating RNAs would have faced the extreme challenge of possessing a mutation rate low enough both to sustain their own information and to compete successfully against molecular parasites with limited evolvability. Thus theoretical analyses suggest that networks of interacting molecules were more likely to develop and sustain life-like behaviour. Here we show that mixtures of RNA fragments that self-assemble into self-replicating ribozymes spontaneously form cooperative catalytic cycles and networks. We find that a specific three-membered network has highly cooperative growth dynamics. When such cooperative networks are competed directly against selfish autocatalytic cycles, the former grow faster, indicating an intrinsic ability of RNA populations to evolve greater complexity through cooperation. We can observe the evolvability of networks through in vitro selection. Our experiments highlight the advantages of cooperative behaviour even at the molecular stages of nascent life.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Cooperative covalent assembly of recombinase ribozymes.
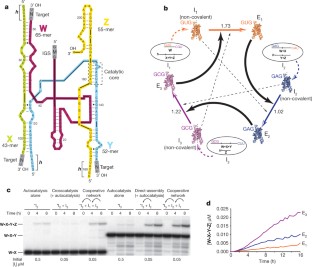
The alternative text for this image may have been generated using AI.
Figure 2: Cooperative chemistry out-competes selfish chemistry when directly competed.
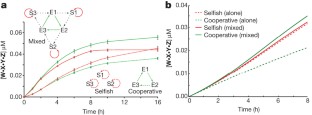
The alternative text for this image may have been generated using AI.
Figure 3: The randomization experiment.
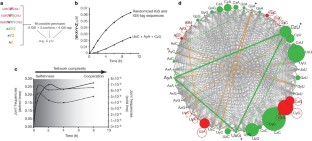
The alternative text for this image may have been generated using AI.
Figure 4: The serial transfer experiment.
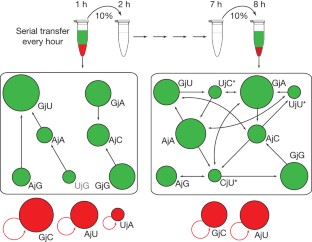
The alternative text for this image may have been generated using AI.
Figure 5: Growth curve of a four-piece system.
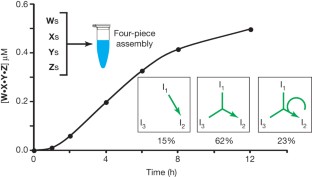
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
Change history
31 October 2012
A minor typo in Fig. 1 was corrected.
References
- Joyce, G. F. RNA evolution and the origins of life. Nature 338, 217–224 (1989)
Article ADS CAS Google Scholar - Zaher, H. S. & Unrau, P. J. Selection of an improved RNA polymerase ribozyme with superior extension and fidelity. RNA 13, 1017–1026 (2007)
Article CAS Google Scholar - Wochner, A., Attwater, J., Coulson, A. & Holliger, P. Ribozyme-catalyzed transcription of an active ribozyme. Science 332, 209–212 (2011)
Article ADS CAS Google Scholar - Eigen, M. & Schuster, P. The hypercycle. A principle of natural self-organization. Part A: emergence of the hypercycle. Naturwissenschaften 64, 541–565 (1977)
Article ADS CAS Google Scholar - Kauffman, S. A. The Origins of Order: Self-Organization and Selection in Evolution (Oxford Univ. Press, 1993)
Google Scholar - Sievers, D. & von Kiedrowski, G. Self-replication of complementary nucleotide-based oligomers. Nature 369, 221–224 (1994)
Article ADS CAS Google Scholar - Levy, M. & Ellington, A. D. The descent of polymerization. Nature Struct. Biol. 8, 580–582 (2001)
Article CAS Google Scholar - Szathmáry, E. The origin of replicators and reproducers. Phil. Trans. Royal Soc. B 361, 1761–1776 (2006)
Article Google Scholar - Nowak, M. A. & Ohtsuki, H. Preevolutionary dynamics and the origin of evolution. Proc. Natl Acad. Sci. USA 105, 14924–14927 (2008)
Article ADS CAS Google Scholar - Eigen, M. Selforganization of matter and the evolution of biological macromolecules. Naturwissenschaften 58, 465–523 (1971)
Article ADS CAS Google Scholar - Maynard Smith, J. Hypercycles and the origin of life. Nature 280, 445–446 (1979)
Article Google Scholar - Kun, Á., Santos, M. & Szathmáry, E. Real ribozymes suggest a relaxed error threshold. Nature Genet. 37, 1008–1011 (2005)
Article CAS Google Scholar - Doudna, J. A. & Szostak, J. W. RNA-catalysed synthesis of complementary-strand RNA. Nature 339, 519–522 (1989)
Article ADS CAS Google Scholar - Joyce, G. F. Forty years of in vitro evolution. Angew. Chem. Int. Ed. 46, 6420–6436 (2007)
Article CAS Google Scholar - Lincoln, T. A. & Joyce, G. F. Self-sustained replication of an RNA enzyme. Science 323, 1229–1232 (2009)
Article ADS CAS Google Scholar - Levy, M. & Ellington, A. D. Exponential growth by cross-catalytic cleavage of deoxyribozymogens. Proc. Natl Acad. Sci. USA 100, 6416–6421 (2003)
Article ADS CAS Google Scholar - Lee, D. H., Severin, K., Yokobayashi, Y. & Ghadiri, M. R. Emergence of symbiosis in peptide self-replication through a hypercyclic network. Nature 390, 591–594 (1997)
Article ADS CAS Google Scholar - Lee, D. H., Severin, K. & Ghadiri, M. R. Autocatalytic networks: the transition from molecular self-replication to molecular ecosystems. Curr. Opin. Chem. Biol. 1, 491–496 (1997)
Article CAS Google Scholar - Voytek, S. B. & Joyce, G. F. Niche partitioning in the coevolution of 2 distinct RNA enzymes. Proc. Natl Acad. Sci. USA 106, 7780–7785 (2009)
Article ADS CAS Google Scholar - Kim, D.-E. & Joyce, G. F. Cross-catalytic replication of an RNA ligase ribozyme. Chem. Biol. 11, 1505–1512 (2004)
Article CAS Google Scholar - Eigen, M. & Schuster, P. The hypercycle. A principle of natural self-organization. Part C: the realistic hypercycle. Naturwissenschaften 65, 341–369 (1978)
Article ADS CAS Google Scholar - Reinhold-Hurek, B. & Shub, D. A. Self-splicing introns in tRNA genes of widely divergent bacteria. Nature 357, 173–176 (1992)
Article ADS CAS Google Scholar - Hayden, E. J. & Lehman, N. Self-assembly of a group I intron from inactive oligonucleotide fragments. Chem. Biol. 13, 909–918 (2006)
Article CAS Google Scholar - Hayden, E. J., von Kiedrowski, G. & Lehman, N. Systems chemistry on ribozyme self-construction: evidence for anabolic autocatalysis in a recombination network. Angew. Chem. Int. Ed. 47, 8424–8428 (2008)
Article CAS Google Scholar - Draper, W. E., Hayden, E. J. & Lehman, N. Mechanisms of covalent self-assembly of the Azoarcus ribozyme from four fragment oligonucleotides. Nucleic Acids Res. 36, 520–531 (2008)
Article CAS Google Scholar - Nowak, M. A. Evolutionary Dynamics: Exploring the Equations of Life (Harvard Univ. Press, 2006)
Book Google Scholar - Hordijk, W. & Steel, M. Detecting autocatalytic, self-containing sets in chemical reaction systems. J. Theor. Biol. 227, 451–461 (2004)
Article CAS Google Scholar - Weiner, A. M. & Maizels, N. 3′ terminal tRNA-like structures tag genomic RNA molecules for replication: implications for the origin of protein synthesis. Proc. Natl Acad. Sci. USA 84, 7383–7387 (1987)
Article ADS CAS Google Scholar - Boerlijst, M. C. & Hogeweg, P. Spiral wave structures in prebiotic evolution: hypercycles stable against parasites. Physica D 48, 17–28 (1991)
Article ADS Google Scholar - Szathmáry, E. & Demeter, L. Group selection of early replicators and the origin of life. J. Theor. Biol. 128, 463–486 (1987)
Article Google Scholar - Lynch, M., Burger, R., Butcher, D. & Gabriel, W. The mutational meltdown in asexual populations. J. Hered. 84, 339–344 (1993)
Article CAS Google Scholar - Eigen, M. & Schuster, P. The hypercycle. A principle of natural self-organization. Part B: the abstract hypercycle. Naturwissenschaften 65, 7–41 (1978)
Article ADS Google Scholar
Acknowledgements
We would like to thank A. Burton, R. Ghadiri, P. Higgs, B. Larson, K. Chacón and A. López García de Lomana for help during preparation of this manuscript. This work was supported by NASA grant NNX10AR15G to N.L., the Center for Life in Extreme Environments at Portland State University, and a fellowship from the Human Frontier Science Program Organization to R.X.-B.
Author information
Author notes
- Irene A. Chen
Present address: Present address: Department of Chemistry and Biochemistry, University of California, Santa Barbara, California 93106, USA.,
Authors and Affiliations
- Department of Chemistry, Portland State University, PO Box 751, Portland, Oregon 97207, USA,
Nilesh Vaidya & Niles Lehman - School of Engineering and Applied Sciences and Program for Evolutionary Dynamics, Harvard University, Cambridge, 02138, Massachusetts, USA
Michael L. Manapat - FAS Center for Systems Biology, Harvard University, Cambridge, 02138, Massachusetts, USA
Irene A. Chen & Ramon Xulvi-Brunet - Department of Bioengineering, Stanford University, Stanford, 94305, California, USA
Eric J. Hayden
Authors
- Nilesh Vaidya
- Michael L. Manapat
- Irene A. Chen
- Ramon Xulvi-Brunet
- Eric J. Hayden
- Niles Lehman
Contributions
N.L. and N.V. conceived the basic idea and conducted the experiments; E.J.H. and I.A.C. contributed to the evaluation of the results; I.A.C., M.L.M. and R.X.-B. constructed the mathematical models; N.L. wrote the manuscript.
Corresponding author
Correspondence toNiles Lehman.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information (download PDF )
This file contains Supplementary Methods, Supplementary Text and Data, a Supplementary Discussion, Supplementary Figures 1-14, Supplementary Tables 1-3 and additional references. (PDF 4848 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Vaidya, N., Manapat, M., Chen, I. et al. Spontaneous network formation among cooperative RNA replicators.Nature 491, 72–77 (2012). https://doi.org/10.1038/nature11549
- Received: 11 March 2011
- Accepted: 06 September 2012
- Published: 17 October 2012
- Issue date: 01 November 2012
- DOI: https://doi.org/10.1038/nature11549
This article is cited by
Quasispecies productivity
- Esteban Domingo
- Guenther Witzany
The Science of Nature (2024)
Multispecies autocatalytic RNA reaction networks in coacervates
- Sandeep Ameta
- Manoj Kumar
- Shashi Thutupalli
Communications Chemistry (2023)
Self-organization of primitive metabolic cycles due to non-reciprocal interactions
- Vincent Ouazan-Reboul
- Jaime Agudo-Canalejo
- Ramin Golestanian
Nature Communications (2023)
Emergence of autocatalytic sets in a simple model of technological evolution
- Wim Hordijk
- Stuart Kauffman
- Roger Koppl
Journal of Evolutionary Economics (2023)
Charge-density reduction promotes ribozyme activity in RNA–peptide coacervates via RNA fluidization and magnesium partitioning
- Juan M. Iglesias-Artola
- Björn Drobot
- Moritz Kreysing
Nature Chemistry (2022)
Comments
Commenting on this article is now closed.
- Mike Miller 31 March 2013, 11:07
Thank you for an excellent article.
The following comments pertain to the application of this study to abiogenesis:
There are still several considerations before such an article should be cited as evidence for abiogenesis.
- Start with a pre-existing system, (The ~200-nucleotide (nt) Azoarcus group I intron ribozyme22 can be broken into fragments that can covalently self-assemble by catalysing recombination reactions in an autocatalytic fashion.
I'm sure someone will cite this as evidence of complex systems originating from simpler systems thus making abiogenesis appear as a sound scientific theory. Wait, what; they took an enzyme and fragmented it, and then noticed that many of the components have a similar function.... Is that similar to several indistinct components spontaneously forming and then organizing into a system that forms an enzyme? Not to mention the lack of thermocycler etc. Not to mention that these sequences do not even code for any proteins, which are hundreds of amino acids in length, and just by chance happen to have relevant cellular functions, and happen to be assembled with the adjunct of chaperones; and happen to facilitate the production of a membrane with modified lipids and integral proteins. In summary, abiogenesis remains as tangible as a man getting safely to the moon after being stranded on a desert island.
Editorial Summary
Cooperativity in an RNA world
In models of early life characterized by an RNA-only world, it has been suggested that life and evolution would be more easily achieved if the RNA molecules could interact, rather than function independently. They can. Niles Lehman and colleagues demonstrate the validity of the concept in vitro using a model system containing RNA fragments that can assemble into a ribozyme. The authors show that the cooperative networks formed by these fragments can outcompete self-catalytic RNA fragments. This work indicates that RNA populations have an intrinsic ability to evolve greater complexity through cooperation, and suggests that the benefits of such behaviour were established early in the development of life on Earth.