Mutations in pericentrin cause Seckel syndrome with defective ATR-dependent DNA damage signaling (original) (raw)
- Letter
- Published: 23 December 2007
- Sarah Walker2 na1,
- Carol-Anne Martin1,
- Paola Vagnarelli3,
- Tom Stiff2,
- Bertrand Vernay1,
- Nouriya Al Sanna4,
- Anand Saggar5,
- Ben Hamel6,
- William C Earnshaw3,
- Penny A Jeggo2,
- Andrew P Jackson1 &
- …
- Mark O'Driscoll2
Nature Genetics volume 40, pages 232–236 (2008)Cite this article
- 2808 Accesses
- 292 Citations
- 12 Altmetric
- Metrics details
Abstract
Large brain size is one of the defining characteristics of modern humans. Seckel syndrome (MIM 210600), a disorder of markedly reduced brain and body size1,2, is associated with defective ATR-dependent DNA damage signaling3. Only a single hypomorphic mutation of ATR has been identified in this genetically heterogeneous condition4. We now report that mutations in the gene encoding pericentrin (PCNT)—resulting in the loss of pericentrin from the centrosome, where it has key functions anchoring both structural and regulatory proteins—also cause Seckel syndrome5,6. Furthermore, we find that cells of individuals with Seckel syndrome due to mutations in PCNT (PCNT-Seckel) have defects in ATR-dependent checkpoint signaling, providing the first evidence linking a structural centrosomal protein with DNA damage signaling. These findings also suggest that other known microcephaly genes implicated in either DNA repair responses7 or centrosomal function8,9 may act in common developmental pathways determining human brain and body size.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Schematic of the Sckl4 critical region and the PCNT gene depicting location of identified mutations.
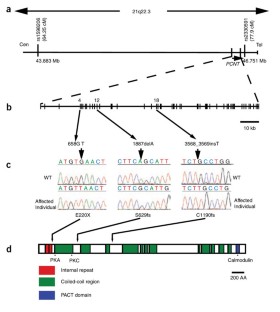
The alternative text for this image may have been generated using AI.
Figure 2: Pericentrin localization and function is disrupted in PCNT-Seckel cell lines.
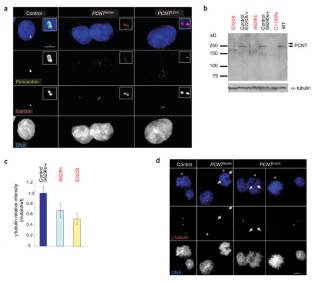
The alternative text for this image may have been generated using AI.
Figure 3: PCNT is required for ATR-dependent DNA damage signaling.
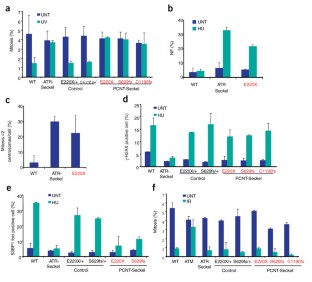
The alternative text for this image may have been generated using AI.
Figure 4: Model of pericentrin's role in ATR-dependent G2-M checkpoint arrest.
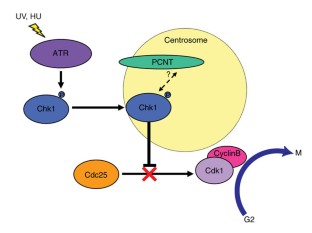
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
Accession codes
Accessions
GenBank/EMBL/DDBJ
References
- Majewski, F. et al. Studies of microcephalic primordial dwarfism I: approach to a delineation of the Seckel syndrome. Am. J. Med. Genet. 12, 7–21 (1982).
Article CAS Google Scholar - Seckel, H.P.G. Bird-headed Dwarfs: Studies in Developmental Anthropology Including Human Proportions (Charles C. Thomas, Springfield, Illinois, 1960).
Google Scholar - Alderton, G.K. et al. Seckel syndrome exhibits cellular features demonstrating defects in the ATR-signalling pathway. Hum. Mol. Genet. 13, 3127–3138 (2004).
Article CAS Google Scholar - O'Driscoll, M. et al. A splicing mutation affecting expression of ataxia-telangiectasia and Rad3-related protein (ATR) results in Seckel syndrome. Nat. Genet. 33, 497–501 (2003).
Article CAS Google Scholar - Diviani, D. et al. Pericentrin anchors protein kinase A at the centrosome through a newly identified RII-binding domain. Curr. Biol. 10, 417–420 (2000).
Article CAS Google Scholar - Zimmerman, W.C. et al. Mitosis-specific anchoring of gamma tubulin complexes by pericentrin controls spindle organization and mitotic entry. Mol. Biol. Cell 15, 3642–3657 (2004).
Article CAS Google Scholar - Alderton, G.K. et al. Regulation of mitotic entry by microcephalin and its overlap with ATR signalling. Nat. Cell Biol. 8, 725–733 (2006).
Article CAS Google Scholar - Bond, J. et al. ASPM is a major determinant of cerebral cortical size. Nat. Genet. 32, 316–320 (2002).
Article CAS Google Scholar - Bond, J. et al. A centrosomal mechanism involving CDK5RAP2 and CENPJ controls brain size. Nat. Genet. 37, 353–355 (2005).
Article CAS Google Scholar - Shiloh, Y. ATM and ATR: networking cellular responses to DNA damage. Curr. Opin. Genet. Dev. 11, 71–77 (2001).
Article CAS Google Scholar - Flory, M.R. et al. The centrosomal proteins pericentrin and kendrin are encoded by alternatively spliced products of one gene. Genomics 82, 401–405 (2003).
Article CAS Google Scholar - Miyoshi, K. et al. Characterization of pericentrin isoforms in vivo. Biochem. Biophys. Res. Commun. 351, 745–749 (2006).
Article CAS Google Scholar - Li, Q. et al. Kendrin/pericentrin-B, a centrosome protein with homology to pericentrin that complexes with PCM-1. J. Cell Sci. 114, 797–809 (2001).
CAS PubMed Google Scholar - Dictenberg, J.B. et al. Pericentrin and gamma-tubulin form a protein complex and are organized into a novel lattice at the centrosome. J. Cell Biol. 141, 163–174 (1998).
Article CAS Google Scholar - Doxsey, S.J. et al. Pericentrin, a highly conserved centrosome protein involved in microtubule organization. Cell 76, 639–650 (1994).
Article CAS Google Scholar - Purohit, A. et al. Direct interaction of pericentrin with cytoplasmic dynein light intermediate chain contributes to mitotic spindle organization. J. Cell Biol. 147, 481–492 (1999).
Article CAS Google Scholar - Martinez-Campos, M. et al. The Drosophila pericentrin-like protein is essential for cilia/flagella function, but appears to be dispensable for mitosis. J. Cell Biol. 165, 673–683 (2004).
Article CAS Google Scholar - Chen, D. et al. Centrosomal anchoring of protein kinase C betaII by pericentrin controls microtubule organization, spindle function, and cytokinesis. J. Biol. Chem. 279, 4829–4839 (2004).
Article CAS Google Scholar - Sengupta, S. et al. Functional interaction between BLM helicase and 53BP1 in a Chk1-mediated pathway during S-phase arrest. J. Cell Biol. 166, 801–813 (2004).
Article CAS Google Scholar - Woods, C.G. et al. Autosomal recessive primary microcephaly (MCPH): a review of clinical, molecular, and evolutionary findings. Am. J. Hum. Genet. 76, 717–728 (2005).
Article CAS Google Scholar - Ponting, C. et al. Evolution of primary microcephaly genes and the enlargement of primate brains. Curr. Opin. Genet. Dev. 15, 241–248 (2005).
Article CAS Google Scholar - Brown, P. et al. A new small-bodied hominin from the Late Pleistocene of Flores, Indonesia. Nature 431, 1055–1061 (2004).
Article CAS Google Scholar - Martin, R.D. et al. Flores hominid: new species or microcephalic dwarf? Anat. Rec. A Discov. Mol. Cell. Evol. Biol. 288, 1123–1145 (2006).
Article Google Scholar - Doxsey, S. et al. Centrosomes in cellular regulation. Annu. Rev. Cell Dev. Biol. 21, 411–434 (2005).
Article CAS Google Scholar - Zhang, S. et al. Centrosomal localization of DNA damage checkpoint proteins. J. Cell. Biochem. 101, 451–465 (2007).
Article CAS Google Scholar - Loffler, H. et al. DNA damage–induced accumulation of centrosomal Chk1 contributes to its checkpoint function. Cell Cycle 6, 2541–2518 (2007).
Article Google Scholar - Kramer, A. et al. Centrosome-associated Chk1 prevents premature activation of cyclin-B-Cdk1 kinase. Nat. Cell Biol. 6, 884–891 (2004).
Article Google Scholar - Niida, H. et al. Specific role of Chk1 phosphorylations in cell survival and checkpoint activation. Mol. Cell. Biol. 27, 2572–2581 (2007).
Article CAS Google Scholar - Ruschendorf, F. et al. ALOHOMORA: a tool for linkage analysis using 10K SNP array data. Bioinformatics 21, 2123–2125 (2005).
Article Google Scholar - Dodson, H. et al. Centrosome amplification induced by DNA damage occurs during a prolonged G2 phase and involves ATM. EMBO J. 23, 3864–3873 (2004).
Article CAS Google Scholar
Acknowledgements
We thank the families and their clinicians for their participation in this study; C. Hayward for contributing control samples; S. McKay and the MRC HGU core sequencing service for advice and technical support; C. Nicol for help with figure preparation; X. Fant, V. Van Heyningen and N. Hastie for discussions and comments; W. Fergusson (St. Mary's Hospital, Manchester) for LCL transformation; and A. Merdes and J. Salisbury for kindly sharing antibody reagents. A.P.J.'s laboratory is funded by the MRC, M.O'D.'s laboratory is funded by CRUK and the MRC, and P.A.J.'s laboratory is funded by the MRC, UK LRF, IACR and EU grants (FIGH-CT-200200207) (DNA repair) and FI6R-CT-2003-508842 (RiscRad). P.V. and W.C.E. are funded by the Wellcome Trust, of which W.C.E. is a Principal Research Fellow. A.P.J. is an MRC Senior Clinical Fellow and M.O'D. is a CRUK Senior Cancer Research Fellow.
Author information
Author notes
- Elen Griffith and Sarah Walker: These authors contributed equally to this work.
Authors and Affiliations
- Medical Research Council (MRC) Human Genetics Unit, Western General Hospital, Crewe Road, Edinburgh, EH4 2XU, UK
Elen Griffith, Carol-Anne Martin, Bertrand Vernay & Andrew P Jackson - Genome Damage and Stability Centre, University of Sussex, East Sussex, BN1 9RQ, UK
Sarah Walker, Tom Stiff, Penny A Jeggo & Mark O'Driscoll - Wellcome Trust Centre for Cell Biology, School of Biological Sciences, University of Edinburgh, Kings Buildings, Mayfield Road, Edinburgh, EH9 3JR, UK
Paola Vagnarelli & William C Earnshaw - Pediatric Services Division, Box 76, Dhahran Health Center, Saudi Arabia
Nouriya Al Sanna - Southwest Thames Regional Genetics Service, St. George's Hospital Medical School, London, SW17 0RE, UK
Anand Saggar - Department of Human Genetics 417, Radboud University Nijmegen Medical Center, Geert Grooteplein 20, Nijmegen, 6525GA, The Netherlands
Ben Hamel
Authors
- Elen Griffith
- Sarah Walker
- Carol-Anne Martin
- Paola Vagnarelli
- Tom Stiff
- Bertrand Vernay
- Nouriya Al Sanna
- Anand Saggar
- Ben Hamel
- William C Earnshaw
- Penny A Jeggo
- Andrew P Jackson
- Mark O'Driscoll
Contributions
E.G., C.-A.M. and A.P.J. performed the mutation screening and sequencing of controls; A.P.J., linkage analysis; P.V., immunostaining analysis of LCLs and C.-A.M., immunoblotting. S.W., T.S., M.O'D. and P.A.J. designed and performed the DNA damage response assays and RNAi experiments. N.A.S., A.S. and B.H. provided clinical samples and data. E.G. and A.P.J. wrote the paper with M.O'D., P.A.J., B.V. and W.C.E.
Corresponding author
Correspondence toAndrew P Jackson.
Supplementary information
Rights and permissions
About this article
Cite this article
Griffith, E., Walker, S., Martin, CA. et al. Mutations in pericentrin cause Seckel syndrome with defective ATR-dependent DNA damage signaling.Nat Genet 40, 232–236 (2008). https://doi.org/10.1038/ng.2007.80
- Received: 29 October 2007
- Accepted: 13 December 2007
- Published: 23 December 2007
- Issue date: February 2008
- DOI: https://doi.org/10.1038/ng.2007.80