The contribution of rare variants to risk of schizophrenia in individuals with and without intellectual disability (original) (raw)
References
- van Os, J. & Kapur, S. Schizophrenia. Lancet 374, 635–645 (2009).
Article CAS PubMed Google Scholar - American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders (DSM-5) (American Psychiatric Publishing, 2013).
- Tandon, R. et al. Definition and description of schizophrenia in the DSM-5. Schizophr. Res. 150, 3–10 (2013).
Article PubMed Google Scholar - Owen, M.J. New approaches to psychiatric diagnostic classification. Neuron 84, 564–571 (2014).
Article CAS PubMed Google Scholar - Owen, M.J., Sawa, A. & Mortensen, P.B. Schizophrenia. Lancet 388, 86–97 (2016).
Article PubMed PubMed Central Google Scholar - Howes, O.D. & Kapur, S. The dopamine hypothesis of schizophrenia: version III—the final common pathway. Schizophr. Bull. 35, 549–562 (2009).
Article PubMed PubMed Central Google Scholar - Pocklington, A.J. et al. Novel findings from CNVs implicate inhibitory and excitatory signaling complexes in schizophrenia. Neuron 86, 1203–1214 (2015).
Article CAS PubMed PubMed Central Google Scholar - Sekar, A. et al. Schizophrenia risk from complex variation of complement component 4. Nature 530, 177–183 (2016).
Article CAS PubMed PubMed Central Google Scholar - Owen, M.J., O'Donovan, M.C., Thapar, A. & Craddock, N. Neurodevelopmental hypothesis of schizophrenia. Br. J. Psychiatry 198, 173–175 (2011).
Article PubMed PubMed Central Google Scholar - Rapoport, J.L., Giedd, J.N. & Gogtay, N. Neurodevelopmental model of schizophrenia: update 2012. Mol. Psychiatry 17, 1228–1238 (2012).
Article CAS PubMed PubMed Central Google Scholar - Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 511, 421–427 (2014).
- Loh, P.-R. et al. Contrasting genetic architectures of schizophrenia and other complex diseases using fast variance-components analysis. Nat. Genet. 47, 1385–1392 (2015).
Article CAS PubMed PubMed Central Google Scholar - Fromer, M. et al. De novo mutations in schizophrenia implicate synaptic networks. Nature 506, 179–184 (2014).
Article CAS PubMed PubMed Central Google Scholar - Kirov, G. et al. De novo CNV analysis implicates specific abnormalities of postsynaptic signalling complexes in the pathogenesis of schizophrenia. Mol. Psychiatry 17, 142–153 (2012).
Article CAS PubMed Google Scholar - International Schizophrenia Consortium. Rare chromosomal deletions and duplications increase risk of schizophrenia. Nature 455, 237–241 (2008).
- Rees, E. et al. Analysis of copy number variations at 15 schizophrenia-associated loci. Br. J. Psychiatry 204, 108–114 (2014).
Article PubMed PubMed Central Google Scholar - Zhu, X., Need, A.C., Petrovski, S. & Goldstein, D.B. One gene, many neuropsychiatric disorders: lessons from Mendelian diseases. Nat. Neurosci. 17, 773–781 (2014).
Article CAS PubMed Google Scholar - Singh, T. et al. Rare loss-of-function variants in SETD1A are associated with schizophrenia and developmental disorders. Nat. Neurosci. 19, 571–577 (2016).
Article CAS PubMed PubMed Central Google Scholar - Szatkiewicz, J.P. et al. Copy number variation in schizophrenia in Sweden. Mol. Psychiatry 19, 762–773 (2014).
Article CAS PubMed PubMed Central Google Scholar - Network and Pathway Analysis Subgroup of Psychiatric Genomics Consortium. Psychiatric genome-wide association study analyses implicate neuronal, immune and histone pathways. Nat. Neurosci. 18, 199–209 (2015).
- Lee, S.H. et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat. Genet. 45, 984–994 (2013).
CAS PubMed Google Scholar - Robinson, E.B. et al. Genetic risk for autism spectrum disorders and neuropsychiatric variation in the general population. Nat. Genet. 48, 552–555 (2016).
Article CAS PubMed PubMed Central Google Scholar - Sanders, S.J. et al. Insights into autism spectrum disorder genomic architecture and biology from 71 risk loci. Neuron 87, 1215–1233 (2015).
Article CAS PubMed PubMed Central Google Scholar - Samocha, K.E. et al. A framework for the interpretation of de novo mutation in human disease. Nat. Genet. 46, 944–950 (2014).
Article CAS PubMed PubMed Central Google Scholar - Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Article CAS PubMed PubMed Central Google Scholar - Genovese, G. et al. Increased burden of ultra-rare protein-altering variants among 4,877 individuals with schizophrenia. Nat. Neurosci. 19, 1433–1441 (2016).
Article CAS PubMed PubMed Central Google Scholar - Kirov, G. et al. The penetrance of copy number variations for schizophrenia and developmental delay. Biol. Psychiatry 75, 378–385 (2014).
Article CAS PubMed Google Scholar - Iossifov, I. et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 515, 216–221 (2014).
Article CAS PubMed PubMed Central Google Scholar - Rees, E. et al. CNV analysis in a large schizophrenia sample implicates deletions at 16p12.1 and SLC1A1 and duplications at 1p36.33 and CGNL1. Hum. Mol. Genet. 23, 1669–1676 (2014).
Article CAS PubMed Google Scholar - Price, A.L. et al. Pooled association tests for rare variants in exon-resequencing studies. Am. J. Hum. Genet. 86, 832–838 (2010).
Article PubMed PubMed Central Google Scholar - Raychaudhuri, S. et al. Accurately assessing the risk of schizophrenia conferred by rare copy-number variation affecting genes with brain function. PLoS Genet. 6, e1001097 (2010).
Article PubMed PubMed Central Google Scholar - De Rubeis, S. et al. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 515, 209–215 (2014).
Article CAS PubMed PubMed Central Google Scholar - Firth, H.V. et al. DECIPHER: Database of Chromosomal Imbalance and Phenotype in Humans Using Ensembl Resources. Am. J. Hum. Genet. 84, 524–533 (2009).
Article CAS PubMed PubMed Central Google Scholar - Deciphering Developmental Disorders Study. Prevalence and architecture of de novo mutations in developmental disorders. Nature 542, 433–438 (2017).
- Purcell, S.M. et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature 506, 185–190 (2014).
Article CAS PubMed PubMed Central Google Scholar - Cotney, J. et al. The autism-associated chromatin modifier CHD8 regulates other autism risk genes during human neurodevelopment. Nat. Commun. 6, 6404 (2015).
Article CAS PubMed Google Scholar - Weyn-Vanhentenryck, S.M. et al. HITS-CLIP and integrative modeling define the Rbfox splicing-regulatory network linked to brain development and autism. Cell Rep. 6, 1139–1152 (2014).
Article CAS PubMed PubMed Central Google Scholar - Darnell, J.C. et al. FMRP stalls ribosomal translocation on mRNAs linked to synaptic function and autism. Cell 146, 247–261 (2011).
Article CAS PubMed PubMed Central Google Scholar - Ascano, M. Jr. et al. FMRP targets distinct mRNA sequence elements to regulate protein expression. Nature 492, 382–386 (2012).
Article CAS PubMed PubMed Central Google Scholar - Ganna, A. et al. Ultra-rare disruptive and damaging mutations influence educational attainment in the general population. Nat. Neurosci. 19, 1563–1565 (2016).
Article CAS PubMed PubMed Central Google Scholar - Deciphering Developmental Disorders Study. Large-scale discovery of novel genetic causes of developmental disorders. Nature 519, 223–228 (2015).
- Pardiñas, A.F. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and maintained by background selection. Preprint at bioRxiv http://dx.doi.org/10.1101/068593 (2016).
- Ben-Shachar, S. et al. 22q11.2 distal deletion: a recurrent genomic disorder distinct from DiGeorge syndrome and velocardiofacial syndrome. Am. J. Hum. Genet. 82, 214–221 (2008).
Article CAS PubMed PubMed Central Google Scholar - Michaelovsky, E. et al. Genotype–phenotype correlation in 22q11.2 deletion syndrome. BMC Med. Genet. 13, 122 (2012).
Article CAS PubMed PubMed Central Google Scholar - Guipponi, M. et al. Exome sequencing in 53 sporadic cases of schizophrenia identifies 18 putative candidate genes. PLoS One 9, e112745 (2014).
Article PubMed PubMed Central Google Scholar - Girard, S.L. et al. Increased exonic de novo mutation rate in individuals with schizophrenia. Nat. Genet. 43, 860–863 (2011).
Article CAS PubMed Google Scholar - McCarthy, S.E. et al. De novo mutations in schizophrenia implicate chromatin remodeling and support a genetic overlap with autism and intellectual disability. Mol. Psychiatry 19, 652–658 (2014).
Article CAS PubMed PubMed Central Google Scholar - Takata, A. et al. Loss-of-function variants in schizophrenia risk and SETD1A as a candidate susceptibility gene. Neuron 82, 773–780 (2014).
Article CAS PubMed PubMed Central Google Scholar - Xu, B. et al. Exome sequencing supports a de novo mutational paradigm for schizophrenia. Nat. Genet. 43, 864–868 (2011).
Article CAS PubMed PubMed Central Google Scholar - Xu, B. et al. De novo gene mutations highlight patterns of genetic and neural complexity in schizophrenia. Nat. Genet. 44, 1365–1369 (2012).
Article CAS PubMed PubMed Central Google Scholar - Do, R. et al. Exome sequencing identifies rare LDLR and APOA5 alleles conferring risk for myocardial infarction. Nature 518, 102–106 (2015).
Article CAS PubMed Google Scholar - Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Article CAS PubMed PubMed Central Google Scholar - Doody, G.A., Johnstone, E.C., Sanderson, T.L., Owens, D.G. & Muir, W.J. 'Pfropfschizophrenie' revisited. Schizophrenia in people with mild learning disability. Br. J. Psychiatry 173, 145–153 (1998).
Article CAS PubMed Google Scholar
Acknowledgements
We gratefully thank all participants in these studies. We thank T. Touloupoulou, M. Picchioni, C. Nosarti, F. Gaughran, and O. Howes for contributing clinical data used in this study. The UK10K project was funded by Wellcome Trust grant WT091310. The INTERVAL sequencing studies are funded by Wellcome Trust grant WT098051. T.S. is supported by the Williams College Dr. Herchel Smith Fellowship. A.P. is supported by Academy of Finland grants 251704 and 286500, NIMH grant U01MH105666, and the Sigrid Juselius Foundation. The work at Cardiff University was funded by Medical Research Council (MRC) Centre (G0801418) and Program (G0800509) grants. P.F.S. gratefully acknowledges support from the Swedish Research Council (Vetenskapsrådet, award D0886501). Creation of the Sweden schizophrenia study data was supported by NIMH grant R01 MH077139 and the Stanley Center of the Broad Institute. Participants in INTERVAL were recruited with the active collaboration of NHS Blood and Transplant England, which has supported fieldwork and other elements of the trial. DNA extraction and genotyping were funded by the National Institute of Health Research (NIHR), the NIHR BioResource, and the NIHR Cambridge Biomedical Research Centre. The academic coordinating center for INTERVAL was supported by core funding from the following: the NIHR Blood and Transplant Research Unit in Donor Health and Genomics, the UK MRC (G0800270), and the British Heart Foundation (SP/09/002). For the CNV analysis, we would like to acknowledge the contribution of data from outside sources: (i) Genetic Architecture of Smoking and Smoking Cessation accessed through dbGaP (study accession phs000404.v1.p1). Funding support for genotyping, which was performed at the Center for Inherited Disease Research (CIDR), was provided by 1 X01 HG005274-01. CIDR is fully funded through a federal contract from the National Institutes of Health to the Johns Hopkins University, contract number HHSN268200782096C. Assistance with genotype cleaning, as well as with general study coordination, was provided by the Gene Environment Association Studies (GENEVA) Coordinating Center (U01 HG004446). Funding support for collection of data sets and samples was provided by the Collaborative Genetic Study of Nicotine Dependence (COGEND; P01 CA089392) and the University of Wisconsin Transdisciplinary Tobacco Use Research Center (P50 DA019706 and P50 CA084724). (ii) High-Density SNP Association Analysis of Melanoma: Case–Control and Outcomes Investigation (dbGaP study accession phs000187.v1.p1). Research support to collect data and develop an application to support this project was provided by 3P50CA093459, 5P50CA097007, 5R01ES011740, and 5R01CA133996. (iii) Genetic Epidemiology of Refractive Error in the KORA Study (dbGaP study accession phs000303.v1.p1). Principal investigators: D. Stambolian (University of Pennsylvania) and H.E. Wichmann (Institut für Humangenetik, Helmholtz Zentrum München; National Eye Institute, National Institutes of Health). Funding was provided by R01 EY020483 from the National Institutes of Health. (iv) WTCCC2 study. Samples were downloaded from EGA (http://www.ebi.ac.uk/ega/) and include samples from the National Blood Donors Cohort (EGAD00000000024) and samples from the 1958 British Birth Cohort (EGAD00000000022). Funding for these projects was provided by the Wellcome Trust Case Control Consortium 2 project (085475/B/08/Z and 085475/Z/08/Z), the Wellcome Trust (072894/Z/03/Z, 090532/Z/09/Z, and 075491/Z/04/B), and NIMH grants (MH 41953 and MH083094).
Author information
Authors and Affiliations
- Wellcome Trust Sanger Institute, Wellcome Trust Genome Campus, Hinxton, UK
Tarjinder Singh & Jeffrey C Barrett - Division of Psychological Medicine and Clinical Neurosciences, MRC Centre for Neuropsychiatric Genetics and Genomics, School of Medicine, Cardiff University, Cardiff, UK
James T R Walters, Elliott Rees, Georg Kirov, Michael C O'Donovan & Michael J Owen - Division of Psychiatry, University of Edinburgh, Royal Edinburgh Hospital, Edinburgh, UK
Mandy Johnstone & Douglas Blackwood - University College London Genetics Institute, University College London, London, UK
David Curtis - Centre for Psychiatry, Barts and the London School of Medicine and Dentistry, London, UK
David Curtis - National Institute for Health and Welfare, Helsinki, Finland
Jaana Suvisaari & Minna Torniainen - Institute of Psychiatry, King's College London, London, UK
Conrad Iyegbe, Robin M Murray & Marta Di Forti - Centre for Cognitive Ageing and Cognitive Epidemiology, University of Edinburgh, Edinburgh, UK
Andrew M McIntosh - Department of Human Genetics, David Geffen School of Medicine, University of California, Los Angeles, Los Angeles, California, USA
Daniel Geschwind & Michael Gandal - Division of Psychiatry, University College London, London, UK
Elvira Bramon - Department of Medical Epidemiology and Biostatistics, Karolinska Institutet, Stockholm, Sweden
Christina M Hultman - Division of Psychiatric Genomics, Department of Psychiatry, Icahn School of Medicine at Mount Sinai, New York, New York, USA
Pamela Sklar - Institute for Molecular Medicine Finland, University of Helsinki, Helsinki, Finland
Aarno Palotie - Program in Medical and Population Genetics and Genetic Analysis Platform, Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA
Aarno Palotie - Department of Medical Epidemiology and Biostatistics, Karolinska Institutet, Stockholm, Sweden
Patrick F Sullivan - Departments of Genetics and Psychiatry, University of North Carolina, Chapel Hill, North Carolina, USA
Patrick F Sullivan
Authors
- Tarjinder Singh
You can also search for this author inPubMed Google Scholar - James T R Walters
You can also search for this author inPubMed Google Scholar - Mandy Johnstone
You can also search for this author inPubMed Google Scholar - David Curtis
You can also search for this author inPubMed Google Scholar - Jaana Suvisaari
You can also search for this author inPubMed Google Scholar - Minna Torniainen
You can also search for this author inPubMed Google Scholar - Elliott Rees
You can also search for this author inPubMed Google Scholar - Conrad Iyegbe
You can also search for this author inPubMed Google Scholar - Douglas Blackwood
You can also search for this author inPubMed Google Scholar - Andrew M McIntosh
You can also search for this author inPubMed Google Scholar - Georg Kirov
You can also search for this author inPubMed Google Scholar - Daniel Geschwind
You can also search for this author inPubMed Google Scholar - Robin M Murray
You can also search for this author inPubMed Google Scholar - Marta Di Forti
You can also search for this author inPubMed Google Scholar - Elvira Bramon
You can also search for this author inPubMed Google Scholar - Michael Gandal
You can also search for this author inPubMed Google Scholar - Christina M Hultman
You can also search for this author inPubMed Google Scholar - Pamela Sklar
You can also search for this author inPubMed Google Scholar - Aarno Palotie
You can also search for this author inPubMed Google Scholar - Patrick F Sullivan
You can also search for this author inPubMed Google Scholar - Michael C O'Donovan
You can also search for this author inPubMed Google Scholar - Michael J Owen
You can also search for this author inPubMed Google Scholar - Jeffrey C Barrett
You can also search for this author inPubMed Google Scholar
Consortia
INTERVAL Study
UK10K Consortium
Contributions
T.S. and J.C.B. conceived and designed the experiments. T.S. performed the statistical analysis. T.S., J.T.R.W., M.J., D.C., J.S., M.T., E.R., and P.F.S. analyzed the data. T.S., J.T.R.W., M.J., J.S., M.T., E.R., C.I., D.B., A.M.M., G.K., D.G., R.M.M., M.D.F., E.B., M.G., C.M.H., P.S., A.P., M.C.O'D., M.J.O., and J.C.B. contributed reagents, materials, or analysis tools. T.S., D.C., M.J.O., and J.C.B. wrote the manuscript.
Corresponding authors
Correspondence toTarjinder Singh or Jeffrey C Barrett.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Integrated supplementary information
Supplementary Figure 1 The use of frequency and size cutoffs in CNV gene set enrichment tests to reduce genomic inflation.
Quantile–quantile plots were generated based on P values from CNV enrichment tests of random gene sets, using different MAF cutoffs (<0.1%, 1%) and CNV size cutoffs (removing the top 5% and 10% of CNVs overlapping the most genes). Each dot represents a different gene set. The 95% CI assuming uniformly distributed P values is displayed as the gray shaded area. The genomic inflation factor (λ) is provided for each distribution. Inflation followed the reasonable null distribution when more stringent MAF thresholds and size cutoffs were applied (see MAF < 0.1% and removing the 10% of CNVs overlapping the most genes).
Supplementary Figure 2 Quantile–quantile plots of P values from enrichment tests of 1,766 gene sets.
Top left, case–control SNVs from whole-exome sequence data. Top right, de novo mutations from 1,077 trios. Bottom left, case–control CNVs. Bottom right, meta-analysis P values from Fisher’s method (dark blue). Tailored enrichment tests were applied to each variant type (Online Methods). Each dot represents a different gene set. The 95% CI assuming uniformly distributed P values is displayed as the gray shaded area. The genomic inflation factor (λ) is provided for each distribution. General inflation of P values from tests of disruptive variants (loss-of-function and CNVs) was observed, but no inflation was observed for tests of synonymous variants. Damaging missense, missense variants with CADD Phred > 15.
Supplementary Figure 3 Quantile–quantile plots of P values from enrichment tests of random gene sets.
Top left, case–control SNVs from whole-exome sequence data. Top right, de novo mutations from 1,077 trios. Bottom left, case–control CNVs. Bottom right, meta-analysis P values from Fisher’s method (dark blue). Genes were randomly sampled from the genome to create gene sets with the same size distribution as the 1,766 tested gene sets. Each dot represents a different gene set. The 95% CI assuming uniformly distributed P values is displayed as the gray shaded area. Tailored enrichment tests were applied to each variant type (Online Methods). The genomic inflation factor (λ) is provided for each distribution. No inflation of test statistics was observed in the meta-analysis P values. Damaging missense, missense variants with CADD Phred > 15.
Supplementary Figure 4 Enrichment of de novo mutations in genes with near-complete depletion of truncating variants across schizophrenia and neurodevelopmental disorders.
In schizophrenia, ASD, and severe neurodevelopmental disorders, de novo mutations were enriched in a subset of genes intolerant of loss-of-function variants, with no excess of polygenic burden in the remaining genes. To generate 95% CIs and P values, the rates of de novo mutations in affected trios (1,077 schizophrenia trios, 4,038 trios with ASD, and 1,133 trios with severe neurodevelopmental disorders) were compared against the rate in unaffected control trios (2,038 trios) using Poisson exact tests. Plotted P values are from the Poisson test of loss-of-function mutations. Damaging missense, missense variants with CADD Phred > 15.
Supplementary Figure 5 Enrichment of damaging rare variants in genes ordered and grouped by the degree of loss-of-function intolerance in schizophrenia, ASD, and severe neurodevelopmental disorders.
(a) Schizophrenia cases compared to controls for rare SNVs and indels. (b) Rates of de novo mutation in schizophrenia, ASD, and severe neurodevelopmental disorder probands as compared to control probands. Genes are ordered by their degree of loss-of-function intolerance (pLI score) and grouped into six categories: the 10% with the highest pLI score, the top 10–20% as ranked by pLI score, 20–40% as ranked by pLI score, and so on. Calculation of the 95% CIs and P values for the trio data followed the same method as in Supplementary Figure 4. A significant enrichment of rare, damaging variants was only observed in the 20% of genes with the highest pLI score, while no signal was observed in the remaining genes. Error bars are 95% CIs of the estimates. Damaging missense, missense variants with CADD Phred > 15.
Supplementary Figure 6 Non-random sampling of genes in the 1,766 tested gene sets.
(a) Genes are ranked and plotted based on the number of gene sets to which they belong. The top 1,000 genes were over-represented in gene sets from public databases, and genes outside the top 5,000 genes were under-represented. (b) Distribution of overlap coefficients with the set of loss-of-function-intolerant genes. The overlap coefficients between each of the 1,766 discovery gene sets and the set of loss-of-function-intolerant genes were calculated. The overlap coefficients between randomly sampled gene sets and loss-of-function-intolerant genes were similarly computed. These values are displayed as two density plots. The overlap coefficient is a similarity measure defined as 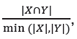 where X and Y are sets of genes.
where X and Y are sets of genes.
Supplementary Figure 7 Heat map of overlap coefficients calculated between the 35 significant gene sets (FDR < 5%).
The overlap coefficients of the 35 gene sets enriched for rare coding variants conferring risk for schizophrenia were computed and are clustered and displayed as a heat map. The overlap coefficient is a similarity measure defined as  where X and Y are sets of genes. The overlap coefficients between each gene set and the set of loss-of-function-intolerant genes are displayed as rounded values. See Supplementary Table 2 and the Supplementary Note for more information on each gene set.
where X and Y are sets of genes. The overlap coefficients between each gene set and the set of loss-of-function-intolerant genes are displayed as rounded values. See Supplementary Table 2 and the Supplementary Note for more information on each gene set.
Supplementary Figure 8 Summary of cognition and educational attainment data available for the schizophrenia whole-exome data set.
(a) Individuals diagnosed with schizophrenia (cases). (b) Individuals without a diagnosis of schizophrenia (controls). Information on population is also provided (UK, Finland, and Sweden).
Supplementary Figure 9 Enrichment of rare loss-of-function variants in loss-of-function-intolerant genes after excluding known developmental disorder–associated genes in schizophrenia cases stratified by information on cognitive function as compared to controls.
The P values shown were calculated using the variant threshold method comparing the burden of loss-of-function variants between the corresponding cases and controls. Error bars represent the 95% CIs of the point estimates. Damaging missense, missense variants with CADD Phred > 15.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Note. (PDF 2042 kb)
Supplementary Table 1
Full results from enrichment analyses of 1,766 gene sets. (XLSX 111 kb)
Supplementary Table 2
Gene sets enriched for rare coding variants conferring risk for schizophrenia at FDR < 5%. (XLSX 14 kb)
Supplementary Table 3
Results from enrichment analyses of FDR < 5% gene sets, conditional on brain-expressed and ExAC loss-of-function-intolerant genes. (XLSX 13 kb)
Supplementary Table 4
Results from enrichment analyses of rare loss-of-function variants in loss-of-function-intolerant genes and developmental disorder–associated genes comparing schizophrenia cases stratified by information on cognitive function and matched controls. (XLSX 10 kb)
Rights and permissions
About this article
Cite this article
Singh, T., Walters, J., Johnstone, M. et al. The contribution of rare variants to risk of schizophrenia in individuals with and without intellectual disability.Nat Genet 49, 1167–1173 (2017). https://doi.org/10.1038/ng.3903
- Received: 29 June 2016
- Accepted: 01 June 2017
- Published: 26 June 2017
- Issue Date: 01 August 2017
- DOI: https://doi.org/10.1038/ng.3903