Isolation and functional assessment of mouse skeletal stem cell lineage (original) (raw)
- Protocol
- Published: 10 May 2018
- Matthew P Murphy1,2 na1,
- Owen Marecic1,2 na1,
- Michael Lopez1,2 na1,
- Rachel E Brewer1,2,
- Lauren S Koepke1,2,
- Anoop Manjunath1,2,
- Ryan C Ransom1,2,
- Ankit Salhotra1,2,
- Irving L Weissman ORCID: orcid.org/0000-0002-9077-74671,3,4,
- Michael T Longaker1,2 &
- …
- Charles K F Chan1,2
Nature Protocols volume 13, pages 1294–1309 (2018)Cite this article
- 5542 Accesses
- 59 Citations
- 23 Altmetric
- Metrics details
Subjects
Abstract
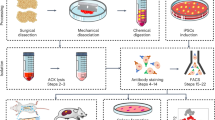
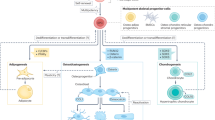
There are limited methods available to study skeletal stem, progenitor, and progeny cell activity in normal and diseased contexts. Most protocols for skeletal stem cell isolation are based on the extent to which cells adhere to plastic or whether they express a limited repertoire of surface markers. Here, we describe a flow cytometry–based approach that does not require in vitro selection and that uses eight surface markers to distinguish and isolate mouse skeletal stem cells (mSSCs); bone, cartilage, and stromal progenitors (mBCSPs); and five downstream differentiated subtypes, including chondroprogenitors, two types of osteoprogenitors, and two types of hematopoiesis-supportive stroma. We provide instructions for the optimal mechanical and chemical digestion of bone and bone marrow, as well as the subsequent flow-cytometry-activated cell sorting (FACS) gating schemes required to maximally yield viable skeletal-lineage cells. We also describe a methodology for renal subcapsular transplantation and in vitro colony-formation assays on the isolated mSSCs. The isolation of mSSCs can be completed in 9 h, with at least 1 h more required for transplantation. Experience with flow cytometry and mouse surgical procedures is recommended before attempting the protocol. Our system has wide applications and has already been used to study skeletal response to fracture, diabetes, and osteoarthritis, as well as hematopoietic stem cell–niche interactions in the bone marrow.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Similar content being viewed by others
Insights into skeletal stem cells
Article Open access 19 October 2022
References
- Reya, T., Morrison, S., Clarke, M. & Weissman, I. Stem cells, cancer, and cancer stem cells. Nature 414, 105–111 (2001).
Article CAS PubMed Google Scholar - Fuchs, E. & Segre, J. Stem cells: a new lease on life. Cell 100, 143–155 (2000).
Article CAS PubMed Google Scholar - Zhu, H. et al. A protocol for isolation and culture of mesenchymal stem cells from mouse compact bone. Nat. Protoc. 5, 550–560 (2010).
Article CAS PubMed Google Scholar - Soleimani, M. & Nadri, S. A protocol for isolation and culture of mesenchymal stem cells from mouse bone marrow. Nat. Protoc. 4, 102–106 (2009).
Article CAS PubMed Google Scholar - Caplan, A. Mesenchymal stem cells. J. Orthop. Res. 9, 641–650 (1991).
Article CAS PubMed Google Scholar - Pittenger, M. Multilineage potential of adult human mesenchymal stem cells. Science 284, 143–147 (1999).
Article CAS PubMed Google Scholar - Sacchetti, B. et al. Self-renewing osteoprogenitors in bone marrow sinusoids can organize a hematopoietic microenvironment. Cell 131, 324–336 (2007).
Article CAS PubMed Google Scholar - Zhou, B., Yue, R., Murphy, M., Peyer, J. & Morrison, S. Leptin-receptor-expressing mesenchymal stromal cells represent the main source of bone formed by adult bone marrow. Cell Stem Cell 15, 154–168 (2014).
Article CAS PubMed PubMed Central Google Scholar - Méndez-Ferrer, S. et al. Mesenchymal and haematopoietic stem cells form a unique bone marrow niche. Nature 466, 829–834 (2010).
Article PubMed PubMed Central Google Scholar - Pinho, S. et al. PDGFRα and CD51 mark human Nestin+ sphere-forming mesenchymal stem cells capable of hematopoietic progenitor cell expansion. J. Exp. Med. 210, 1351–1367 (2013).
Article CAS PubMed PubMed Central Google Scholar - Park, D. et al. Endogenous bone marrow MSCs are dynamic, fate-restricted participants in bone maintenance and regeneration. Cell Stem Cell 10, 259–272 (2012).
Article CAS PubMed PubMed Central Google Scholar - Phinney, D. Functional heterogeneity of mesenchymal stem cells: implications for cell therapy. J. Cell. Biochem. 113, 2806–2812 (2012).
Article CAS PubMed Google Scholar - Chan, C. et al. Identification and specification of the mouse skeletal stem cell. Cell 160, 285–298 (2015).
Article CAS PubMed PubMed Central Google Scholar - Chan, C. et al. Endochondral ossification is required for haematopoietic stem-cell niche formation. Nature 457, 490–494 (2008).
Article PubMed PubMed Central Google Scholar - Chan, C. et al. Clonal precursor of bone, cartilage, and hematopoietic niche stromal cells. Proc. Natl. Acad. Sci. USA 110, 12643–12648 (2013).
Article CAS PubMed PubMed Central Google Scholar - Ambrosi, T. et al. Adipocyte accumulation in the bone marrow during obesity and aging impairs stem cell-based hematopoietic and bone regeneration. Cell Stem Cell 20, 771–784.e6 (2017).
Article CAS PubMed PubMed Central Google Scholar - Morikawa, S. et al. Prospective identification, isolation, and systemic transplantation of multipotent mesenchymal stem cells in murine bone marrow. J. Exp. Med. 206, 2483–2496 (2009).
Article CAS PubMed PubMed Central Google Scholar - Worthley, D. et al. Gremlin 1 identifies a skeletal stem cell with bone, cartilage, and reticular stromal potential. Cell 160, 269–284 (2015).
Article CAS PubMed PubMed Central Google Scholar - Szade, K. et al. Where hematopoietic stem cells live: the bone marrow niche. Antioxid. Redox Signal. http://dx.doi.org/10.1089/ars.2017.7419 (2018).
- Reinisch, A. et al. A humanized bone marrow ossicle xenotransplantation model enables improved engraftment of healthy and leukemic human hematopoietic cells. Nat. Med. 22, 812–821 (2016).
Article CAS PubMed PubMed Central Google Scholar - Reinisch, A., Hernandez, D., Schallmoser, K. & Majeti, R. Generation and use of a humanized bone-marrow-ossicle niche for hematopoietic xenotransplantation into mice. Nat. Protoc. 12, 2169–2188 (2017).
Article CAS PubMed PubMed Central Google Scholar - Kunisaki, Y. et al. Arteriolar niches maintain haematopoietic stem cell quiescence. Nature 502, 637–643 (2013).
Article CAS PubMed PubMed Central Google Scholar - Morrison, S. & Scadden, D. The bone marrow niche for haematopoietic stem cells. Nature 505, 327–334 (2014).
Article CAS PubMed PubMed Central Google Scholar - Calvi, L. et al. Osteoblastic cells regulate the haematopoietic stem cell niche. Nature 425, 841–846 (2003).
Article CAS PubMed Google Scholar - Sugiyama, T., Kohara, H., Noda, M. & Nagasawa, T. Maintenance of the hematopoietic stem cell pool by CXCL12-CXCR4 chemokine signaling in bone marrow stromal cell niches. Immunity 25, 977–988 (2006).
Article CAS PubMed Google Scholar - Omatsu, Y. et al. The essential functions of adipo-osteogenic progenitors as the hematopoietic stem and progenitor cell niche. Immunity 33, 387–399 (2010).
Article CAS PubMed Google Scholar - Shiozawa, Y. et al. Human prostate cancer metastases target the hematopoietic stem cell niche to establish footholds in mouse bone marrow. J. Clin. Invest. 121, 1298–1312 (2011).
Article CAS PubMed PubMed Central Google Scholar - Cogle, C. et al. Bone marrow niche in the myelodysplastic syndromes. Leuk. Res. 39, 1020–1027 (2015).
Article PubMed Google Scholar - Schepers, K., Campbell, T. & Passegué, E. Normal and leukemic stem cell niches: insights and therapeutic opportunities. Cell Stem Cell 16, 254–267 (2015).
Article CAS PubMed PubMed Central Google Scholar - Raaijmakers, M. et al. Bone progenitor dysfunction induces myelodysplasia and secondary leukaemia. Nature 464, 852–857 (2010).
Article CAS PubMed PubMed Central Google Scholar - Lee, N. et al. Endocrine regulation of energy metabolism by the skeleton. Cell 130, 456–469 (2007).
Article CAS PubMed PubMed Central Google Scholar - Marecic, O. et al. Identification and characterization of an injury-induced skeletal progenitor. Proc. Natl. Acad. Sci. USA 112, 9920–9925 (2015).
Article CAS PubMed PubMed Central Google Scholar - Tevlin, R. et al. Pharmacological rescue of diabetic skeletal stem cell niches. Sci. Transl. Med. 9, eaag2809 (2017).
Article PubMed PubMed Central Google Scholar - Ueno, H. & Weissman, I. Clonal analysis of mouse development reveals a polyclonal origin for yolk sac blood islands. Dev. Cell 11, 519–533 (2006).
Article CAS PubMed Google Scholar - Lee, R., Hannig, J., Matthews, K., Myerov, A. & Chen, C. Pharmaceutical therapies for sealing of permeabilized cell membranes in electrical injuries. Ann. N.Y. Acad. Sci. 888, 266–273 (1999).
Article CAS PubMed Google Scholar - Garvey, W., Fathi, A., Bigelow, F., Carpenter, B. & Jimenez, C. Improved Movat pentachrome stain. Stain Technol. 61, 60–62 (1986).
Article CAS PubMed Google Scholar
Acknowledgements
We thank A. McCarty and C. Wang for mouse-colony management; P. Pereira, T. Storm, E. Seo, and T. Naik for laboratory management; and P. Lovelace, J. Ho, and S. Weber for FACS management. This study was supported by the National Institutes of Health (NIH; grants R56 DE025597, R01 DE021683, R21 DE024230, R01 DE019434, RC2 DE020771, U01 HL099776, and R21 DE019274 to M.T.L.; grants U01HL099999, 5 R01 CA86065, and 5 R01 L058770 to I.L.W.); a Siebel Fellowship from the Thomas and Stacey Siebel Foundation, a Prostate Cancer Foundation Young Investigator Award, and a National Institute on Aging Research Career Development Award (grant 1K99AG049958-01A1) to C.K.F.C.; the California Institute for Regenerative Medicine (CIRM; grant TR1-01249), the Oak Foundation, the Hagey Laboratory for Pediatric Regenerative Medicine, and the Gunn/Olivier Research Fund to M.T.L.; a Howard Hughes Medical Institute Medical Student Research Fellowship to G.S.G.; and The Plastic Surgery Research Foundation National Endowment for Plastic Surgery to M.P.M.
Author information
Author notes
- Gunsagar S Gulati, Matthew P Murphy, Owen Marecic and Michael Lopez: These authors contributed equally to this work.
Authors and Affiliations
- Stanford Stem Cell Biology and Regenerative Medicine Institute, Stanford University, Stanford, CA, USA
Gunsagar S Gulati, Matthew P Murphy, Owen Marecic, Michael Lopez, Rachel E Brewer, Lauren S Koepke, Anoop Manjunath, Ryan C Ransom, Ankit Salhotra, Irving L Weissman, Michael T Longaker & Charles K F Chan - Plastic and Reconstructive Surgery Division, Department of Surgery, Hagey Laboratory for Pediatric Regenerative Medicine, Stanford University, Stanford, CA, USA
Gunsagar S Gulati, Matthew P Murphy, Owen Marecic, Michael Lopez, Rachel E Brewer, Lauren S Koepke, Anoop Manjunath, Ryan C Ransom, Ankit Salhotra, Michael T Longaker & Charles K F Chan - Department of Pathology, Stanford University School of Medicine, Stanford, CA, USA
Irving L Weissman - Department of Developmental Biology, Stanford University School of Medicine, Stanford, CA, USA
Irving L Weissman
Authors
- Gunsagar S Gulati
You can also search for this author inPubMed Google Scholar - Matthew P Murphy
You can also search for this author inPubMed Google Scholar - Owen Marecic
You can also search for this author inPubMed Google Scholar - Michael Lopez
You can also search for this author inPubMed Google Scholar - Rachel E Brewer
You can also search for this author inPubMed Google Scholar - Lauren S Koepke
You can also search for this author inPubMed Google Scholar - Anoop Manjunath
You can also search for this author inPubMed Google Scholar - Ryan C Ransom
You can also search for this author inPubMed Google Scholar - Ankit Salhotra
You can also search for this author inPubMed Google Scholar - Irving L Weissman
You can also search for this author inPubMed Google Scholar - Michael T Longaker
You can also search for this author inPubMed Google Scholar - Charles K F Chan
You can also search for this author inPubMed Google Scholar
Contributions
C.K.F.C., G.S.G., M.T.L., and I.L.W. conceived the isolation strategy and functional assays. M.T.L. and I.L.W. supervised the project. G.S.G., M.P.M., O.M., and M.L. developed the protocol, performed the experiments, and analyzed the data. G.S.G. wrote the manuscript. R.E.B., L.S.K., R.C.R., A.S., and A.M. assisted with flow cytometry, in vitro assays, and manuscript preparation.
Corresponding author
Correspondence toCharles K F Chan.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Effect of successive chemical digests on total cell yield.
Total cell yield from crushed bones increases with each successive digest but saturates after three digests [Steps 11-15]. Briefly, crushed 8-week mouse skeletal tissue was separated from whole bone marrow and digested in digest buffer containing 3000 U ml−1 type II collagenase. Total cell number before and after each successive digest was counted by hemocytometer and mean and standard deviation for n = 4 was calculated. All animal experiments in this figure are in accordance with the Stanford's Administrative Panel on Laboratory Animal Care (APLAC) and received approval from the Institutional Review Board (IRB).
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1. (PDF 199 kb)
Supplementary Data
Estimated total number of cells obtained from individual 8-week-old C57BL/6 mice. The data in this file were used to obtain the averages ± standard deviation stated in Table 1. (XLSX 9 kb)
Rights and permissions
About this article
Cite this article
Gulati, G., Murphy, M., Marecic, O. et al. Isolation and functional assessment of mouse skeletal stem cell lineage.Nat Protoc 13, 1294–1309 (2018). https://doi.org/10.1038/nprot.2018.041
- Published: 10 May 2018
- Issue Date: June 2018
- DOI: https://doi.org/10.1038/nprot.2018.041