Formaldehyde catabolism is essential in cells deficient for the Fanconi anemia DNA-repair pathway (original) (raw)
- Brief Communication
- Published: 13 November 2011
Nature Structural & Molecular Biology volume 18, pages 1432–1434 (2011)Cite this article
- 5120 Accesses
- 184 Citations
- 7 Altmetric
- Metrics details
Subjects
Abstract
Metabolism is predicted to generate formaldehyde, a toxic, simple, reactive aldehyde that can damage DNA. Here we report a synthetic lethal interaction in avian cells between ADH5, encoding the main formaldehyde-detoxifying enzyme, and the Fanconi anemia (FA) DNA-repair pathway. These results define a fundamental role for the combined action of formaldehyde catabolism and DNA cross-link repair in vertebrate cell survival.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: The FA DNA-repair pathway ensures genome stability after exogenous formaldehyde exposure.

The alternative text for this image may have been generated using AI.
Figure 2: FANCD2 and ADH5 are synthetically lethal in DT40 cells.
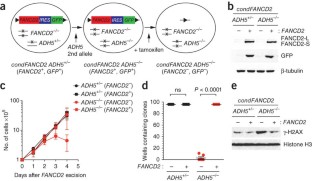
The alternative text for this image may have been generated using AI.
Figure 3: The FA core complex gene FANCL and ADH5 are synthetically lethal in DT40 cells.

The alternative text for this image may have been generated using AI.
Similar content being viewed by others
References
- Muller, L.U. & Williams, D.A. Mutat. Res. 668, 141–149 (2009).
Article PubMed PubMed Central Google Scholar - Crossan, G.P. et al. Nat. Genet. 43, 147–152 (2011).
Article CAS PubMed PubMed Central Google Scholar - Stoepker, C. et al. Nat. Genet. 43, 138–141 (2011).
Article CAS PubMed Google Scholar - Kim, Y. et al. Nat. Genet. 43, 142–146 (2011).
Article CAS PubMed PubMed Central Google Scholar - Joenje, H. & Patel, K.J. Nat. Rev. Genet. 2, 446–457 (2001).
Article CAS PubMed Google Scholar - Patel, K.J. & Joenje, H. DNA Repair (Amst.) 6, 885–890 (2007).
Article CAS Google Scholar - de Winter, J.P. & Joenje, H. Mutat. Res. 668, 11–19 (2009).
Article CAS PubMed Google Scholar - Rosado, I.V., Niedzwiedz, W., Alpi, A.F. & Patel, K.J. Nucleic Acids Res. 37, 4360–4370 (2009).
Article CAS PubMed PubMed Central Google Scholar - Langevin, F., Crossan, G.P., Rosado, I.V., Arends, M.J. & Patel, K.J. Nature 475, 53–58 (2011).
Article CAS PubMed Google Scholar - Mosammaparast, N. & Shi, Y. Annu. Rev. Biochem. 79, 155–179 (2010).
Article CAS PubMed Google Scholar - Ridpath, J.R. et al. Cancer Res. 67, 11117–11122 (2007).
Article CAS PubMed Google Scholar - Noda, T. et al. Biochem. Biophys. Res. Commun. 404, 206–210 (2011).
Article CAS PubMed Google Scholar - Nomura, Y., Adachi, N. & Koyama, H. Genes Cells 12, 1111–1122 (2007).
Article CAS PubMed Google Scholar - Trewick, S.C., Henshaw, T.F., Hausinger, R.P., Lindahl, T. & Sedgwick, B. Nature 419, 174–178 (2002).
Article CAS PubMed Google Scholar - Falnes, P.O., Johansen, R.F. & Seeberg, E. Nature 419, 178–182 (2002).
Article CAS PubMed Google Scholar - Iborra, F.J. et al. J. Histochem. Cytochem. 40, 1865–1878 (1992).
Article CAS PubMed Google Scholar - Mosedale, G. et al. Nat. Struct. Mol. Biol. 12, 763–771 (2005).
Article CAS PubMed Google Scholar - Zhang, X.Y. et al. PLoS Genet. 5, e1000645 (2009).
Article PubMed PubMed Central Google Scholar
Acknowledgements
We thank N. Adachi and H. Koyama (Kihara Institute for Biological Research, Japan) for kindly providing NALM-6 human cell lines. We are grateful to J. Sutherland and K. Lang (MRC Laboratory of Molecular Biology) for chemical insight and to M. Daly and F. Zhang (MRC Laboratory of Molecular Biology) for invaluable help with flow cytometry. I.V.R. and F.L. are funded by the Fanconi Anaemia Research Fund and Children's Leukaemia Trust, respectively.
Author information
Authors and Affiliations
- Medical Research Council Laboratory of Molecular Biology, Cambridge, UK
Ivan V Rosado, Frédéric Langevin, Gerry P Crossan & Ketan J Patel - Department of Late Effect Studies, Laboratory of DNA Damage Signaling, Radiation Biology Center, Kyoto University, Kyoto, Japan
Minoru Takata - Department of Medicine, University of Cambridge, Addenbrooke's Hospital, Cambridge, UK
Ketan J Patel
Authors
- Ivan V Rosado
- Frédéric Langevin
- Gerry P Crossan
- Minoru Takata
- Ketan J Patel
Contributions
I.V.R. and K.J.P. designed the study and the experiments, and wrote the paper. I.V.R. performed the majority of the experiments presented. F.L. contributed to DT40 clonogenic assays and assisted in the generation of _ADH5_-deficient cell lines. G.P.C. helped with analysis of chromosome breaks. M.T. generated and provided the FANCD2 inducible cell line.
Corresponding author
Correspondence toKetan J Patel.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Rosado, I., Langevin, F., Crossan, G. et al. Formaldehyde catabolism is essential in cells deficient for the Fanconi anemia DNA-repair pathway.Nat Struct Mol Biol 18, 1432–1434 (2011). https://doi.org/10.1038/nsmb.2173
- Received: 29 June 2011
- Accepted: 16 September 2011
- Published: 13 November 2011
- Issue date: December 2011
- DOI: https://doi.org/10.1038/nsmb.2173