Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs (original) (raw)
- Letter
- Published: 30 January 2005
- Nelson C. Lau2,
- Philip Garrett-Engele1,
- Andrew Grimson2,
- Janell M. Schelter1,
- John Castle1,
- David P. Bartel2,
- Peter S. Linsley1 &
- …
- Jason M. Johnson1
Nature volume 433, pages 769–773 (2005)Cite this article
- 24k Accesses
- 4306 Citations
- 21 Altmetric
- Metrics details
Abstract
MicroRNAs (miRNAs) are a class of noncoding RNAs that post-transcriptionally regulate gene expression in plants and animals1,2. To investigate the influence of miRNAs on transcript levels, we transfected miRNAs into human cells and used microarrays to examine changes in the messenger RNA profile. Here we show that delivering miR-124 causes the expression profile to shift towards that of brain, the organ in which miR-124 is preferentially expressed, whereas delivering miR-1 shifts the profile towards that of muscle, where miR-1 is preferentially expressed. In each case, about 100 messages were downregulated after 12 h. The 3′ untranslated regions of these messages had a significant propensity to pair to the 5′ region of the miRNA, as expected if many of these messages are the direct targets of the miRNAs3. Our results suggest that metazoan miRNAs can reduce the levels of many of their target transcripts, not just the amount of protein deriving from these transcripts. Moreover, miR-1 and miR-124, and presumably other tissue-specific miRNAs, seem to downregulate a far greater number of targets than previously appreciated, thereby helping to define tissue-specific gene expression in humans.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Tissue-specific gene expression rankings for downregulated genes.

The alternative text for this image may have been generated using AI.
Figure 2: Over-represented motifs in the 3′ UTRs of downregulated genes.
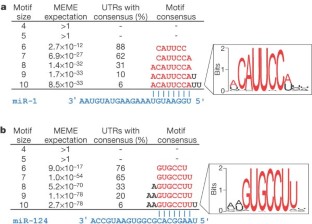
The alternative text for this image may have been generated using AI.
Figure 3: Microarray analysis of the effects of miRNA mutations.
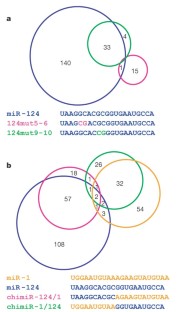
The alternative text for this image may have been generated using AI.
Figure 4: MicroRNA-directed repression of renilla luciferase reporter genes bearing 3′ UTR segments from predicted target genes.
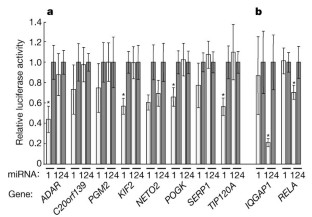
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
References
- Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116, 281–297 (2004)
Article CAS Google Scholar - Carrington, J. C. & Ambros, V. Role of microRNAs in plant and animal development. Science 301, 336–338 (2003)
Article ADS CAS Google Scholar - Lewis, B. P., Shih, I. H., Jones-Rhoades, M. W., Bartel, D. P. & Burge, C. B. Prediction of mammalian microRNA targets. Cell 115, 787–798 (2003)
Article CAS Google Scholar - Palatnik, J. F. et al. Control of leaf morphogenesis by microRNAs. Nature 425, 257–263 (2003)
Article ADS CAS Google Scholar - Jackson, A. L. et al. Expression profiling reveals off-target gene regulation by RNAi. Nature Biotechnol. 21, 635–637 (2003)
Article CAS Google Scholar - Hutvagner, G. & Zamore, P. D. A microRNA in a multiple-turnover RNAi enzyme complex. Science 297, 2056–2060 (2002)
Article ADS CAS Google Scholar - Zeng, Y., Yi, R. & Cullen, B. R. MicroRNAs and small interfering RNAs can inhibit mRNA expression by similar mechanisms. Proc. Natl Acad. Sci. USA 100, 9779–9784 (2003)
Article ADS CAS Google Scholar - Doench, J. G., Petersen, C. P. & Sharp, P. A. siRNAs can function as miRNAs. Genes Dev. 17, 438–442 (2003)
Article CAS Google Scholar - Yekta, S., Shih, I. H. & Bartel, D. P. MicroRNA-directed cleavage of HOXB8 mRNA. Science 304, 594–596 (2004)
Article ADS CAS Google Scholar - Lagos-Quintana, M. et al. Identification of tissue-specific microRNAs from mouse. Curr. Biol. 12, 735–739 (2002)
Article CAS Google Scholar - Sempere, L. F. et al. Expression profiling of mammalian microRNAs uncovers a subset of brain-expressed microRNAs with possible roles in murine and human neuronal differentiation. Genome Biol. 5, R13 (2004)
Article Google Scholar - Pruitt, K. D. & Maglott, D. R. RefSeq and LocusLink: NCBI gene-centered resources. Nucleic Acids Res. 29, 137–140 (2001)
Article CAS Google Scholar - Johnson, J. M. et al. Genome-wide survey of human alternative pre-mRNA splicing with exon junction microarrays. Science 302, 2141–2144 (2003)
Article ADS CAS Google Scholar - Bailey, T. L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 2, 28–36 (1994)
CAS PubMed Google Scholar - Lim, L. P. et al. The microRNAs of Caenorhabditis elegans . Genes Dev. 17, 991–1008 (2003)
Article CAS Google Scholar - Wightman, B., Ha, I. & Ruvkun, G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans . Cell 75, 855–862 (1993)
Article CAS Google Scholar - Lai, E. C. Micro RNAs are complementary to 3′ UTR sequence motifs that mediate negative post-transcriptional regulation. Nature Genet. 30, 363–364 (2002)
Article CAS Google Scholar - Doench, J. G. & Sharp, P. A. Specificity of microRNA target selection in translational repression. Genes Dev. 18, 504–511 (2004)
Article CAS Google Scholar - John, B. et al. Human MicroRNA targets. PLoS Biol. 2, e363 (2004)
Article Google Scholar - Rhoades, M. W. et al. Prediction of plant microRNA targets. Cell 110, 513–520 (2002)
Article CAS Google Scholar - Haley, B. & Zamore, P. D. Kinetic analysis of the RNAi enzyme complex. Nature Struct. Mol. Biol. 11, 599–606 (2004)
Article CAS Google Scholar - Martinez, J. & Tuschl, T. RISC is a 5′ phosphomonoester-producing RNA endonuclease. Genes Dev. 18, 975–980 (2004)
Article CAS Google Scholar - Janz, R. & Sudhof, T. C. Cellugyrin, a novel ubiquitous form of synaptogyrin that is phosphorylated by pp60c-src. J. Biol. Chem. 273, 2851–2857 (1998)
Article CAS Google Scholar - Vartiainen, M. K., Sarkkinen, E. M., Matilainen, T., Salminen, M. & Lappalainen, P. Mammals have two twinfilin isoforms whose subcellular localizations and tissue distributions are differentially regulated. J. Biol. Chem. 278, 34347–34355 (2003)
Article CAS Google Scholar - Bartel, D. P. & Chen, C. Z. Micromanagers of gene expression: the potentially widespread influence of metazoan microRNAs. Nature Rev. Genet. 5, 396–400 (2004)
Article CAS Google Scholar - Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nature Genet. 25, 25–29 (2000)
Article CAS Google Scholar - Schwarz, D. S. et al. Asymmetry in the assembly of the RNAi enzyme complex. Cell 115, 199–208 (2003)
Article CAS Google Scholar - Khvorova, A., Reynolds, A. & Jayasena, S. D. Functional siRNAs and miRNAs exhibit strand bias. Cell 115, 209–216 (2003)
Article CAS Google Scholar - Hughes, T. R. et al. Functional discovery via a compendium of expression profiles. Cell 102, 109–126 (2000)
Article CAS Google Scholar - Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994)
Article CAS Google Scholar
Acknowledgements
Thanks to S. Baskerville, M. Cleary and P. Sharp for comments on the manuscript, C. Armour, S. Bartz, J. Burchard, G. Cavet, D. Haynor, A. Jackson, M. Pellegrini, E. Schadt and Y. Wang for their assistance, the Rosetta Gene Expression Laboratory for microarray work, M. Jones-Rhoades for primer design, and W. Johnston for plasmid construction.
Author information
Authors and Affiliations
- Rosetta Inpharmatics (wholly owned subsidiary of Merck and Co.), 401 Terry Avenue N, Seattle, Washington, 98109, USA
Lee P. Lim, Philip Garrett-Engele, Janell M. Schelter, John Castle, Peter S. Linsley & Jason M. Johnson - Whitehead Institute for Biomedical Research and Department of Biology, Massachusetts Institute of Technology, 9 Cambridge Center, Cambridge, Massachusetts, 02142, USA
Nelson C. Lau, Andrew Grimson & David P. Bartel
Authors
- Lee P. Lim
- Nelson C. Lau
- Philip Garrett-Engele
- Andrew Grimson
- Janell M. Schelter
- John Castle
- David P. Bartel
- Peter S. Linsley
- Jason M. Johnson
Corresponding author
Correspondence toLee P. Lim.
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Lim, L., Lau, N., Garrett-Engele, P. et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs.Nature 433, 769–773 (2005). https://doi.org/10.1038/nature03315
- Received: 22 July 2004
- Accepted: 22 December 2004
- Published: 30 January 2005
- Issue date: 17 February 2005
- DOI: https://doi.org/10.1038/nature03315
This article is cited by
Editorial Summary
microRNAs edit the message
New work on microRNAs (miRNAs) sheds light on the nature of tissue-specific gene expression in humans. miRNAs act in plants and animals to regulate gene expression. Microarray analysis of messenger RNAs expressed in human cells containing miR-124, an miRNA expressed in the brain, or miR-1, found in muscle, shows that the miRNAs skew a cell's expression profile to resemble that of the miRNA's ‘home’ cells, regardless of source. This involves the downregulation of many target mRNA transcripts, as well as reduction of the amount of protein produced.