Programming cells by multiplex genome engineering and accelerated evolution (original) (raw)
- Letter
- Published: 26 July 2009
- Farren J. Isaacs1 na1,
- Peter A. Carr4,5,
- Zachary Z. Sun6,
- George Xu6,
- Craig R. Forest7 &
- …
- George M. Church1
Nature volume 460, pages 894–898 (2009)Cite this article
- 59k Accesses
- 1445 Citations
- 144 Altmetric
- Metrics details
Abstract
The breadth of genomic diversity found among organisms in nature allows populations to adapt to diverse environments1,2. However, genomic diversity is difficult to generate in the laboratory and new phenotypes do not easily arise on practical timescales3. Although in vitro and directed evolution methods4,5,6,7,8,9 have created genetic variants with usefully altered phenotypes, these methods are limited to laborious and serial manipulation of single genes and are not used for parallel and continuous directed evolution of gene networks or genomes. Here, we describe multiplex automated genome engineering (MAGE) for large-scale programming and evolution of cells. MAGE simultaneously targets many locations on the chromosome for modification in a single cell or across a population of cells, thus producing combinatorial genomic diversity. Because the process is cyclical and scalable, we constructed prototype devices that automate the MAGE technology to facilitate rapid and continuous generation of a diverse set of genetic changes (mismatches, insertions, deletions). We applied MAGE to optimize the 1-deoxy-d-xylulose-5-phosphate (DXP) biosynthesis pathway in Escherichia coli to overproduce the industrially important isoprenoid lycopene. Twenty-four genetic components in the DXP pathway were modified simultaneously using a complex pool of synthetic DNA, creating over 4.3 billion combinatorial genomic variants per day. We isolated variants with more than fivefold increase in lycopene production within 3 days, a significant improvement over existing metabolic engineering techniques. Our multiplex approach embraces engineering in the context of evolution by expediting the design and evolution of organisms with new and improved properties.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Multiplex automated genome engineering enables the rapid and continuous generation of sequence diversity at many targeted chromosomal locations across a large population of cells through the repeated introduction of synthetic DNA.
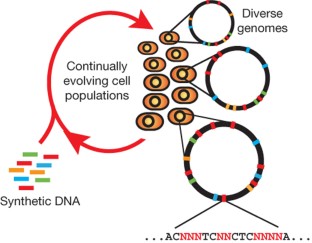
The alternative text for this image may have been generated using AI.
Figure 2: Characterization of allelic replacement efficiency as a function of the type and scale of genetic modifications.
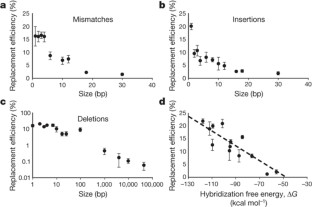
The alternative text for this image may have been generated using AI.
Figure 3: Sequence diversity generated across three separate cell populations as a function of the number of MAGE cycles.
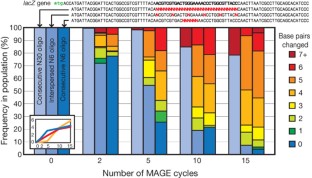
The alternative text for this image may have been generated using AI.
Figure 4: MAGE automation.
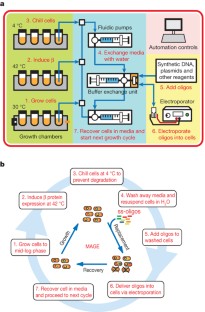
The alternative text for this image may have been generated using AI.
Figure 5: Optimization of the DXP biosynthesis pathway for lycopene production.
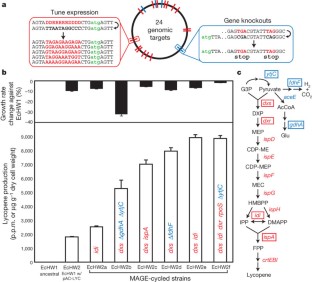
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
References
- Venter, J. C. et al. Environmental genome shotgun sequencing of the Sargasso Sea. Science 304, 66–74 (2004)
Article ADS CAS Google Scholar - Tringe, S. G. et al. Comparative metagenomics of microbial communities. Science 308, 554–557 (2005)
Article ADS CAS Google Scholar - Elena, S. F. & Lenski, R. E. Evolution experiments with microorganisms: the dynamics and genetic bases of adaptation. Nature Rev. Genet. 4, 457–469 (2003)
Article CAS Google Scholar - Ellington, A. D. & Szostak, J. W. In vitro selection of RNA molecules that bind specific ligands. Nature 346, 818–822 (1990)
Article ADS CAS Google Scholar - Crameri, A., Raillard, S.-A., Bermudez, E., Stemmer, W. P. & C DNA shuffling of a family of genes from diverse species accelerates directed evolution. Nature 391, 288–291 (1998)
Article ADS CAS Google Scholar - Joo, H., Lin, Z. & Arnold, F. H. Laboratory evolution of peroxide-mediated cytochrome P450 hydroxylation. Nature 399, 670–673 (1999)
Article ADS CAS Google Scholar - Zhang, Y. X. et al. Genome shuffling leads to rapid phenotypic improvement in bacteria. Nature 415, 644–646 (2002)
Article ADS CAS Google Scholar - Pfleger, B. F., Pitera, D. J., Smolke, C. D. & Keasling, J. D. Combinatorial engineering of intergenic regions in operons tunes expression of multiple genes. Nature Biotechnol. 24, 1027–1032 (2006)
Article CAS Google Scholar - Cadwell, R. C. & Joyce, G. F. Randomization of genes by PCR mutagenesis. PCR Methods Appl. 2, 28–33 (1992)
Article CAS Google Scholar - Shendure, J. et al. Accurate multiplex polony sequencing of an evolved bacterial genome. Science 309, 1728–1732 (2005)
Article ADS CAS Google Scholar - Zhang, Y., Buchholz, F., Muyrers, J. P. & Stewart, A. F. A new logic for DNA engineering using recombination in Escherichia coli . Nature Genet. 20, 123–128 (1998)
Article CAS Google Scholar - Costantino, N. & Court, D. L. Enhanced levels of λ Red-mediated recombinants in mismatch repair mutants. Proc. Natl Acad. Sci. USA 100, 15748–15753 (2003)
Article ADS CAS Google Scholar - Sharan, S. K., Thomason, L. C., Kuznetsov, S. G. & Court, D. L. Recombineering: a homologous recombination-based method of genetic engineering. Nature Protocols 4, 206–223 (2009)
Article CAS Google Scholar - Ellis, H. M., Yu, D., DiTizio, T. & Court, D. L. High efficiency mutagenesis, repair, and engineering of chromosomal DNA using single-stranded oligonucleotides. Proc. Natl Acad. Sci. USA 98, 6742–6746 (2001)
Article ADS CAS Google Scholar - Markham, N. R. & Zuker, M. DINAMelt web server for nucleic acid melting prediction. Nucleic Acids Res. 33, W577–W581 (2005)
Article CAS Google Scholar - Jin, Y. S. & Stephanopoulos, G. Multi-dimensional gene target search for improving lycopene biosynthesis in Escherichia coli . Metab. Eng. 9, 337–347 (2007)
Article CAS Google Scholar - Kang, M. J. et al. Identification of genes affecting lycopene accumulation in Escherichia coli using a shot-gun method. Biotechnol. Bioeng. 91, 636–642 (2005)
Article CAS Google Scholar - Chen, H., Bjerknes, M., Kumar, R. & Jay, E. Determination of the optimal aligned spacing between the Shine – Dalgarno sequence and the translation initiation codon of Escherichia coli mRNAs. Nucleic Acids Res. 22, 4953–4957 (1994)
Article CAS Google Scholar - Alper, H., Jin, Y. S., Moxley, J. F. & Stephanopoulos, G. Identifying gene targets for the metabolic engineering of lycopene biosynthesis in Escherichia coli . Metab. Eng. 7, 155–164 (2005)
Article CAS Google Scholar - Alper, H., Miyaoku, K. & Stephanopoulos, G. Construction of lycopene-overproducing E. coli strains by combining systematic and combinatorial gene knockout targets. Nature Biotechnol. 23, 612–616 (2005)
Article CAS Google Scholar - Farmer, W. R. & Liao, J. C. Precursor balancing for metabolic engineering of lycopene production in Escherichia coli . Biotechnol. Prog. 17, 57–61 (2001)
Article CAS Google Scholar - Kim, S. W. & Keasling, J. D. Metabolic engineering of the nonmevalonate isopentenyl diphosphate synthesis pathway in Escherichia coli enhances lycopene production. Biotechnol. Bioeng. 72, 408–415 (2001)
Article CAS Google Scholar - Yuan, L. Z., Rouviere, P. E., Larossa, R. A. & Suh, W. Chromosomal promoter replacement of the isoprenoid pathway for enhancing carotenoid production in E. coli . Metab. Eng. 8, 79–90 (2006)
Article CAS Google Scholar - Khosla, C. & Keasling, J. D. Metabolic engineering for drug discovery and development. Nature Rev. Drug Discov. 2, 1019–1025 (2003)
Article CAS Google Scholar - Cropp, T. A. & Schultz, P. G. An expanding genetic code. Trends Genet. 20, 625–630 (2004)
Article CAS Google Scholar - Gibson, D. G. et al. Complete chemical synthesis, assembly, and cloning of a Mycoplasma genitalium genome. Science 319, 1215–1220 (2008)
Article ADS CAS Google Scholar - Metzgar, D. et al. Acinetobacter sp. ADP1: an ideal model organism for genetic analysis and genome engineering. Nucleic Acids Res. 32, 5780–5790 (2004)
Article CAS Google Scholar - Nakayama, M. & Ohara, O. Improvement of recombination efficiency by mutation of Red proteins. Biotechniques 38, 917–924 (2005)
Article CAS Google Scholar - Datta, S., Costantino, N., Zhou, X. & Court, D. L. Identification and analysis of recombineering functions from Gram-negative and Gram-positive bacteria and their phages. Proc. Natl Acad. Sci. USA 105, 1626–1631 (2008)
Article ADS CAS Google Scholar - Tian, J. et al. Accurate multiplex gene synthesis from programmable DNA microchips. Nature 432, 1050–1054 (2004)
Article ADS CAS Google Scholar - Yu, D. et al. An efficient recombination system for chromosome engineering in Escherichia coli . Proc. Natl Acad. Sci. USA 97, 5978–5983 (2000)
Article ADS CAS Google Scholar - Cunningham, F. X., Sun, Z., Chamovitz, D., Hirschberg, J. & Gantt, E. Molecular structure and enzymatic function of lycopene cyclase from the cyanobacterium Synechococcus sp strain PCC7942. Plant Cell 6, 1107–1121 (1994)
Article CAS Google Scholar
Acknowledgements
We are grateful to J. Jacobson for his insights and advice throughout this work. We thank D. Court for his insights and sharing strain DY330, N. Reppas for advice and sharing strain EcNR2, F. X. Cunningham for sharing pAC-LYC, and B. H. Sterling for assistance in constructing the EcFI5 strain. We also thank M. Jewett, J. Aach, D. Bang, S. Kosuri and members of the Church laboratory for advice and discussions. We thank the NSF, DOE, DARPA, the Wyss Institute for Biologically Inspired Engineering and training fellowships from the NIH and NDSEG (H.H.W.) for supporting this research.
Author Contributions H.H.W., F.J.I. and G.M.C. conceived the study jointly with P.A.C.; H.H.W. and F.J.I. designed and performed experiments with assistance from P.A.C., Z.Z.S., G.X. and C.R.F.; H.H.W. and F.J.I. wrote the manuscript; G.M.C. supervised all aspects of the study.
Author information
Author notes
- Harris H. Wang and Farren J. Isaacs: These authors contributed equally to this work.
Authors and Affiliations
- Department of Genetics, Harvard Medical School, Boston, Massachusetts 02115, USA,
Harris H. Wang, Farren J. Isaacs & George M. Church - Program in Biophysics, Harvard University, Cambridge, Massachusetts 02138, USA ,
Harris H. Wang - Harvard-MIT Division of Health Sciences and Technology,, Program in Medical Engineering Medical Physics,
Harris H. Wang - The Center for Bits and Atoms,,
Peter A. Carr - Media Lab, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA ,
Peter A. Carr - Harvard College, Cambridge, Massachusetts 02138, USA ,
Zachary Z. Sun & George Xu - George W. Woodruff School of Mechanical Engineering, Georgia Institute of Technology, Atlanta, Georgia 30332, USA ,
Craig R. Forest
Authors
- Harris H. Wang
- Farren J. Isaacs
- Peter A. Carr
- Zachary Z. Sun
- George Xu
- Craig R. Forest
- George M. Church
Corresponding authors
Correspondence toHarris H. Wang or Farren J. Isaacs.
Ethics declarations
Competing interests
we wish to disclose that three authors (G.M.C., H.H.W, F.J.I.) have a pending patent application whose value may be affected by the publication of this paper. G.M.C. also discloses various associations with companies as outlined at http://arep.med.harvard.edu/gmc/tech.html.
Supplementary information
PowerPoint slides
Rights and permissions
About this article
Cite this article
Wang, H., Isaacs, F., Carr, P. et al. Programming cells by multiplex genome engineering and accelerated evolution.Nature 460, 894–898 (2009). https://doi.org/10.1038/nature08187
- Received: 06 March 2009
- Accepted: 29 May 2009
- Published: 26 July 2009
- Issue date: 13 August 2009
- DOI: https://doi.org/10.1038/nature08187
This article is cited by
Editorial Summary
Generating genomic diversity
Genomic diversity is difficult to generate in the laboratory in an efficient way. A new technique called MAGE (multiplex automated genome engineering), described here, simultaneously targets many locations on the chromosome for modification in a single cell or across a population of cells, thereby producing combinatorial genomic diversity. This is an automated and efficient approach that expedites the design and evolution of organisms with new and improved properties.