Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis (original) (raw)
- Letter
- Published: 15 April 2010
- Nilay Shah4,
- Kevin C. Wang1,
- Jeewon Kim2,
- Hugo M. Horlings6,
- David J. Wong1,
- Miao-Chih Tsai1,
- Tiffany Hung1,
- Pedram Argani5,
- John L. Rinn7,
- Yulei Wang8,
- Pius Brzoska8,
- Benjamin Kong8,
- Rui Li3,
- Robert B. West3,
- Marc J. van de Vijver6,
- Saraswati Sukumar4 &
- …
- Howard Y. Chang1
Nature volume 464, pages 1071–1076 (2010)Cite this article
- 44k Accesses
- 4793 Citations
- 178 Altmetric
- Metrics details
Subjects
Abstract
Large intervening non-coding RNAs (lincRNAs) are pervasively transcribed in the genome1,2,3 yet their potential involvement in human disease is not well understood4,5. Recent studies of dosage compensation, imprinting, and homeotic gene expression suggest that individual lincRNAs can function as the interface between DNA and specific chromatin remodelling activities6,7,8. Here we show that lincRNAs in the HOX loci become systematically dysregulated during breast cancer progression. The lincRNA termed HOTAIR is increased in expression in primary breast tumours and metastases, and HOTAIR expression level in primary tumours is a powerful predictor of eventual metastasis and death. Enforced expression of HOTAIR in epithelial cancer cells induced genome-wide re-targeting of Polycomb repressive complex 2 (PRC2) to an occupancy pattern more resembling embryonic fibroblasts, leading to altered histone H3 lysine 27 methylation, gene expression, and increased cancer invasiveness and metastasis in a manner dependent on PRC2. Conversely, loss of HOTAIR can inhibit cancer invasiveness, particularly in cells that possess excessive PRC2 activity. These findings indicate that lincRNAs have active roles in modulating the cancer epigenome and may be important targets for cancer diagnosis and therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: HOX lincRNAs are systematically dysregulated in breast carcinoma and have prognostic value for metastasis and survival.
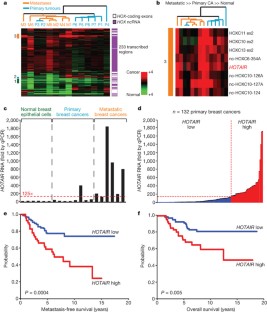
The alternative text for this image may have been generated using AI.
Figure 2: HOTAIR promotes invasion of breast carcinoma cells.

The alternative text for this image may have been generated using AI.
Figure 3: HOTAIR promotes selective re-targeting of PRC2 and H3K27me3 genome-wide.
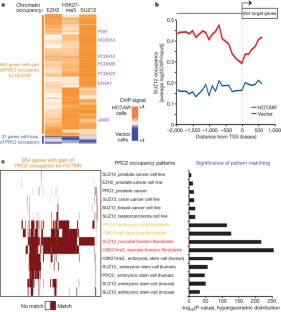
The alternative text for this image may have been generated using AI.
Figure 4: HOTAIR -induced matrix invasion and global gene expression changes requires PRC2.

The alternative text for this image may have been generated using AI.
Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
Microarray data are deposited in Gene Expression Omnibus (GEO) under accession numbers GSE20435 and GSE20737.
References
- Amaral, P. P., Dinger, M. E., Mercer, T. R. & Mattick, J. S. The eukaryotic genome as an RNA machine. Science 319, 1787–1789 (2008)
Article ADS CAS Google Scholar - The FANTOM Consortium. The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005)
- Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009)
Article ADS CAS Google Scholar - Calin, G. A. et al. Ultraconserved regions encoding ncRNAs are altered in human leukemias and carcinomas. Cancer Cell 12, 215–229 (2007)
Article CAS Google Scholar - Yu, W. et al. Epigenetic silencing of tumour suppressor gene p15 by its antisense RNA. Nature 451, 202–206 (2008)
Article ADS CAS Google Scholar - Ponting, C. P., Oliver, P. L. & Reik, W. Evolution and functions of long noncoding RNAs. Cell 136, 629–641 (2009)
Article CAS Google Scholar - Rinn, J. L. et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311–1323 (2007)
Article CAS Google Scholar - Khalil, A. M. et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl Acad. Sci. USA 106, 11667–11672 (2009)
Article ADS CAS Google Scholar - Raman, V. et al. Compromised HOXA5 function can limit p53 expression in human breast tumours. Nature 405, 974–978 (2000)
Article ADS CAS Google Scholar - Wu, X. et al. HOXB7, a homeodomain protein, is overexpressed in breast cancer and confers epithelial-mesenchymal transition. Cancer Res. 66, 9527–9534 (2006)
Article CAS Google Scholar - Sparmann, A. & van Lohuizen, M. Polycomb silencers control cell fate, development and cancer. Natl. Rev. 6, 846–856 (2006)
Article CAS Google Scholar - Kleer, C. G. et al. EZH2 is a marker of aggressive breast cancer and promotes neoplastic transformation of breast epithelial cells. Proc. Natl Acad. Sci. USA 100, 11606–11611 (2003)
Article ADS CAS Google Scholar - van de Vijver, M. J. et al. A gene-expression signature as a predictor of survival in breast cancer. N. Engl. J. Med. 347, 1999–2009 (2002)
Article CAS Google Scholar - Ma, L., Teruya-Feldstein, J. & Weinberg, R. A. Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature 449, 682–688 (2007)
Article ADS CAS Google Scholar - Novak, P. et al. Agglomerative epigenetic aberrations are a common event in human breast cancer. Cancer Res. 68, 8616–8625 (2008)
Article CAS Google Scholar - Naik, M. U., Naik, T. U., Suckow, A. T., Duncan, M. K. & Naik, U. P. Attenuation of junctional adhesion molecule-A is a contributing factor for breast cancer cell invasion. Cancer Res. 68, 2194–2203 (2008)
Article CAS Google Scholar - Fox, B. P. & Kandpal, R. P. Invasiveness of breast carcinoma cells and transcript profile: Eph receptors and ephrin ligands as molecular markers of potential diagnostic and prognostic application. Biochem. Biophys. Res. Commun. 318, 882–892 (2004)
Article CAS Google Scholar - Herath, N. I., Doecke, J., Spanevello, M. D., Leggett, B. A. & Boyd, A. W. Epigenetic silencing of EphA1 expression in colorectal cancer is correlated with poor survival. Br. J. Cancer 100, 1095–1102 (2009)
Article CAS Google Scholar - The Gene Ontology Consortium. Gene ontology: tool for the unification of biology. Nature Genet. 25, 25–29 (2000)
- Segal, E., Friedman, N., Koller, D. & Regev, A. A module map showing conditional activity of expression modules in cancer. Nature Genet. 36, 1090–1098 (2004)
Article CAS Google Scholar - Srinivasan, D. & Plattner, R. Activation of Abl tyrosine kinases promotes invasion of aggressive breast cancer cells. Cancer Res. 66, 5648–5655 (2006)
Article CAS Google Scholar - Olmeda, D. et al. SNAI1 is required for tumor growth and lymph node metastasis of human breast carcinoma MDA-MB-231 cells. Cancer Res. 67, 11721–11731 (2007)
Article CAS Google Scholar - Marinkovich, M. P. Tumour microenvironment: laminin 332 in squamous-cell carcinoma. Natl. Rev. 7, 370–380 (2007)
Article CAS Google Scholar - Tan, J. et al. Pharmacologic disruption of Polycomb-repressive complex 2-mediated gene repression selectively induces apoptosis in cancer cells. Genes Dev. 21, 1050–1063 (2007)
Article CAS Google Scholar - Sen, G. L., Webster, D. E., Barragan, D. I., Chang, H. Y. & Khavari, P. A. Control of differentiation in a self-renewing mammalian tissue by the histone demethylase JMJD3. Genes Dev. 22, 1865–1870 (2008)
Article CAS Google Scholar - Bergstraesser, L. M. & Weitzman, S. A. Culture of normal and malignant primary human mammary epithelial cells in a physiological manner simulates in vivo growth patterns and allows discrimination of cell type. Cancer Res. 53, 2644–2654 (1993)
CAS PubMed Google Scholar - Wu, J. M. et al. Heterogeneity of breast cancer metastases: comparison of therapeutic target expression and promoter methylation between primary tumors and their multifocal metastases. Clin. Cancer Res. 14, 1938–1946 (2008)
Article CAS Google Scholar - Eisen, M. B., Spellman, P. T., Brown, P. O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl Acad. Sci. USA 95, 14863–14868 (1998)
Article ADS CAS Google Scholar - Rinn, J. L., Bondre, C., Gladstone, H. B., Brown, P. O. & Chang, H. Y. Anatomic demarcation by positional variation in fibroblast gene expression programs. PLoS Genet. 2, e119 (2006)
Article Google Scholar - Adler, A. S. et al. Genetic regulators of large-scale transcriptional signatures in cancer. Nature Genet. 38, 421–430 (2006)
Article CAS Google Scholar
Acknowledgements
We thank Y. Chen-Tsai, M. Guttman, G. Sen, T. Ridky, P. Khavari, V. Band and Y. Kang for advice and reagents. Supported by National Institutes of Health (NIH), Emerald Foundation, and American Cancer Society (H.Y.C.), Dermatology Foundation (R.A.G., K.C.W. and D.J.W.), Susan Komen Foundation (M.-C.T.), NSF (T.H.), and Department of Defense BCRP (S.S.). H.Y.C. is an Early Career Scientist of the Howard Hughes Medical Institute.
Author Contributions R.A.G. measured lincRNAs in cancer samples and performed all gene transfer and knockdown experiments. R.A.G. and N.S. performed cell growth, invasion, and in vivo xenograft assays. R.A.G., K.C.W., M.-C.T. and T.H. performed ChIP-chip studies and analyses. R.A.G., J.L.R. and D.J.W. performed bioinformatic analyses. J.K. performed in vivo bioluminescence studies. H.M.H., P.A. and M.J.v.d.V. procured and analysed human tumour samples. Y.W., P.B. and B.K. designed lincRNA Taqman probes and analysed tumour RNAs by qRT–PCR. R.L. and R.B.W. performed in situ hybridization studies. R.A.G., N.S., S.S. and H.Y.C. designed the experiments and interpreted the results. R.A.G. and H.Y.C. wrote the paper.
Author information
Authors and Affiliations
- Howard Hughes Medical Institute and Program in Epithelial Biology,,
Rajnish A. Gupta, Kevin C. Wang, David J. Wong, Miao-Chih Tsai, Tiffany Hung & Howard Y. Chang - Stanford Cancer Center and Transgenic Mouse Research Center,,
Jeewon Kim - Department of Pathology, Stanford University School of Medicine, Stanford, California 94305, USA,
Rui Li & Robert B. West - Sidney Kimmel Comprehensive Cancer Center,,
Nilay Shah & Saraswati Sukumar - Department of Pathology, Johns Hopkins University School of Medicine, Baltimore, Maryland 21231, USA,
Pedram Argani - Department of Pathology, Academic Medical Center, Meibergdreef 9, 1105AZ, Amsterdam, The Netherlands,
Hugo M. Horlings & Marc J. van de Vijver - The Broad Institute of Harvard and Massachusetts Institute of Technology, Cambridge, Massachusetts 02142, USA ,
John L. Rinn - Applied Biosystems, Foster City, California 94404, USA ,
Yulei Wang, Pius Brzoska & Benjamin Kong
Authors
- Rajnish A. Gupta
- Nilay Shah
- Kevin C. Wang
- Jeewon Kim
- Hugo M. Horlings
- David J. Wong
- Miao-Chih Tsai
- Tiffany Hung
- Pedram Argani
- John L. Rinn
- Yulei Wang
- Pius Brzoska
- Benjamin Kong
- Rui Li
- Robert B. West
- Marc J. van de Vijver
- Saraswati Sukumar
- Howard Y. Chang
Corresponding author
Correspondence toHoward Y. Chang.
Supplementary information
PowerPoint slides
Rights and permissions
About this article
Cite this article
Gupta, R., Shah, N., Wang, K. et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis.Nature 464, 1071–1076 (2010). https://doi.org/10.1038/nature08975
- Received: 06 August 2009
- Accepted: 26 February 2010
- Issue date: 15 April 2010
- DOI: https://doi.org/10.1038/nature08975
This article is cited by
Editorial Summary
Chromatin in cancer metastasis
Long non-coding RNAs (lincRNAs), a relatively recently recognized class of widely transcribed genes, are thought to affect chromatin state and epigenetic regulation, but their mechanisms of action and potential roles in human disease are poorly understood. This study shows that long non-coding RNAs in the human HOX loci are systematically dysregulated during breast cancer progression, and that expression levels of the lincRNA termed HOTAIR can predict cancer metastasis. Elevated levels of HOTAIR can lead to altered patterns of Polycomb binding to the genome. These findings indicate that lincRNAs have active roles in modulating the cancer epigenome and may be important targets for cancer diagnosis and therapy.