Solid-phase extraction of N-linked glycopeptides (original) (raw)
- Protocol
- Published: 01 March 2007
Nature Protocols volume 2, pages 334–339 (2007)Cite this article
- 5972 Accesses
- 281 Citations
- 15 Altmetric
- Metrics details
Abstract
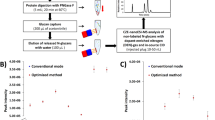
Protein glycosylation is a common post-translational modification and has been increasingly recognized as one of the most prominent biochemical alterations associated with malignant transformation and tumorigenesis. _N_-linked glycosylation is prevalent in proteins on the extracellular membrane, and many clinical biomarkers and therapeutic targets are glycoproteins. Here, we describe a protocol for solid-phase extraction of _N_-linked glycopeptides and subsequent identification of _N_-linked glycosylation sites (_N_-glycosites) by tandem mass spectrometry. The method oxidizes the carbohydrates in glycopeptides into aldehydes, which can be immobilized on a solid support. The _N_-linked glycopeptides are then optionally labeled with a stable isotope using deuterium-labeled succinic anhydride and the peptide moieties are released by peptide-_N_-glycosidase. In a single analysis, the method identifies hundreds of _N_-linked glycoproteins, the site(s) of _N_-linked glycosylation and the relative quantity of the identified glycopeptides.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Schematic diagram of SPEG.

The alternative text for this image may have been generated using AI.
Figure 2

The alternative text for this image may have been generated using AI.
Figure 3

The alternative text for this image may have been generated using AI.
Similar content being viewed by others

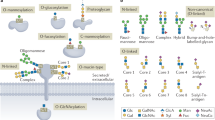
Glycoproteomics
Article 23 June 2022

References
- Spiro, R.G. Protein glycosylation: nature, distribution, enzymatic formation, and disease implications of glycopeptide bonds. Glycobiology 12, 43R–56R (2002).
Article CAS Google Scholar - Durand, G. & Seta, N. Protein glycosylation and diseases: blood and urinary oligosaccharides as markers for diagnosis and therapeutic monitoring. Clin. Chem. 46, 795–805 (2000).
CAS PubMed Google Scholar - Freeze, H.H. Update and perspectives on congenital disorders of glycosylation. Glycobiology 11, 129R–143R (2001).
Article CAS Google Scholar - Kaji, H. et al. Lectin affinity capture, isotope-coded tagging and mass spectrometry to identify _N_-linked glycoproteins. Nat. Biotechnol. 21, 667–672 (2003).
Article CAS Google Scholar - Zhang, H., Li, X.J., Martin, D.B. & Aebersold, R. Identification and quantification of _N_-linked glycoproteins using hydrazide chemistry, stable isotope labeling and mass spectrometry. Nat. Biotechnol. 21, 660–666 (2003).
Article CAS Google Scholar - Zhang, H. et al. High throughput quantitative analysis of serum proteins using glycopeptide capture and liquid chromatography mass spectrometry. Mol. Cell. Proteomics 4, 144–155 (2005).
Article CAS Google Scholar - Anderson, N.L. & Anderson, N.G. The human plasma proteome: history, character, and diagnostic prospects. Mol. Cell. Proteomics 1, 845–867 (2002).
Article CAS Google Scholar - Roth, J. Protein _N_-glycosylation along the secretory pathway: relationship to organelle topography and function, protein quality control, and cell interactions. Chem. Rev. 102, 285–303 (2002).
Article CAS Google Scholar - Bause, E. Structural requirements of _N_-glycosylation of proteins. Studies with proline peptides as conformational probes. Biochem. J. 209, 331–336 (1983).
Article CAS PubMed Central Google Scholar - Zhang, H. et al. UniPep, a database for human N-linked glycosites: a resource for biomarker discovery. Genome Biol. 7, R73 (2006).
Article PubMed Central Google Scholar - Deutsch, E.W. et al. Human Plasma PeptideAtlas. Proteomics 5, 3497–3500 (2005).
Article CAS Google Scholar - Li, X.J. et al. A software suite for the generation and comparison of Peptide arrays from sets of data collected by liquid chromatography–mass spectrometry. Mol. Cell. Proteomics 4, 1328–1340 (2005).
Article CAS Google Scholar - Liu, T. et al. Human plasma _N_-glycoproteome analysis by immunoaffinity subtraction, hydrazide chemistry, and mass spectrometry. J. Proteome Res. 4, 2070–2080 (2005).
Article CAS PubMed Central Google Scholar - Ramachandran, P. et al. Identification of _N_-linked glycoproteins in human saliva by glycoprotein capture and mass spectrometry. J. Proteome Res. 5, 1493–1503 (2006).
Article CAS Google Scholar - Pan, S. et al. Identification of glycoproteins in human cerebrospinal fluid with a complementary proteomic approach. J. Proteome Res. 5, 2769–2779 (2006).
Article CAS Google Scholar - Zhang, H. et al. Mass spectrometric detection of tissue proteins in plasma. Mol. Cell. Proteomics. 6, 64–71 (2007).
Article CAS Google Scholar - Lewandrowski, U., Moebius, J., Walter, U. & Sickmann, A. Elucidation of _N_-glycosylation sites on human platelet proteins: a glycoproteomic approach. Mol. Cell. Proteomics 5, 226–233 (2006).
Article CAS Google Scholar - Liu, A.Y., Zhang, H., Sorensen, C.M. & Diamond, D.L. Analysis of prostate cancer by proteomics using tissue specimens. J. Urol. 173, 73–78 (2005).
Article CAS Google Scholar - Gygi, S.P. et al. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat. Biotechnol. 17, 994–999 (1999).
Article CAS Google Scholar - Aebi, M. & Hennet, T. Congenital disorders of glycosylation: genetic model systems lead the way. Trends Cell Biol. 11, 136–141 (2001).
Article CAS Google Scholar - Eng, J., McCormack, A.L. & Yates, J.R., III. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Article CAS PubMed Central Google Scholar - Keller, A., Nesvizhskii, A.I., Kolker, E. & Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002).
Article CAS PubMed Central Google Scholar
Acknowledgements
This work was supported with federal funds from the National Cancer Institute, National Institutes of Health, by grant R21-CA-114852 (to H.Z.) and by Federal funds from the National Heart, Lung, and Blood Institute, National Institutes of Health, under contract no. N01-HV-8179 (to R. A.).
Author information
Authors and Affiliations
- Institute for Systems Biology, Seattle, 98103, Washington, USA
Yuan Tian, Yong Zhou, Sarah Elliott & Hui Zhang - Department of Pathology, Johns Hopkins Medical Institutions, Baltimore, 21287, Maryland, USA
Yuan Tian & Hui Zhang - Swiss Federal Institute of Technology (ETH) Zurich and Faculty of Natural Sciences, University of Zurich, Zurich, CH-8093, Switzerland
Ruedi Aebersold
Authors
- Yuan Tian
- Yong Zhou
- Sarah Elliott
- Ruedi Aebersold
- Hui Zhang
Corresponding author
Correspondence toHui Zhang.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Tian, Y., Zhou, Y., Elliott, S. et al. Solid-phase extraction of _N_-linked glycopeptides.Nat Protoc 2, 334–339 (2007). https://doi.org/10.1038/nprot.2007.42
- Published: 01 March 2007
- Issue date: February 2007
- DOI: https://doi.org/10.1038/nprot.2007.42