Creating custom synthetic genomes in Escherichia coli with REXER and GENESIS (original) (raw)
- Protocol
- Published: 26 April 2021
Nature Protocols volume 16, pages 2345–2380 (2021)Cite this article
- 7966 Accesses
- 32 Citations
- 28 Altmetric
- Metrics details
Subjects
This article has been updated
Abstract
We previously developed REXER (Replicon EXcision Enhanced Recombination); this method enables the replacement of >100 kb of the Escherichia coli genome with synthetic DNA in a single step and allows the rapid identification of non-viable or otherwise problematic sequences with nucleotide resolution. Iterative repetition of REXER (GENESIS, GENomE Stepwise Interchange Synthesis) enables stepwise replacement of longer contiguous sections of genomic DNA with synthetic DNA, and even the replacement of the entire E. coli genome with synthetic DNA. Here we detail protocols for REXER and GENESIS. A standard REXER protocol typically takes 7–10 days to complete. Our description encompasses (i) synthetic DNA design, (ii) assembly of synthetic DNA constructs, (iii) utilization of CRISPR–Cas9 coupled to lambda-red recombination and positive/negative selection to enable the high-fidelity replacement of genomic DNA with synthetic DNA (or insertion of synthetic DNA), (iv) evaluation of the success of the integration and replacement and (v) identification of non-tolerated synthetic DNA sequences with nucleotide resolution. This protocol provides a set of precise genome engineering methods to create custom synthetic E. coli genomes.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Fig. 1: Schematic of REXER and GENESIS.
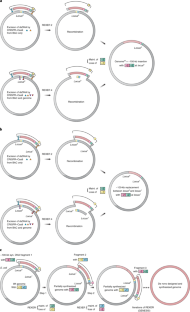
The alternative text for this image may have been generated using AI.
Fig. 2: BAC assembly and delivery.

The alternative text for this image may have been generated using AI.
Fig. 3: Components for REXER.

The alternative text for this image may have been generated using AI.
Fig. 4: REXER workflow.

The alternative text for this image may have been generated using AI.
Fig. 5: Identifying deleterious sequences in synthetic DNA.

The alternative text for this image may have been generated using AI.
Fig. 6: Testing potential fixes for deleterious synthetic sequences.
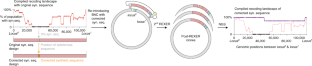
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
Data availability
No new data were generated or analyzed while preparing this protocol. Representative data are available in refs. 3,14.
Code availability
Code used for next-generation sequencing analysis is freely available at https://github.com/TiongSun/iSeq.
Change history
21 November 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41596-024-01114-8
References
- Gibson, D. G. et al. Creation of a bacterial cell controlled by a chemically synthesized genome. Science 329, 52–56 (2010).
Article CAS PubMed Google Scholar - Hutchison, C. A. 3rd et al. Design and synthesis of a minimal bacterial genome. Science 351, aad6253 (2016).
Article PubMed Google Scholar - Fredens, J. et al. Total synthesis of Escherichia coli with a recoded genome. Nature 569, 514–518 (2019).
Article CAS PubMed PubMed Central Google Scholar - Richardson, S. M. et al. Design of a synthetic yeast genome. Science 355, 1040 (2017).
Article CAS PubMed Google Scholar - Venetz, J. E. et al. Chemical synthesis rewriting of a bacterial genome to achieve design flexibility and biological functionality. Proc. Natl. Acad. Sci. USA 116, 8070 (2019).
Article CAS PubMed PubMed Central Google Scholar - Krishnakumar, R. et al. Simultaneous non-contiguous deletions using large synthetic DNA and site-specific recombinases. Nucleic Acids Res. 42, e111–e111 (2014).
Article PubMed PubMed Central Google Scholar - Santos, C. N., Regitsky, D. D. & Yoshikuni, Y. Implementation of stable and complex biological systems through recombinase-assisted genome engineering. Nat. Commun. 4, 2503 (2013).
Article PubMed Google Scholar - Santos, C. N. & Yoshikuni, Y. Engineering complex biological systems in bacteria through recombinase-assisted genome engineering. Nat. Protoc. 9, 1320–1336 (2014).
Article CAS PubMed Google Scholar - Wang, G. et al. CRAGE enables rapid activation of biosynthetic gene clusters in undomesticated bacteria. Nat. Microbiol. 4, 2498–2510 (2019).
Article PubMed Google Scholar - Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 97, 6640 (2000).
Article CAS PubMed PubMed Central Google Scholar - Wannier, T. M. et al. Improved bacterial recombineering by parallelized protein discovery. Proc. Natl. Acad. Sci. USA 117, 13689 (2020).
Article CAS PubMed PubMed Central Google Scholar - Zhang, Y., Buchholz, F., Muyrers, J. P. P. & Stewart, A. F. A new logic for DNA engineering using recombination in Escherichia coli. Nat. Genet. 20, 123–128 (1998).
Article CAS PubMed Google Scholar - Lau, Y. H. et al. Large-scale recoding of a bacterial genome by iterative recombineering of synthetic DNA. Nucleic Acids Res. 45, 6971–6980 (2017).
Article CAS PubMed PubMed Central Google Scholar - Wang, K. et al. Defining synonymous codon compression schemes by genome recoding. Nature 539, 59–64 (2016).
Article CAS PubMed PubMed Central Google Scholar - Kouprina, N. & Larionov, V. Selective isolation of genomic loci from complex genomes by transformation-associated recombination cloning in the yeast Saccharomyces cerevisiae. Nat. Protoc. 3, 371–377 (2008).
Article CAS PubMed Google Scholar - Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Article CAS PubMed PubMed Central Google Scholar - Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Article PubMed PubMed Central Google Scholar - Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform. 14, 178–192 (2013).
Article CAS PubMed Google Scholar - Mosberg, J. A., Lajoie, M. J. & Church, G. M. Lambda red recombineering in Escherichia coli occurs through a fully single-stranded intermediate. Genetics 186, 791–799 (2010).
Article CAS PubMed PubMed Central Google Scholar
Acknowledgements
This work was supported by the Medical Research Council (MRC), UK (MC_U105181009 and MC_UP_A024_1008), the Medical Research Foundation (MRF-109-0003-RG-CHIN/C0741) and an ERC Advanced Grant SGCR, all to J.W.C., and by the Lundbeck Foundation (R232-2016-3474) to J.F. K.W. was supported by the NIH DP2 Award GM140937. J.W.C. thanks H. Pelham for supporting this project. We are grateful to the LMB Media Kitchen for help with preparing materials.
Author information
Author notes
- These authors contributed equally: Wesley E. Robertson, Louise F. H. Funke, Daniel de la Torre, Julius Fredens.
Authors and Affiliations
- Medical Research Council Laboratory of Molecular Biology, Cambridge, England, UK
Wesley E. Robertson, Louise F. H. Funke, Daniel de la Torre, Julius Fredens & Jason W. Chin - Division of Biology and Biological Engineering, California Institute of Technology, Pasadena, CA, USA
Kaihang Wang
Authors
- Wesley E. Robertson
- Louise F. H. Funke
- Daniel de la Torre
- Julius Fredens
- Kaihang Wang
- Jason W. Chin
Contributions
All authors contributed to developing this protocol and writing this paper. J.W.C. supervised the project.
Corresponding authors
Correspondence toKaihang Wang or Jason W. Chin.
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Byung-Kwan Cho, Bogumil Karas and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Wang, K. et al. Nature 539, 59–64 (2016): https://doi.org/10.1038/nature20124
Fredens, J. et al. Nature 569, 514–518 (2019): https://doi.org/10.1038/s41586-019-1192-5
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Robertson, W.E., Funke, L.F.H., de la Torre, D. et al. Creating custom synthetic genomes in Escherichia coli with REXER and GENESIS.Nat Protoc 16, 2345–2380 (2021). https://doi.org/10.1038/s41596-020-00464-3
- Received: 02 May 2019
- Accepted: 16 November 2020
- Published: 26 April 2021
- Version of record: 26 April 2021
- Issue date: May 2021
- DOI: https://doi.org/10.1038/s41596-020-00464-3