AGA: Interactive pipeline for reproducible genomics analyses (original) (raw)
Software Tool Article
[version 1; peer review: 2 approved]
1, Hilary Parker2, Yingying Wei2, [...] Xaio Xia3, Leslie Cope1, Michael Ochs4, Elana Fertig1
Michael Considine1, Hilary Parker2, [...] Yingying Wei2, Xaio Xia3, Leslie Cope1, Michael Ochs4, Elana Fertig1
1 Department of Oncology Biostatistics & Bioinformatics, Johns Hopkins University School of Medicine, Baltimore, MD, 21205, USA
2 Department of Biostatistics, Johns Hopkins Bloomberg School of Public Health, Baltimore, MD, 21205, USA
3 Department of Statistics and Biostatistics, Rutgers University, New Brunswick, NJ, 08901, USA
4 Department of Mathematics and Statistics, The College of New Jersey, Ewing Township, NJ, 08618, USA
OPEN PEER REVIEW
REVIEWER STATUS
Abstract
Automated Genomics Analysis (AGA) is an interactive program to analyze high-throughput genomic data sets on a variety of platforms. An easy to use, point and click, guided pipeline is implemented to combine, define, and compare datasets, and customize their outputs. In contrast to other automated programs, AGA enables flexible selection of sample groups for comparison from complex sample annotations. Batch correction techniques are also included to further enable the combination of datasets from diverse studies in this comparison. AGA also allows users to save plots, tables and data, and log files containing key portions of the R script run for reproducible analyses. The link between the interface and R supports collaborative research, enabling advanced R users to extend preliminary analyses generated from bioinformatics novices.
Keywords
automated, genomic, analysis, datasets, DNA, methylation, expression, arrays
Corresponding authors: Michael Considine, Elana Fertig Competing interests: No competing interests were disclosed.
Grant information: Funding was provided by the JHU Head and Neck SPORE, NCI (CA141053) to EJF, and NLM (LM011000) to MFO.
The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Copyright: © 2015 Considine M et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. How to cite: Considine M, Parker H, Wei Y et al. AGA: Interactive pipeline for reproducible genomics analyses [version 1; peer review: 2 approved]. F1000Research 2015, 4:28 (https://doi.org/10.12688/f1000research.6030.1) First published: 28 Jan 2015, 4:28 (https://doi.org/10.12688/f1000research.6030.1) Latest published: 21 Oct 2015, 4:28 (https://doi.org/10.12688/f1000research.6030.2)
Introduction
While high dimensional genetic data have increased in availability at reduced cost, robust analyses remain labor intensive and costly. Numerous automated software pipelines have been developed in an effort to increase the rate and decrease the costs at which analyses can be completed, including SVAw10, Partek3, InSilicoDB17, and cBioPortal4. Automated Genomics Analysis (AGA) provides a more dynamic experience, allowing the user to start with raw data and a text file containing corresponding sample annotations from either a single or multiple studies. AGA performs all necessary normalization and batch correction, and then enables the user to interactively determine the samples to contrast in the analysis based on the sample annotations. AGA is implemented in R to facilitate adaptation of state-of-the art genomics analysis techniques. Linking R to a web browser-based interface through RStudio’s shiny also facilitates collaborative analyses in research teams with diverse bioinformatics expertise.
AGA bridges the gap between interactive and reproducible analyses for several platforms, including expression arrays, methylation arrays, and processed RNAseq data. Through the interface, the user determines the size and scope of the analyses. AGA first performs data normalization, including the ComBat6 and SVA8 batch correction algorithms to enable comparison across multiple datasets for non-methylation platforms. The software then performs differential analysis15, and gene set analyses1,15 based upon defined sample groups. Users obtain standard visualization of genomics data, including hierarchical clustering, boxplots and heatmaps as part of the default analysis. Plots and tables summarizing the results from each analysis are customizable through the interface. The figures and tables in AGA are interactive and customizable. In contrast to other point and click software, AGA logs the R code, and exports the workspace with each figure and table, ensuring that each analysis can be reproduced and further customized.
Methods
The AGA application is run through R and interactive through web browsers. AGA is implemented with RStudio’s shiny12, integrating the R code used in the analysis with HTML and JavaScript, for the interactive user interface. Usage requires R version 3.0.1 or higher, and either Mozilla Firefox or Google Chrome, and R packages described in the AGA User’s Manual. The program is divided into seven tabs. Clicking the respective Update button generates the results to be displayed in each tab and clicking the Download buttons save the plots and data.
Data platforms
AGA supports analyses of DNA methylation and gene expression data. Currently, AGA supports DNA methylation analysis on Illumina 450k arrays. It also supports gene expression analysis of any human Affymetrix expression platform, including exon arrays, and normalized gene counts from RNAseq data. Notably, the flexible format for normalized RNAseq data may be adapted to analyze normalized data from other platforms measuring continuous data, many of which we plan to incorporate in future versions of AGA.
Initiation
Users of AGA select to load annotation files and high throughput genomic data from files in a specified directory. AGA accepts raw CEL files and iDat files for Affymetrix and DNA methylation arrays, respectively. It is assumed that normalized RNAseq data are formatted as individual text files for each sample, containing gene names and normalized counts for each sample. More details about the format for each data type are provided in the User’s manual. Sample annotations are specified in a CSV file, whose first column matches the names of the data files. By default, it is assumed the annotation file defines the sample batch; however, this can be updated by editing the annotation files to contain a ‘Batch’ column with unique identifiers for each respective batch within the dataset. Further details about the sample annotations are also provided in the User’s manual.
Sample selection for differential analysis
After loading in the annotation files, AGA users select categories from the annotation for differential expression analysis. AGA automatically groups samples with common levels in each category as groups for differential analysis. Samples may be further subset from the complete dataset from the criteria selected for each group. When selected, AGA updates the display to output the sample size for each group. Samples are set for analysis by clicking the “Run the Analysis!” button. In cases for which samples span multiple batches, the analysis automatically performs ComBat and SVA batch correction protecting for the biological groups in the annotation selected by the user. Help boxes are available to clarify each input field with further details in the User’s manual.
Interactive plots and tables
The Dendrogram Plot tab in displays unsupervised hierarchical clustering based upon the complete correlation between values of genes (rows) and samples (columns). The Heatmap Plot tab provides an interactive Javascript heatmap of the genomic data, allowing users to customize genes plotted and color rows by sample annotations. For both Dendrograms and Heatmaps, an option is available to view the pre-batch corrected data to show the effects of batch on and efficacy of correction of the data. The Gene Box Plot tab creates boxplots to summarize values of a user-selected gene in the selected groups.
The Differential Results tab displays the results from the differential analysis using empirical Bayes moderated t-statistics with the Bio-conductor Package limma15. Statistics are computed on data that have been batch corrected by combining ComBat with SVA, protecting for the biological groups selected for comparison9. The p-values are adjusted utilizing the Benjamini-Hotchberg method for multiple hypothesis testing7. Optionally, gene set statistics can be performed for each gene set defined in Biocarta and Gene Ontology using a Wilcoxon rank-sum test comparing the t-statistics from the most differentially expressed probe for genes in the set to similarly selected t-statistics for genes outside of the set. If selected, results from gene set analysis are displayed in the GSA Results tab.
Example
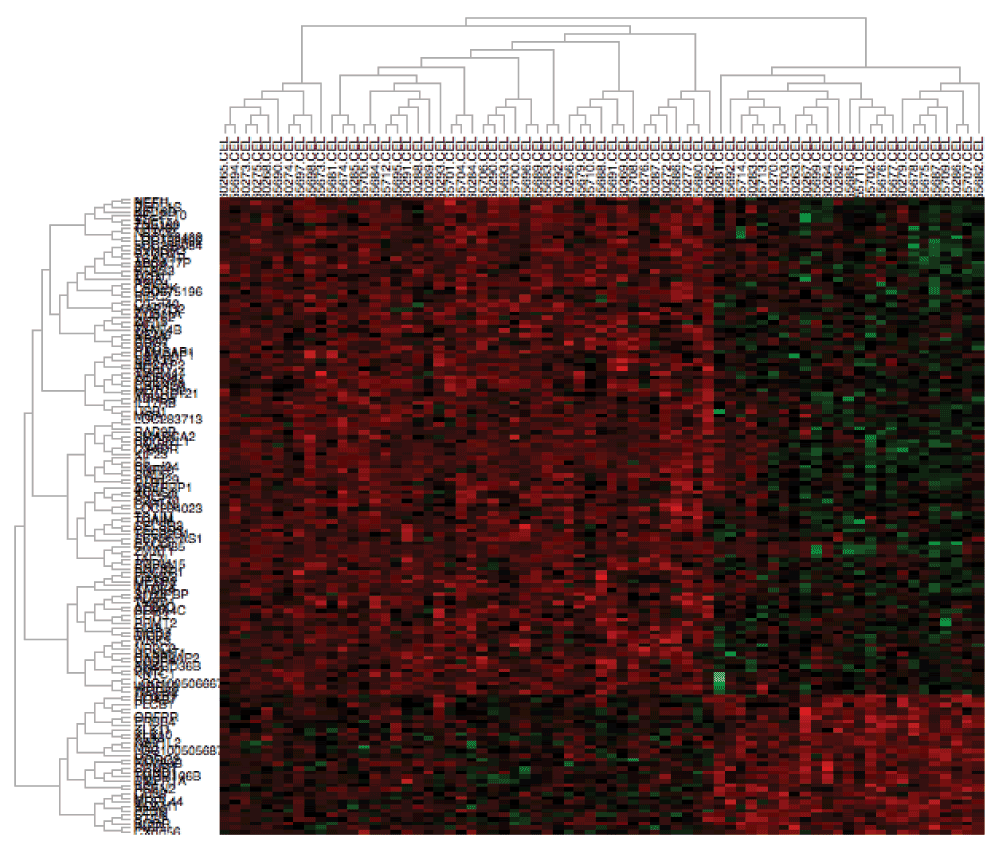
As an example, we perform analysis on sample datasets containing gene expression of primary head and neck squamous cell carcinoma (HNSCC) tumors. We downloaded measurements from a combination of frozen tumor samples from two distinct studies in GEO available under accession numbers GSE103002 and GSE679111, representing two distinct batches. Raw CEL files and annotation csv files were obtained as described in the User’s manual. We initialize AGA by selecting the directory containing these data. Once loaded, we check the HPV and Tumor.Source.Type columns to group the samples into primary HPV-positive and HPV-negative tumors for differential expression analysis. We then click “Run the Analysis” to normalize the CEL files with fRMA5, batch correct the data with ComBat and SVA, and perform differential expression analysis. The plot in the Dendrogram Plot tab confirms that the batch effects are apparent between these datasets but removed after batch. The heatmap generated in the Heatmap Plot tab (Figure 1) demonstrates that the batch correction nonetheless preserves gene expression difference between HPV-positive and HPV-negative tumors. Moreover, performing differential expression analysis comparing HPV-positive and HPV-negative HNSCC in the “Differential Analysis” tab confirms the well-established overexpression (p=8.74e-9) of CDKN2A (p16) in HPV-positive HNSCC13,14.
Figure 1. Heatmap displaying the relative expression of the 150 probes with the lowest p values from the example analysis, including CDKN2A.
Discussion
AGA provides an interface to enable users who may be unfamiliar with R to perform reproducible genomics class comparison analysis. Unlike other automated pipelines, experienced R users can reproduce, extend or modify preliminary analyses. Thus, AGA facilitates collaborations between novice and expert R users for genomics analysis. Future work will extend the AGA pipeline to encode normalization routines to DNA methylation, and analysis routines for other genomics platforms, including copy number data.
Software availability
Author contributions
MFO and EJF conceived the software and EJF and MC designed the web interface. MC designed and coded implemented the software application, and prepared the manuscript. HSP researched and composed cross-study normalization techniques. XXX standardized annotation files for the two example data sets. YW and LC assisted by providing the initial coding for alternative analyses. All authors helped prepare the manuscript.
Competing interests
No competing interests were disclosed.
Grant information
Funding was provided by the JHU Head and Neck SPORE, NCI (CA141053) to EJF, and NLM (LM011000) to MFO.
I confirm that the funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Acknowledgements
We would like to thank Joe Cheng and Winston Chang of RStudio for their support with shiny. Alla Guseynova, Michael Fox and Louis Franceschi are very much appreciated their technical support and implementations of various iterations of the project. We thank Thomas Considine for his assistance in proofreading this manuscript; and Bahman Afsari and Thomas Considine for testing the application and User Manual. Finally, we also thank Luigi Marchionni and Jean-Philippe Fortin for collaborative efforts.
Faculty Opinions recommended
References
- 1. Ashburner M, Ball CA, Blake JA, et al.: Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000; 25(1): 25–29. PubMed Abstract | Publisher Full Text | Free Full Text
- 2. Cohen EE, Zhu H, Lingen MW, et al.: A feed-forward loop involving protein kinase Calpha and microRNAs regulates tumor cell cycle. Cancer Res. 2009; 69(1): 65–74. PubMed Abstract | Publisher Full Text | Free Full Text
- 3. Downey T: Analysis of a multifactor microarray study using Partek genomics solution. Methods Enzymol. 2006; 411: 256–270. PubMed Abstract | Publisher Full Text
- 4. Gao JB, Aksoy A, Dogrusoz U, et al.: Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013; 6(269): pl1. PubMed Abstract | Publisher Full Text | Free Full Text
- 5. Irizarry RA, Hobbs B, Collin F, et al.: Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics. 2003; 4(2): 249–264. PubMed Abstract | Publisher Full Text
- 6. Johnson WE, Li C, Rabinovic A: Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics. 2007; 8(1): 118–127. PubMed Abstract | Publisher Full Text
- 7. Klipper-Aurbach Y, Wasserman M, Braunspiegel-Weintrob N, et al.: Mathematical formulae for the prediction of the residual beta cell function during the first two years of disease in children and adolescents with insulin-dependent diabetes mellitus. Med Hypotheses. 1995; 45(5): 486–490. PubMed Abstract | Publisher Full Text
- 8. Leek JT, Johnson WE, Parker HS, et al.: The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics. 2012; 28(6): 882–883. PubMed Abstract | Publisher Full Text | Free Full Text
- 9. Parker H: Practical Statispractical Statistical Issues in Translational Genomical Issues in Translational Genomics (doctoral dissertation). Johns Hopkins University, Baltimore. 2013.
- 10. Pirooznia M, Seifuddin F, Goes FS, et al.: SVAw - a web-based application tool for automated surrogate variable analysis of gene expression studies. Source Code Biol Med. 2013; 8(1): 8. PubMed Abstract | Publisher Full Text | Free Full Text
- 11. Pyeon D, Newton MA, Lambert PF, et al.: Fundamental differences in cell cycle deregulation in human papillomavirus-positive and human papillomavirus-negative head/neck and cervical cancers. Cancer Res. 2007; 67(10): 4605–4619. PubMed Abstract | Publisher Full Text | Free Full Text
- 12. RStudio and Inc: shiny: Web Application for R. R package version 0.7.0. 2012. Reference Source
- 13. Robinson M, Sloan P, Shaw R: Refining the diagnosis of oropharyngeal squamous cell carcinoma using human papillomavirus testing. Oral Oncol. 2010; 46(7): 492–6. PubMed Abstract | Publisher Full Text
- 14. Smeets SJ, et al.: A novel algorithm for reliable detection of human papillomavirus in paraffin embedded head and neck cancer specimen. Int J Cancer. 2007; 121(11): 2465–72. PubMed Abstract | Publisher Full Text
- 15. Smyth GK: Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol. 2012; 3: 1. Article3. PubMed Abstract | Publisher Full Text
- 16. Subramanian A, Tamayo P, Mootha VK, et al.: Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A. 2005; 102(43): 15545–15550. PubMed Abstract | Publisher Full Text | Free Full Text
- 17. Taminau J, Steenhoff D, Coletta A, et al.: inSilicoDb: an R/Bioconductor package for accessing human Affymetrix expert-curated datasets from GEO. Bioinformatics. 2011; 27(22): 3204–3205. PubMed Abstract | Publisher Full Text
- 18. Considine M, Parker HS, Wei Y, et al.: Automated Genomics Analysis. Zenodo. 2015. Data Source
Comments on this article Comments (0)
Version 2
VERSION 2 PUBLISHED 28 Jan 2015
Funding was provided by the JHU Head and Neck SPORE, NCI (CA141053) to EJF, and NLM (LM011000) to MFO.
The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
© 2015 Considine M et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Open Peer Review
Current Reviewer Status: ?
Key to Reviewer Statuses VIEW HIDE
ApprovedThe paper is scientifically sound in its current form and only minor, if any, improvements are suggested
Approved with reservations A number of small changes, sometimes more significant revisions are required to address specific details and improve the papers academic merit.
Not approvedFundamental flaws in the paper seriously undermine the findings and conclusions
Version 1
VERSION 1
PUBLISHED 28 Jan 2015
Reviewer Report 22 Jun 2015
Matthew McCall, Department of Biostatistics and Computational Biology, University of Rochester, Rochester, NY, USA
Approved
VIEWS 0
Competing Interests: No competing interests were disclosed.
Close
Reviewer Report 02 Feb 2015
Subha Madhavan, Innovation Center for Biomedical Informatics, Georgetown University, Washington, DC, USA
Approved
VIEWS 0
Competing Interests: No competing interests were disclosed.
Close
Comments on this article Comments (0)
Version 2
VERSION 2 PUBLISHED 28 Jan 2015
Open Peer Review
Reviewer Status
Alongside their report, reviewers assign a status to the article:
Approved
The paper is scientifically sound in its current form and only minor, if any, improvements are suggested
Approved with reservations
A number of small changes, sometimes more significant revisions are required to address specific details and improve the papers academic merit.
Not approved
Fundamental flaws in the paper seriously undermine the findings and conclusions
Reviewer Reports
| Invited Reviewers | ||
|---|---|---|
| 1 | 2 | |
| Version 2 (revision) 21 Oct 15 | ||
| Version 1 28 Jan 15 | read | read |
- Subha Madhavan, Georgetown University, Washington, USA
- Matthew McCall, University of Rochester, Rochester, USA
Comments on this article
Sign up for content alerts
Browse by related subjects
Alongside their report, reviewers assign a status to the article:
Approved - the paper is scientifically sound in its current form and only minor, if any, improvements are suggested
Approved with reservations - A number of small changes, sometimes more significant revisions are required to address specific details and improve the papers academic merit.
Not approved - fundamental flaws in the paper seriously undermine the findings and conclusions
Adjust parameters to alter display
View on desktop for interactive features ![]()
Includes Interactive Elements
View on desktop for interactive features ![]()