Impact of Regulatory Variation from RNA to Protein (original) (raw)
. Author manuscript; available in PMC: 2015 Aug 6.
Published in final edited form as: Science. 2014 Dec 18;347(6222):664–667. doi: 10.1126/science.1260793
Abstract
The phenotypic consequences of expression quantitative trait loci (eQTLs) are presumably due to their effects on protein expression levels. Yet, the impact of genetic variation, including eQTLs, on protein levels remains poorly understood. To address this, we mapped genetic variants that are associated with eQTLs, ribosome occupancy (rQTLs), or protein abundance (pQTLs). We found that most QTLs are associated with transcript expression levels, with consequent effects on ribosome and protein levels. However, eQTLs tend to have significantly reduced effect sizes on protein levels, suggesting that their potential impact on downstream phenotypes is often attenuated or buffered. Additionally, we identified a class of cis QTLs that affect protein abundance with little or no effect on mRNA or ribosome levels, suggesting that they may arise from differences in post-translational regulation.
To understand the links between genetic and phenotypic variation it may be essential to first understand how genetic variation impacts the regulation of gene expression. Previous studies have evaluated the association between variation and transcript expression in humans (1-3). Yet protein abundances are more direct determinants of cellular functions (4), and the impact of genetic differences on the multi-stage process of gene expression through transcription and translation to steady state protein levels, has not been fully characterized. Studies in model organisms have shown that variation in mRNA and protein expression levels are often uncorrelated (5-8). Comparative studies (9-13) have suggested that protein expression evolves under greater evolutionary constraint than transcript levels (14), and provided evidence consistent with buffering of protein expression with respect to variation introduced at the transcript level. Yet, in contrast to comparative work, there are few reports of QTLs associated with protein levels (pQTLs) in humans (15-17).
Here we present a unified analysis of the association of genetic variation with transcript expression, ribosome profiling (18), and steady state protein levels in a set of HapMap Yoruba (Ibadan, Nigeria) lymphoblastoid cell lines (LCLs). We collected ribosome profiling data for 72 Yoruba LCLs and quantified protein abundance in 62 of these lines. Genome-wide genotypes and RNA-sequencing data were available for all lines (19).
Ribosome profiling is an effective way to measure changes in translational regulation using sequencing (11). We obtained a median coverage of 12 million mapped reads per sample and, as expected, the ribosome profiling reads are highly concentrated within coding regions and show an enrichment of a 3-bp periodicity, reflecting the progression of a translating ribosome (Figs. S1-S3, Table S1).
We collected relative protein expression measurements using a SILAC internal standard sample (20) and quantitative protein mass spectrometry (Fig. S4). To confirm the quality of the proteomics data (Table S2) we evaluated the agreement between measurements of distinct groups of peptides from the same protein. Differences between these measurements can reflect true biological variation (e.g., splicing), or experimental noise. The high correlations (Spearman's rho 0.7-0.9; R2 of 0.3-0.7; depending on the sample) confirmed that we are able to precisely quantify inter-individual variation in protein levels (Fig. S5). We also analyzed quantifications of peptides that overlapped non-synonymous SNPs that were heterozygous in either the analyzed or the internal standard sample (Fig. S6). The median ratios measured from these peptides matched the expected values closely, indicating that our protein measurements were likely not subject to ratio compression (Figs. S7, S8).
As a final quality check, we considered variation in expression levels within and between genes. We found that transcript and protein expression levels – which are the furthest removed processes studied here - are the least correlated (Fig. S9, S10). Our observations are in agreement with most high-throughput studies that considered large number of samples, although smaller studies have often observed higher correlations (18, 21, 22).
We mapped genetic associations with regulatory phenotypes. First, we evaluated QTLs for each phenotype independently by testing for association between the phenotype and all genetic variants with minor allele frequency >10% in a 20 kb window around the corresponding gene. We used a shared standardization, normalization, regression, and permutation pipeline for all three phenotypes. At an FDR of 10% we detected 2,355 eQTLs, 939 rQTLs, and 278 pQTLs (Table 1, Fig. S11).
Table 1. Number of _cis_-QTLs identified at False Discovery Rate (FDR) 10%.
| Measurement | Genes Tested | Number of cell lines | cis-QTLs |
|---|---|---|---|
| Protein Abundance | 4,381 | 62 | 278 |
| Ribosome Occupancy | 15,059 | 72 | 939 |
| mRNA Expression | 16,614 | 75 | 2,355 |
There is substantial overlap among detected QTLs (Fig. S12). Among the 4,322 genes quantified for all three phenotypes, 54% of the genes with pQTLs also have a significant rQTL and/or eQTL. Given the incomplete statistical power to detect QTLs in each dataset independently, we performed replication testing across datasets, using the specific SNP-gene pairs underlying each class of QTLs. This analysis is less sensitive to power limitations than genome-wide testing. The results confirm that many QTLs are shared across all three phenotypes (example in Fig. 1A). In particular, most (90%) genetic variants associated with ribosome occupancy are also associated with transcript levels (Fig. S13). In contrast, eQTLs showed the lowest overlap with pQTLs (35%), as expected (Fig. 1B).
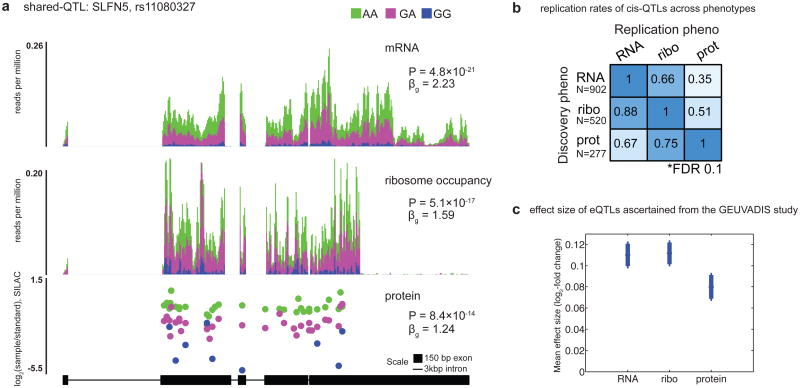
Figure 1. Comparisons of QTLs at three levels of gene regulation.
a Many QTLs exhibit shared effects across mRNA, ribosome occupancy and protein. This example illustrates a shared QTL for the schlafen family member 5 (SLFN5) gene (24). The upper panels show mean sequence depth (per bp) for mRNA and ribosome occupancy, averaged among individuals with each genotype at the QTL SNP. The lower panel shows median log2 SILAC ratios at each detected peptide, relative to the shared internal standard.
b. Replication rates between independently tested _cis_-QTLs for each phenotype pair, at FDR=10%. QTLs detected for the phenotype labeled on each row were tested in the phenotype listed for each column, considering only the 4,322 genes quantified in all three phenotypes.
c. On average, eQTLs exhibit attenuated effects on protein abundance but not on ribosome occupancy. We used eQTLs detected by the GEUVADIS study to avoid ascertainment bias, and we polarized the alleles according to the direction of effect in GEUVADIS. The plot shows mean effect sizes and standard errors on the means, measured as expected fold-change per allele copy on a log2 scale.
Our observation that many SNPs identified as eQTLs are not associated with differences in protein levels is consistent with the notion that, across species, protein levels diverge less than transcript levels (12-17). Yet some QTLs may not replicate at the protein level simply due to incomplete mapping power. To address this, and to avoid over-estimation of effect sizes due to ascertainment bias at significant QTLs, we focused on eQTLs detected previously in European samples by the GEUVADIS study (2). We then attempted to replicate the GEUVADIS eQTLs using our transcript, ribosome profiling, and protein data and considered the mean QTL effect size in each data type. Mean effect sizes calculated in this way are expected to be unbiased with respect to either technical or biological variance.
Using this approach, we observed a reduced mean effect size for the GEUVADIS eQTLs in the protein data compared to either the RNA-seq data (t-test P = 6.7×10-3) or ribosome data (P = 5.6×10-3; Fig. 1C). In contrast, the average effect sizes observed for the RNA-seq and ribosome data are not significantly different from each other, and their effect sizes are highly correlated across the tested eQTLs (Pearson c = 0.79, P < 10-96, Fig. S14). The reduction in effect size observed in protein data is robust with respect to potential technical confounders, including iBAQ intensity level and transcript model complexity (Fig. S15). We thus conclude that the majority of genetic variants affecting transcript levels also alter ribosomal occupancy, typically with a similar magnitude of effect. Yet, both eQTL mapping and effect-size analyses indicate that many eQTLs have attenuated (or absent) effects on steady state protein levels (Fig. S16).
In addition to the observation of generally attenuated effect sizes in pQTLs compared with eQTLs, we identified a subset of variants that appear to affect levels of proteins but not mRNA, and hence are candidates to affect post-transcriptional gene regulation. To evaluate evidence for these, we tested each SNP for association with one regulatory phenotype, while treating one or both of the other phenotypes as covariates (conditional model). Considering protein levels, with RNA levels as a covariate, we identified 146 protein-specific QTLs (psQTLs) at FDR = 10% (Fig. 2A). The identification of psQTLs is generally robust to the choice of technology used to characterize transcript expression (Fig S17).
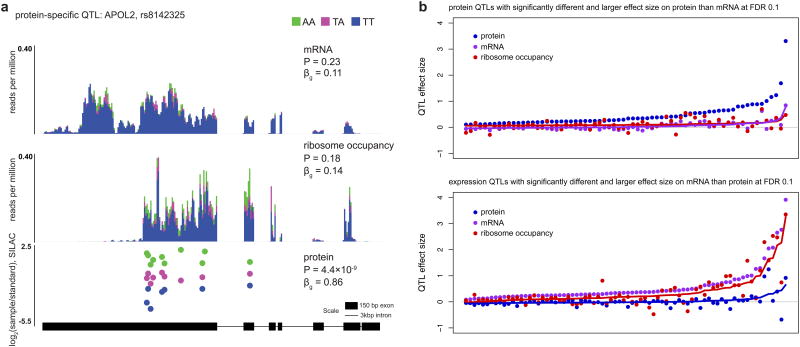
Figure 2. Protein-specific and RNA-specific QTLs.
a An example of a protein-specific QTL, for the apolipoprotein L, 2 (APOL2) gene, detected by both the interaction model and the conditional models, indicating both larger effect (LRT, P = 3.3×10-6, interaction model; P = 5.1×10-13, conditional model) in protein than mRNA, and that the effect on protein is not mediated by either mRNA or ribosome occupancy (LRT, P = 2.1×10-12, conditional model). Plotting details as in Figure 1A. While the causal variant underlying this pQTL is unknown, several linked variants near the 3′ end of APOL2 are all strongly associated with protein levels, including rs8142325 shown here and missense variant rs7285167 (βg = 0.83, P = 9.8×10-9; LRT, P = 2.1×10-5, interaction model; P = 5.5×10-13, conditional model).
b. Effect sizes for ribosome occupancy tend to track with RNA, not protein. Top panel: effect sizes in all three phenotypes are shown for protein-specific QTLs. Effect sizes were estimated using linear regression in each of the phenotypes independently. The signs of the effects were set to be positive in protein. Solid lines reflect predicted effects based on a linear model. Bottom panel: Similarly, effect sizes in all three phenotypes for esQTLs. Here, signs of the effects were set to be positive in RNA.
Using an alternative approach, an interaction model, we identified 68 psQTLs with significantly larger effects in protein than mRNA (LRT; FDR = 10%). We also used the interaction model to identify 76 expression-specific QTLs (esQTLs, interaction model, LRT; FDR = 10%). We then considered the ribosomal data. We found that the effect sizes for ribosomal occupancy are similar to the esQTL effect sizes (Fig. 2B). Yet, for psQTLs, low ribosome effect sizes are observed. Thus, for QTLs with discordant effects between transcript and protein, the ribosome data usually tracked with levels of RNA. Put together, these results allow us to identify loci where genetic variants have specific impacts on protein levels that are not fully mediated by regulation of either transcription or translation and hence may affect rates of protein degradation.
Finally, we performed enrichment analysis in which we considered each tested gene-SNP pair separately, and evaluated the full distribution of p-values from the conditional model (rather than choosing a significance threshold) for different genomic and functional annotations. SNPs within transcribed regions (exonic and UTR) are enriched for more significant psQTL effects, compared to intergenic or intronic SNPs, even within the narrow 20 kb windows tested (Fig. S18, S19, S20). In addition, psQTLs are further enriched for non-synonymous sites (compared to all exonic SNPs, Table 2).
Table 2. Enrichment of genomic annotations among expression and protein-specific QTLs.
Enrichments were evaluated by a continuous test using QTL results from the conditional model (see Supplementary Information). Columns (from left to right) describe the annotation being considered, the number of SNPs matching this annotation, the set of SNPs used as background for the corresponding test, the enrichment p-values for protein-specific QTLs (psQTLs), and expression-specific QTLs (esQTLs), respectively.
| Annotation | #SNPs | Background | psQTLs | esQTLs |
|---|---|---|---|---|
| Exonic | 12568 | Intergenic | 2.8×10-14 | 2.3×10-21 |
| 5′ UTR | 6488 | Intergenic | 3.2×10-5 | 5.9×10-19 |
| 3′ UTR | 15139 | Intergenic | 2.0×10-6 | 1.7×10-16 |
| Intronic | 628591 | Intergenic | 7.1×10-3 | 2.9×10-38(*depl) |
| Non-synonymous | 2099 | Exonic | 5.7×10-3 | 9.7×10-2 |
| Ribo SNitch | 414 | Exonic | 5.2×10-2 | 2.5×10-2 |
| Acetylation site | 22 | Non-synonymous | 3.2×10-2 | 0.62 |
Investigating additional annotations (Table 2, Fig. S21), we found that non-synonymous SNPs near acetylation sites showed nominal enrichment for psQTLs. This possibly reflects the functional role of lysine acetylation in modulating protein degradation (23). Overall, the enrichment results suggest that genetic variants involved in post-transcriptional regulation are functionally distinct from genetic variants that primarily affect transcription – they are more likely to fall within translated regions of the gene, and more likely to occur at non-synonymous sites.
In summary, we have shown that while a substantial fraction of regulatory genetic variants influence gene expression at all levels from mRNA to steady state protein abundance, there are also a number of effects with specific impact on particular expression phenotypes. QTLs affecting mRNA levels are on average attenuated or buffered at the protein level, as has been observed between species (14). Our analysis indicates that this attenuation is not evident at the stage of translation. While the overall phenotypic similarity between ribosome occupancy and protein abundance is high, _cis_-regulatory genetic effects on ribosome occupancy appear to be more strongly shared with mRNA than with protein. These observations, along with the phenotype-specific QTL analysis, indicate a scarcity of translation-specific QTLs, and minimal attenuation of genetic impact between mRNA and ribosome phenotypes.
Supplementary Material
Supplementary Data 1
Supplementary Data 2
Supplementary Data 3
Supplementary Data 4
Supplementary Information
Acknowledgments
We thank M. Stephens, T. Flutre, and members of the Pritchard and Gilad labs for helpful discussions. This work was supported by NIH grants GM077959, HG007036 and MH084703. AB and JKP were supported the Howard Hughes Medical Institute. ZK was supported by F32HG006972. The proteomics data are available at ProteomeXchange (accession PXD001406). The ribosome profiling data are available at GEO (accession GSE61742). JKP is on the Senior Advisory Board for 23andMe and DNANexus and holds stock in both. AB holds stock in Google, Inc.
References
- 1.Battle A, et al. Characterizing the genetic basis of transcriptome diversity through RNA-sequencing of 922 individuals. Genome research. 2014 Jan;24:14. doi: 10.1101/gr.155192.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Lappalainen T, et al. Transcriptome and genome sequencing uncovers functional variation in humans. Nature. 2013 Sep 26;501:506. doi: 10.1038/nature12531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Grundberg E, et al. Mapping cis- and trans-regulatory effects across multiple tissues in twins. Nature genetics. 2012 Oct;44:1084. doi: 10.1038/ng.2394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Vogel C, Marcotte EM. Insights into the regulation of protein abundance from proteomic and transcriptomic analyses. Nature reviews Genetics. 2012 Apr;13:227. doi: 10.1038/nrg3185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ghazalpour A, et al. Comparative analysis of proteome and transcriptome variation in mouse. PLoS genetics. 2011 Jun;7:e1001393. doi: 10.1371/journal.pgen.1001393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Foss EJ, et al. Genetic basis of proteome variation in yeast. Nature genetics. 2007 Nov;39:1369. doi: 10.1038/ng.2007.22. [DOI] [PubMed] [Google Scholar]
- 7.Picotti P, et al. A complete mass-spectrometric map of the yeast proteome applied to quantitative trait analysis. Nature. 2013 Feb 14;494:266. doi: 10.1038/nature11835. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Albert FW, Treusch S, Shockley AH, Bloom JS, Kruglyak L. Genetics of single-cell protein abundance variation in large yeast populations. Nature. 2014 Feb 27;506:494. doi: 10.1038/nature12904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Laurent JM, et al. Protein abundances are more conserved than mRNA abundances across diverse taxa. Proteomics. 2010;10:4209. doi: 10.1002/pmic.201000327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Schrimpf SP, et al. Comparative Functional Analysis of the Caenorhabditis elegans and Drosophila melanogaster Proteomes. PLoS Biol. 2009;7:e1000048. doi: 10.1371/journal.pbio.1000048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Stadler M, Fire A. Conserved translatome remodeling in nematode species executing a shared developmental transition. PLoS genetics. 2013;9:e1003739. doi: 10.1371/journal.pgen.1003739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Artieri CG, Fraser HB. Evolution at two levels of gene expression in yeast. Genome research. 2014 Mar;24:411. doi: 10.1101/gr.165522.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.McManus CJ, May GE, Spealman P, Shteyman A. Ribosome profiling reveals post-transcriptional buffering of divergent gene expression in yeast. Genome research. 2014 Mar;24:422. doi: 10.1101/gr.164996.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Khan Z, et al. Primate transcript and protein expression levels evolve under compensatory selection pressures. Science (New York, N Y) 2013 Nov 29;342:1100. doi: 10.1126/science.1242379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Wu L, et al. Variation and genetic control of protein abundance in humans. Nature. 2013;499:79. doi: 10.1038/nature12223. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Hause RJ, et al. Identification and Validation of Genetic Variants that Influence Transcription Factor and Cell Signaling Protein Levels. American journal of human genetics. 2014 Aug 7;95:194. doi: 10.1016/j.ajhg.2014.07.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Garge N, et al. Identification of quantitative trait loci underlying proteome variation in human lymphoblastoid cells. Molecular & cellular proteomics : MCP. 2010 Jul;9:1383. doi: 10.1074/mcp.M900378-MCP200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ingolia NT, Ghaemmaghami S, Newman JR, Weissman JS. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science (New York, N Y) 2009 Apr 10;324:218. doi: 10.1126/science.1168978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Pickrell JK, et al. Understanding mechanisms underlying human gene expression variation with RNA sequencing. Nature. 2010 Apr 1;464:768. doi: 10.1038/nature08872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ong SE, et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Molecular & cellular proteomics : MCP. 2002 May;1:376. doi: 10.1074/mcp.m200025-mcp200. [DOI] [PubMed] [Google Scholar]
- 21.Wilhelm M, et al. Mass-spectrometry-based draft of the human proteome. Nature. 2014 May 29;509:582. doi: 10.1038/nature13319. [DOI] [PubMed] [Google Scholar]
- 22.Schwanhausser B, et al. Global quantification of mammalian gene expression control. Nature. 2011 May 19;473:337. doi: 10.1038/nature10098. [DOI] [PubMed] [Google Scholar]
- 23.Yang XJ, Seto E. Lysine acetylation: codified crosstalk with other posttranslational modifications. Molecular cell. 2008 Aug 22;31:449. doi: 10.1016/j.molcel.2008.07.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Lee MN, et al. Common genetic variants modulate pathogen-sensing responses in human dendritic cells. Science (New York, N Y) 2014 Mar 7;343:1246980. doi: 10.1126/science.1246980. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Data 1
Supplementary Data 2
Supplementary Data 3
Supplementary Data 4
Supplementary Information