Enzymatic properties of de novo-type mouse DNA (cytosine-5) methyltransferases - PubMed (original) (raw)
Enzymatic properties of de novo-type mouse DNA (cytosine-5) methyltransferases
A Aoki et al. Nucleic Acids Res. 2001.
Abstract
We have purified GST-fused recombinant mouse Dnmt3a and three isoforms of mouse Dnmt3b to near homogeneity. Dnmt3b3, an isoform of Dnmt3b, did not have DNA methylation activity. Dnmt3a, Dnmt3b1 or Dnmt3b2 showed similar activity toward poly(dG-dC)-poly(dG-dC) for measuring de novo methylation activity, and toward poly(dI-dC)-poly(dI-dC) for measuring total activity. This indicates that the enzymes are de novo-type DNA methyltransferases. The enzyme activity was inhibited by NaCl or KCl at concentrations >100 mM. The kinetic parameter, K(m)(AdoMet), for Dnmt3a, Dnmt3b1 and Dnmt3b2 was 0.4, 1.2 and 0.9 microM when poly(dI-dC)-poly(dI-dC) was used, and 0.3, 1.2 and 0.8 microM when poly(dG-dC)-poly(dG-dC) was used, respectively. The K(m)(DNA) values for Dnmt3a, Dnmt3b1 and Dnmt3b2 were 2.7, 1.3 and 1.5 microM when poly(dI-dC)-poly(dI-dC) was used, and 3.5, 1.0 and 0.9 microM when poly(dG-dC)-poly(dG-dC) was used, respectively. For the methylation specificity, Dnmt3a significantly methylated CpG >> CpA. On the other hand, Dnmt3b1 methylated CpG > CpT >/= CpA. Immuno-purified Dnmt3a, Myc-tagged and overexpressed in HEK 293T cells, methylated CpG >> CpA > CpT. Neither Dnmt3a nor Dnmt3b1 methylated the first cytosine of CpC.
Figures
Figure 1
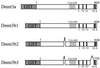
Schematic illustration of Dnmt3a, Dnnmt3b1, Dnmt3b2 and Dnmt3b3. The regions of the conserved motifs in the C-terminal catalytic domain (I∼X) and the Cys-rich region (Cys-rich) are indicated. Dnmt3b2 lacks amino acid residues 363–382, and Dnmt3b3 lacks amino acid residues 363–382 and 731–813, of Dnmt3b1. GST in the N-terminus of each Dnmt3 indicates the fused GST moiety. The numbers above the bars are amino acid numbers.
Figure 2
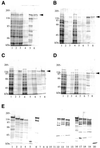
Purification of GST-fused Dnmt3s. An aliquot of a sample obtained at each purification step was subjected to SDS–PAGE and then stained with CBB. (A) Solubilized (lane 1), DEAE–Sepharose unbound (lane 2), glutathione–Sepharose unbound (lane 3), washed (lane 4), eluted (lane 5) and Superdex 200 eluted (lane 6) fractions of GST-fused Dnmt3a. 1/20 000 (lanes 1–4), 1/1000 (lane 5) and 1/100 (lane 6) of the recovered fractions were loaded. (B–D) Solubilized (lane 1), ammonium sulfate-precipitated with 20% saturation (lane 2), 20–30%-saturation (lane 3), supernatant on 30% saturation (lane 4), glutathione–Sepharose unbound (lane 5), washed (lane 6), eluted (lane 7) and Superdex 200 eluted (lane 8) fractions of GST-fused Dnmt3b1 (B), Dnmt3b2 (C) and Dnmt3b3 (D). 1/20 000 (lanes 1–5), 1/1000 (lanes 6 and 7) and 1/100 (lane 8) of the recovered fractions were loaded. Arrowheads indicate the position of Dnmt3. (E) Purified Dnmt3a (lane 1), Dnmt3b1 (lane 2), Dnmt3b2 (lane 3), Dnmt3b3 (lane 4) and GST (lane 5) were electrophoresed in the same gel and then CBB-stained. Purified Dnmt3a (lanes 6, 11 and 16), Dnmt3b1 (lanes 7, 12 and 17), Dnmt3b2 (lanes 8, 13 and 18), Dnmt3b3 (lanes 9, 14 and 19) and GST (lanes 10, 15 and 20) were immunodetected with specific antibodies reactive with Dnmt3a (lanes 6–10), Dnmt3b (lanes 11–15) and GST (lanes 16–20), respectively. Molecular weight sizes (kDa) are indicated beside each gel.
Figure 3
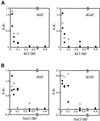
Effects of salts on the DNA methylation activity of Dnmt3s. The effects of KCl (A) and NaCl (B) on the DNA methylation activities [specific activity (s.a.) in mol/h/mol Dnmt3] of Dnmt3a (circles), Dnmt3b1 (open squares) and Dnmt3b2 (filled squares) was determined for methyl acceptors dIdC (left) and dGdC (right).
Figure 4
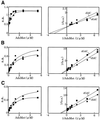
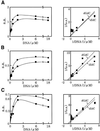
Effect of the AdoMet concentration on the activity of Dnmt3s. The DNA methylation activities [specific activity (s.a.) in mol/h/mol Dnmt3] of GST-fused Dnmt3a (A), Dnmt3b1 (B) and Dnmt3b2 (C) were titrated with AdoMet (left). As methyl acceptors, dIdC (circles) and dGdC (squares) were used. A typical example for each enzyme is demonstrated. Right panels show double reciprocal plots of the respective titration curves.
Figure 5
Effect of the DNA concentration on the activity of Dnmt3s. The DNA methylation activities [specific activity (s.a.) in mol/h/mol Dnmt3] of GST-fused Dnmt3a (A), Dnmt3b1 (B) and Dnmt3b2 (C) were titrated with dIdC (circles) or dGdC (squares). One mole double-stranded DNA with 1 CpI or CpG site was calculated to be 2 mol. A typical example for each enzyme is demonstrated. Right panels show double reciprocal plot of the respective titration curves.
Figure 6
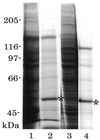
Expression and immuno-purification of Dnmt3s. Myc-tagged Dnmt3a (lanes 1 and 2) and Dnmt3b1 (lanes 3 and 4) were expressed in HEK 293T cells, extracted (lanes 1 and 3), and immunoprecipitated with anti-Myc monoclonal antibody (9E10)-coupled Sepharose (lanes 2 and 4). Asterisks indicate the heavy chain of 9E10. Molecular weight sizes (kDa) are indicated beside the gel.
References
- Antequera F. and Bird,A. (1993) CpG islands in DNA methylation. In Jost,J.P. and Saluz,H.P. (eds), Molecular Biology and Biological Significance. Birkhäuser Verlag, Basel, pp. 169–185.
- Tajima S. and Suetake,I. (1998) Regulation and function of DNA methylation in vertebrates. J. Biochem., 123, 993–999. -PubMed
- Jaenisch R. (1997) DNA methylation and imprinting: Why bother? Trends Genet., 13, 323–329. -PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous