p53 Binding protein 53BP1 is required for DNA damage responses and tumor suppression in mice - PubMed (original) (raw)
p53 Binding protein 53BP1 is required for DNA damage responses and tumor suppression in mice
Irene M Ward et al. Mol Cell Biol. 2003 Apr.
Abstract
53BP1 is a p53 binding protein of unknown function that binds to the central DNA-binding domain of p53. It relocates to the sites of DNA strand breaks in response to DNA damage and is a putative substrate of the ataxia telangiectasia-mutated (ATM) kinase. To study the biological role of 53BP1, we disrupted the 53BP1 gene in the mouse. We show that, similar to ATM(-/-) mice, 53BP1-deficient mice were growth retarded, immune deficient, radiation sensitive, and cancer prone. 53BP1(-/-) cells show a slight S-phase checkpoint defect and prolonged G(2)/M arrest after treatment with ionizing radiation. Moreover, 53BP1(-/-) cells feature a defective DNA damage response with impaired Chk2 activation. These data indicate that 53BP1 acts downstream of ATM and upstream of Chk2 in the DNA damage response pathway and is involved in tumor suppression.
Figures
FIG. 1.
Targeted disruption of the mouse 53BP1 gene. (A) Map of the genomic locus surrounding the targeted exon, the targeting vector containing the PGK-neo cassette, and the targeted locus. A 5′-flanking probe used for screening ES cell clones and mice is indicated. (B) Southern blot analysis of _Eco_RI-digested genomic DNA. (C) Multiplex PCR genotype analysis with a primer pair for the neo gene (resulting in a 455-bp product) and a 5′ external exon (resulting in a 270-bp product). (D) Western blot of cell extracts from mouse testes with an antibody specific for the N terminus of 53BP1. (E) Immunofluorescence analysis of irradiated (1 Gy) 53BP1+/+ and 53BP1−/− MEFs with polyclonal antibodies raised against the N terminus of 53BP1.
FIG. 2.
53BP1 deficiency results in growth retardation, cell cycle defect, radiosensitivity, and chromosomal instability. (A) Body weights of female 53BP1 wild-type, heterozygous, and knockout mice at 5 months of age. (B) G2/M arrest of 53BP1+/+ and 53BP1−/− embryonic cells in response to 0.5 or 5 Gy of IR. Cells were stained with anti-P-Histone 3 antibody 1 h after IR and analyzed by fluorescence-activated cell sorting. (C) G2 accumulation of 53BP1+/+ and 53BP1−/− MEFs several hours after IR. Cells were pulse-labeled for 1 h with BrdU before exposure to 6 Gy of IR. The cell cycle profile of BrdU-positive cells was analyzed by staining with propidium iodide. Consistent data were obtained in three independent experiments. (D) IR-induced intra-S-phase checkpoint in 53BP1+/+ and 53BP1−/− MEFs. DNA synthesis was assessed by [3H]thymidine incorporation 30 min after exposure to 20 Gy of IR. (E) Sensitivity of 10 pairs of female 53BP1−/− and 53BP1+/+ littermates to 8 Gy of whole body IR. Similar results were observed with 10 male pairs. (F) Chromosomal instability in 53BP1−/− MEFs. 100 metaphase spreads from genetically matched passage three 53BP1+/+ and 53BP1−/− MEFs were analyzed. Consistent data were obtained from three different experiments.
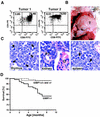
FIG. 3.
53BP1−/− mice are tumor prone. (A) CD4 and CD8 cell surface expression of cells from two different thymic lymphomas was assessed by flow cytometry. (B) Massive thymic lymphoma (T) in a 4-month-old 53BP1−/− mouse. (C) Hematoxylin-and-eosin-stained sections from the thymus, spleen, and kidney of a 53BP1−/− animal with thymic lymphoma. Monomorphic lymphoblastic tumor cells are dominant in all three tissues. The arrows indicate mitotic figures. G, glomerulus; PT, proximal tubulus. (D) Overall survival of 53BP1+/+ (n = 54), 53BP1+/− (n = 97), and 53BP1−/− (n = 101) mice over a period of 10 months.
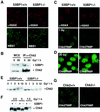
FIG. 4.
Irradiated 53BP−/− MEFs show impaired Chk2 activation after low doses of IR. (A) γ-H2AX and NBS1 focus formation 6 h after exposure to 6 Gy of IR is unaffected in 53BP1−/−MEFs compared to 53BP1 wild-type MEFs, as assessed by immunofluorescence staining. (B) Coimmunoprecipitation of 53BP1 and Chk2 in untreated or irradiated 293T cells. (C) Activated Chk2 phosphorylated at Thr 68 (Chk2T68) forms foci in 53BP1+/+ MEFs 1 h after exposure to 1 Gy of IR. No foci are detectable in 53BP1−/− MEFs at this low dose of IR. γ-H2AX staining is shown as a control. (D) Chk2T68P foci form in both 53BP1+/+ and 53BP1−/− cells in response to 20 Gy of IR. (E) Chk2 mobility shift is reduced in 53BP1−/− MEFs in response to low-dose radiation. Cell lysates from 53BP1+/+ and 53BP1−/− MEFs were prepared 1 h after IR and immunoblotted with anti-Chk2 antibody. (F) For better comparison of the Chk2 mobility shift, lysates from irradiated 53BP1+/+ and 53BP1−/− cells were run side by side. (G) The guinea pig anti-Chk2T68P antibody specifically recognizes Chk2.
Similar articles
- Tumor suppressor p53 binding protein 1 (53BP1) is involved in DNA damage-signaling pathways.
Rappold I, Iwabuchi K, Date T, Chen J. Rappold I, et al. J Cell Biol. 2001 Apr 30;153(3):613-20. doi: 10.1083/jcb.153.3.613. J Cell Biol. 2001. PMID: 11331310 Free PMC article. - 53BP1 functions in an ATM-dependent checkpoint pathway that is constitutively activated in human cancer.
DiTullio RA Jr, Mochan TA, Venere M, Bartkova J, Sehested M, Bartek J, Halazonetis TD. DiTullio RA Jr, et al. Nat Cell Biol. 2002 Dec;4(12):998-1002. doi: 10.1038/ncb892. Nat Cell Biol. 2002. PMID: 12447382 - 53BP1 and NFBD1/MDC1-Nbs1 function in parallel interacting pathways activating ataxia-telangiectasia mutated (ATM) in response to DNA damage.
Mochan TA, Venere M, DiTullio RA Jr, Halazonetis TD. Mochan TA, et al. Cancer Res. 2003 Dec 15;63(24):8586-91. Cancer Res. 2003. PMID: 14695167 - ATM signaling and 53BP1.
Zgheib O, Huyen Y, DiTullio RA Jr, Snyder A, Venere M, Stavridi ES, Halazonetis TD. Zgheib O, et al. Radiother Oncol. 2005 Aug;76(2):119-22. doi: 10.1016/j.radonc.2005.06.026. Radiother Oncol. 2005. PMID: 16024119 Review. - ATM: the protein encoded by the gene mutated in the radiosensitive syndrome ataxia-telangiectasia.
Lavin MF, Khanna KK. Lavin MF, et al. Int J Radiat Biol. 1999 Oct;75(10):1201-14. doi: 10.1080/095530099139359. Int J Radiat Biol. 1999. PMID: 10549596 Review.
Cited by
- RNF4 is required for DNA double-strand break repair in vivo.
Vyas R, Kumar R, Clermont F, Helfricht A, Kalev P, Sotiropoulou P, Hendriks IA, Radaelli E, Hochepied T, Blanpain C, Sablina A, van Attikum H, Olsen JV, Jochemsen AG, Vertegaal AC, Marine JC. Vyas R, et al. Cell Death Differ. 2013 Mar;20(3):490-502. doi: 10.1038/cdd.2012.145. Epub 2012 Nov 30. Cell Death Differ. 2013. PMID: 23197296 Free PMC article. - A role for the p53 tumour suppressor in regulating the balance between homologous recombination and non-homologous end joining.
Moureau S, Luessing J, Harte EC, Voisin M, Lowndes NF. Moureau S, et al. Open Biol. 2016 Sep;6(9):160225. doi: 10.1098/rsob.160225. Open Biol. 2016. PMID: 27655732 Free PMC article. - BRCA1 loss activates cathepsin L-mediated degradation of 53BP1 in breast cancer cells.
Grotsky DA, Gonzalez-Suarez I, Novell A, Neumann MA, Yaddanapudi SC, Croke M, Martinez-Alonso M, Redwood AB, Ortega-Martinez S, Feng Z, Lerma E, Ramon y Cajal T, Zhang J, Matias-Guiu X, Dusso A, Gonzalo S. Grotsky DA, et al. J Cell Biol. 2013 Jan 21;200(2):187-202. doi: 10.1083/jcb.201204053. J Cell Biol. 2013. PMID: 23337117 Free PMC article. - Trolox contributes to Nrf2-mediated protection of human and murine primary alveolar type II cells from injury by cigarette smoke.
Messier EM, Bahmed K, Tuder RM, Chu HW, Bowler RP, Kosmider B. Messier EM, et al. Cell Death Dis. 2013 Apr 4;4(4):e573. doi: 10.1038/cddis.2013.96. Cell Death Dis. 2013. PMID: 23559007 Free PMC article. - MBTD1 is associated with Pr-Set7 to stabilize H4K20me1 in mouse oocyte meiotic maturation.
Luo YB, Ma JY, Zhang QH, Lin F, Wang ZW, Huang L, Schatten H, Sun QY. Luo YB, et al. Cell Cycle. 2013 Apr 1;12(7):1142-50. doi: 10.4161/cc.24216. Epub 2013 Mar 8. Cell Cycle. 2013. PMID: 23475131 Free PMC article.
References
- Abraham, R. T. 2001. Cell cycle checkpoint signaling through the ATM and ATR kinases. Genes Dev. 15:2177-2196. - PubMed
- Barlow, C., S. Hirotsune, R. Paylor, M. Liyanage, M. Eckhaus, F. Collins, Y. Shiloh, J. N. Crawley, T. Ried, D. Tagle, and A. Wynshaw-Boris. 1996. ATM-deficient mice: a paradigm of ataxia telangiectasia. Cell 86:159-171. - PubMed
- Bartek, J., J. Falck, and J. Lukas. 2001. CHK2 kinase: a busy messenger. Nat. Rev. Mol. Cell. Biol. 2:877-886. - PubMed
- Callebaut, I., and J. P. Mornon. 1997. From BRCA1 to RAP1: a widespread BRCT module closely associated with DNA repair. FEBS Lett. 400:25-30. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous