A bacterial one-hybrid system for determining the DNA-binding specificity of transcription factors - PubMed (original) (raw)
A bacterial one-hybrid system for determining the DNA-binding specificity of transcription factors
Xiangdong Meng et al. Nat Biotechnol. 2005 Aug.
Abstract
The DNA-binding specificities of transcription factors can be used to computationally predict cis-regulatory modules (CRMs) that regulate gene expression. However, the absence of specificity data for the majority of transcription factors limits the widespread implementation of this approach. We have developed a bacterial one-hybrid system that provides a simple and rapid method to determine the DNA-binding specificity of a transcription factor. Using this technology, we successfully determined the DNA-binding specificity of seven previously characterized transcription factors and one novel transcription factor, the Drosophila melanogaster factor Odd-skipped. Regulatory targets of Odd-skipped were successfully predicted using this information, demonstrating that the data produced by the bacterial one-hybrid system are relevant to in vivo function.
Figures
Figure 1
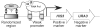
Overview of the bacterial one-hybrid system. a) Schematic representation of the HIS3/URA3 cistron in the pH3U3 prey vector. If the DNA-binding domain (DBD) of the bait recognizes a sequence in the randomized region, the fusion to the alpha subunit will recruit RNA polymerase, to the weak lac promoter and activate transcription of HIS3 and URA3. b) Schematic outline of the B1H selection procedure. The prey library and bait are introduced into the bacterial selection strain. These cells are plated on NM selective media containing the desired concentration of 3-AT to select for bait-prey combinations that activate the reporter. Subsequent steps depend on the prey library (either the original or purified version) that was used for the selection. If the original library is used then prey from the colonies that grow under the selective conditions are isolated and reintroduced into the selection strain in the absence of the bait. These cells are challenged to survive stringent counterselection (5-FOA) to remove any self-activating clones that represent false positives. DNA is isolated from individual colonies that grow under these conditions and the randomized region of each prey is sequenced. If the purified library was used for the selection, then DNA can be isolated and sequenced from colonies on the positive selection plate since the majority of self-activating prey have already been eliminated. The unique DNA sequences recovered from the selection are analyzed by MEME to identify any overrepresented sequence motifs, which should represent the recognition sequence of the bait. c) A purified library can be constructed to simplify the selection of bait-prey combinations that activate the reporter genes. The original prey library is introduced into the selection strain and cells are challenged to survive stringent counterselection conditions (5-FOA) to remove any self-activating clones that represent false positives. Prey vectors from the surviving colonies are isolated as a pool to generate the purified prey library for use with any desired bait.
Figure 1
Overview of the bacterial one-hybrid system. a) Schematic representation of the HIS3/URA3 cistron in the pH3U3 prey vector. If the DNA-binding domain (DBD) of the bait recognizes a sequence in the randomized region, the fusion to the alpha subunit will recruit RNA polymerase, to the weak lac promoter and activate transcription of HIS3 and URA3. b) Schematic outline of the B1H selection procedure. The prey library and bait are introduced into the bacterial selection strain. These cells are plated on NM selective media containing the desired concentration of 3-AT to select for bait-prey combinations that activate the reporter. Subsequent steps depend on the prey library (either the original or purified version) that was used for the selection. If the original library is used then prey from the colonies that grow under the selective conditions are isolated and reintroduced into the selection strain in the absence of the bait. These cells are challenged to survive stringent counterselection (5-FOA) to remove any self-activating clones that represent false positives. DNA is isolated from individual colonies that grow under these conditions and the randomized region of each prey is sequenced. If the purified library was used for the selection, then DNA can be isolated and sequenced from colonies on the positive selection plate since the majority of self-activating prey have already been eliminated. The unique DNA sequences recovered from the selection are analyzed by MEME to identify any overrepresented sequence motifs, which should represent the recognition sequence of the bait. c) A purified library can be constructed to simplify the selection of bait-prey combinations that activate the reporter genes. The original prey library is introduced into the selection strain and cells are challenged to survive stringent counterselection conditions (5-FOA) to remove any self-activating clones that represent false positives. Prey vectors from the surviving colonies are isolated as a pool to generate the purified prey library for use with any desired bait.
Figure 1
Overview of the bacterial one-hybrid system. a) Schematic representation of the HIS3/URA3 cistron in the pH3U3 prey vector. If the DNA-binding domain (DBD) of the bait recognizes a sequence in the randomized region, the fusion to the alpha subunit will recruit RNA polymerase, to the weak lac promoter and activate transcription of HIS3 and URA3. b) Schematic outline of the B1H selection procedure. The prey library and bait are introduced into the bacterial selection strain. These cells are plated on NM selective media containing the desired concentration of 3-AT to select for bait-prey combinations that activate the reporter. Subsequent steps depend on the prey library (either the original or purified version) that was used for the selection. If the original library is used then prey from the colonies that grow under the selective conditions are isolated and reintroduced into the selection strain in the absence of the bait. These cells are challenged to survive stringent counterselection (5-FOA) to remove any self-activating clones that represent false positives. DNA is isolated from individual colonies that grow under these conditions and the randomized region of each prey is sequenced. If the purified library was used for the selection, then DNA can be isolated and sequenced from colonies on the positive selection plate since the majority of self-activating prey have already been eliminated. The unique DNA sequences recovered from the selection are analyzed by MEME to identify any overrepresented sequence motifs, which should represent the recognition sequence of the bait. c) A purified library can be constructed to simplify the selection of bait-prey combinations that activate the reporter genes. The original prey library is introduced into the selection strain and cells are challenged to survive stringent counterselection conditions (5-FOA) to remove any self-activating clones that represent false positives. Prey vectors from the surviving colonies are isolated as a pool to generate the purified prey library for use with any desired bait.
Figure 2
Binding site motifs for seven proteins determined using the B1H system. The binding site motif determined using each bait in the B1H system is displayed as a Sequence logo. The maximum possible information content at each position is 2 bits. The Zif268, PLAG-1 and Runt/Bgb binding motifs were generated using the two-step selection method, whereas motifs for the other factors were generated using the purified library in a single selection step. The previously reported consensus sequence for each factor as determined by SELEX is displayed or listed below each Logo. a) Comparison of the binding site motifs produced for Zif268 by the B1H system and from a previously reported in vitro SELEX experiment. b) PLAG1 can tolerate a 0 to 4 base pair gap between the two recognition motifs based on analysis of the raw sequences by Bioprospector. A one base pair gap (position 8) is shown in its motif, but a 2 base pair gap was most prevalent in the isolated sequences (7 of 18). The PLAG1 motif is consistent with the previously described PLAG1 consensus sequence (GG(g/a)GGCCNNNNNN(g/a)GG(g/t)) determined by SELEX. The most 5' base identified in the SELEX analysis could not be conclusively defined in the B1H data because of overlap of the majority of sequences with the edge of the constant region abutting the library. c) The ZnFp53 motif is consistent with the previously described consensus sequence ((g/a)GACACGT) determined by SELEX for a nearly identical clone. d) The Dorsal motif is consistent with the previously described consensus sequence (GGG(a/t)(a/t)(t/a)(t/c/a)C(c/t)) determined by SELEX. e) The LAG-1 motif is consistent with the previously described consensus sequence (A(c/g)CGTGGGAA(a/c)) for the mouse homolog of LAG-1 (RBP-Jκ) determined by SELEX. f) The Paired motif contains at its core a sequence similar to the consensus sequence ((t/c)CGTCACG(g/c)TT(g/c)) determined by SELEX for the paired domain in the absence of the homeodomain. SELEX selections reported for both DNA-binding domains of Paired resulted a complex mixture of recognition sequences, a subset contained a core homeodomain binding site abutting the 5' end of the paired domain binding site (
AATTA
GTCACGC; where the homeodomain element is underlined), which is similar to the 5' end of our motif. g) The Runt/Bgb motif is consistent with the previously described consensus sequence ((t/c)G(t/c)GGT(t/c)) for CBFα/β determined by SELEX. Raw sequences of the prey from each selection are listed in Supplementary Table 2.
Figure 3
Analysis of the DNA-binding specificity of Odd determined using the B1H system. a) The binding site motifs obtained at 1.5, 2.5 and 5 mM 3-AT were each compiled from more than 20 sequences that contained an overrepresented sequence motif identified by MEME. The tolerance of Odd for both A and T at position 4 (boxed) in its binding site becomes apparent at lower selection stringencies. Raw sequences of the prey from each selection are listed in Supplementary Table 2. b) Effect of point mutations on the recognition by Odd of its consensus sequence. i) Gel shift assay examining the effect of different DNA competitors on formation of a complex between Odd and its labeled consensus binding site. The presence of Odd and the type of cold competitor in each binding reaction is indicated above each lane of the gel. wt = contains the consensus Odd sequence GCTACTGTA. The other competitors have mutations at each of the first five positions of the consensus sequence where the number represents the position and the letter represents the substitution. For example, 1c = cCTACTGTA. ii) Growth rates for bacteria assayed at 2 mM 3-AT containing the Odd bait and either the wild type or a mutant prey corresponding to DNA competitors used in the gel shift analysis. Growth rates were defined for each bait-prey combination based on serial dilutions of cells harboring these vectors on plates containing 3-AT (Supplementary Fig. 3). The growth rates for cells containing the various bait-prey combinations are qualitatively similar to the degree of competition observed in the gel shift experiments.
Figure 3
Analysis of the DNA-binding specificity of Odd determined using the B1H system. a) The binding site motifs obtained at 1.5, 2.5 and 5 mM 3-AT were each compiled from more than 20 sequences that contained an overrepresented sequence motif identified by MEME. The tolerance of Odd for both A and T at position 4 (boxed) in its binding site becomes apparent at lower selection stringencies. Raw sequences of the prey from each selection are listed in Supplementary Table 2. b) Effect of point mutations on the recognition by Odd of its consensus sequence. i) Gel shift assay examining the effect of different DNA competitors on formation of a complex between Odd and its labeled consensus binding site. The presence of Odd and the type of cold competitor in each binding reaction is indicated above each lane of the gel. wt = contains the consensus Odd sequence GCTACTGTA. The other competitors have mutations at each of the first five positions of the consensus sequence where the number represents the position and the letter represents the substitution. For example, 1c = cCTACTGTA. ii) Growth rates for bacteria assayed at 2 mM 3-AT containing the Odd bait and either the wild type or a mutant prey corresponding to DNA competitors used in the gel shift analysis. Growth rates were defined for each bait-prey combination based on serial dilutions of cells harboring these vectors on plates containing 3-AT (Supplementary Fig. 3). The growth rates for cells containing the various bait-prey combinations are qualitatively similar to the degree of competition observed in the gel shift experiments.
Figure 4
Altered gene expression following ectopic expression of Drosophila Odd. Probes used for RNA in situ hybridization are indicated to the left of each pair of panels. (upper panels) Wild type embryos showing the head and trunk stripes of odd expression at the pair rule and segment polarity stages. (lower panels) Expression of putative targets of the Odd repressor was examined in wild type (left) or heat-shock odd embryos (right) that were fixed 19 minutes following a six minute heat shock. In heat-shock odd embryos: stripe 1 of h disappears (arrow); the even stripes of prd are missing (bracket); the segment polarity stripes of gsb are missing or reduced (bracket); and the head stripe of Gsc is fainter (arrow). For analysis of Gsc, the embryos were co-stained with a probe to Ubx (central stripe), which did not significantly change following ectopic expression of odd. The changes observed in h and prd expression following ectopic expression of odd have been previously described.
Similar articles
- Discovering DNA regulatory elements with bacteria.
Bulyk ML. Bulyk ML. Nat Biotechnol. 2005 Aug;23(8):942-4. doi: 10.1038/nbt0805-942. Nat Biotechnol. 2005. PMID: 16082362 Free PMC article. No abstract available. - A systematic characterization of factors that regulate Drosophila segmentation via a bacterial one-hybrid system.
Noyes MB, Meng X, Wakabayashi A, Sinha S, Brodsky MH, Wolfe SA. Noyes MB, et al. Nucleic Acids Res. 2008 May;36(8):2547-60. doi: 10.1093/nar/gkn048. Epub 2008 Mar 10. Nucleic Acids Res. 2008. PMID: 18332042 Free PMC article. - [Recruiting of insulator protein ZIPIC of Drosophila melanogaster to minor binding sites in vivo depends on other DNA-binding transcription factors].
Zolotarev NA, Kyrchanova OV, Maksimenko OG, Georgiev PG. Zolotarev NA, et al. Mol Biol (Mosk). 2015 Nov-Dec;49(6):1016-21. doi: 10.7868/S0026898415060245. Mol Biol (Mosk). 2015. PMID: 26710784 Russian. - Understanding DNA-binding specificity by bacteria hybrid selection.
Xu DJ, Noyes MB. Xu DJ, et al. Brief Funct Genomics. 2015 Jan;14(1):3-16. doi: 10.1093/bfgp/elu048. Epub 2014 Dec 23. Brief Funct Genomics. 2015. PMID: 25539837 Review. - Architectural transcription factors: proteins that remodel DNA.
Werner MH, Burley SK. Werner MH, et al. Cell. 1997 Mar 21;88(6):733-6. doi: 10.1016/s0092-8674(00)81917-0. Cell. 1997. PMID: 9118214 Review. No abstract available.
Cited by
- A Biosensor Strategy for E. coli Based on Ligand-Dependent Stabilization.
Brandsen BM, Mattheisen JM, Noel T, Fields S. Brandsen BM, et al. ACS Synth Biol. 2018 Sep 21;7(9):1990-1999. doi: 10.1021/acssynbio.8b00052. Epub 2018 Aug 14. ACS Synth Biol. 2018. PMID: 30064218 Free PMC article. - Prediction of Transcription Factor Regulators and Gene Regulatory Networks in Tomato Using Binding Site Information.
Manosalva Pérez N, Vandepoele K. Manosalva Pérez N, et al. Methods Mol Biol. 2023;2698:323-349. doi: 10.1007/978-1-0716-3354-0_19. Methods Mol Biol. 2023. PMID: 37682483 - Accurate and sensitive quantification of protein-DNA binding affinity.
Rastogi C, Rube HT, Kribelbauer JF, Crocker J, Loker RE, Martini GD, Laptenko O, Freed-Pastor WA, Prives C, Stern DL, Mann RS, Bussemaker HJ. Rastogi C, et al. Proc Natl Acad Sci U S A. 2018 Apr 17;115(16):E3692-E3701. doi: 10.1073/pnas.1714376115. Epub 2018 Apr 2. Proc Natl Acad Sci U S A. 2018. PMID: 29610332 Free PMC article. - Integrated microfluidic approach for quantitative high-throughput measurements of transcription factor binding affinities.
Glick Y, Orenstein Y, Chen D, Avrahami D, Zor T, Shamir R, Gerber D. Glick Y, et al. Nucleic Acids Res. 2016 Apr 7;44(6):e51. doi: 10.1093/nar/gkv1327. Epub 2015 Dec 3. Nucleic Acids Res. 2016. PMID: 26635393 Free PMC article. - Using protein design algorithms to understand the molecular basis of disease caused by protein-DNA interactions: the Pax6 example.
Alibés A, Nadra AD, De Masi F, Bulyk ML, Serrano L, Stricher F. Alibés A, et al. Nucleic Acids Res. 2010 Nov;38(21):7422-31. doi: 10.1093/nar/gkq683. Epub 2010 Aug 4. Nucleic Acids Res. 2010. PMID: 20685816 Free PMC article.
References
- Roulet E, et al. High-throughput SELEX SAGE method for quantitative modeling of transcription-factor binding sites. Nat Biotechnol. 2002;20:831–835. - PubMed
- Lee TI, et al. Transcriptional regulatory networks in Saccharomyces cerevisiae. Science. 2002;298:799–804. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials