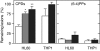Impaired nucleotide excision repair upon macrophage differentiation is corrected by E1 ubiquitin-activating enzyme - PubMed (original) (raw)
Impaired nucleotide excision repair upon macrophage differentiation is corrected by E1 ubiquitin-activating enzyme
Thierry Nouspikel et al. Proc Natl Acad Sci U S A. 2006.
Abstract
Global nucleotide excision repair is greatly attenuated in terminally differentiated mammalian cells. We observed this phenomenon in human neurons and in macrophages, noting that the transcription-coupled repair pathway remains functional and that there is no significant reduction in levels of excision repair enzymes. We have discovered that ubiquitin-activating enzyme E1 complements the repair deficiency in macrophage extracts, and although there is no reduction in the concentration of E1 upon differentiation, our results indicate a reduction in phosphorylation of E1. In preliminary studies, we have identified the basal transcription factor TFIIH as the potential target for ubiquitination. We suggest that this unusual type of regulation at the level of the E1 enzyme is likely to affect numerous cellular processes and may represent a strategy to coordinate multiple phenotypic changes upon differentiation by using E1 as a "master switch."
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Fig. 1.
In vivo repair of UV-induced lesions. HL60 and THP1 cells, naïve (white bars) or differentiated with TPA for 16 h (gray bars) or 48 h (black bars), were irradiated with a dose (10 J/m2) of 254-nm UV light. Cells were harvested either immediately or 24 h later, and DNA was purified, blotted onto a nitrocellulose membrane, and probed with antibodies specific for CPDs (Left) or (6-4)PPs (Right). Results are expressed as percentage of the lesions measured at T0 that remained 24 h after irradiation. Error bars are the SEM of two to four experiments. ∗, P < 0.05; ∗∗, P < 0.01 (Student's t test). More detailed time-course experiments can be found in refs. and .
Fig. 2.
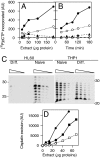
In vitro repair of UV-induced lesions and cisplatin cross-links. (A and B) Two plasmids of different sizes, irradiated or not with 254-nm UV light, were incubated with “Manley” extract from naïve (filled symbols) or differentiated (open symbols) HL60 cells, in the presence of [32P]dCTP. The damaged (circles) and undamaged (triangles) plasmids were then resolved by agarose-gel electrophoresis, and the amount of radioactive precursor incorporated into the DNA was quantified with a PhosphorImager. (C and D) A plasmid carrying a single cisplatin GTG intrastrand cross-link close to a 32P label was incubated with Manley extracts from HL60 (circles) or THP1 (squares) cells, either naïve (filled symbols) or differentiated with TPA for 48 h (open symbols). Excision products spanning the lesion, of 26–35 nucleotides in length, were resolved by acrylamide-gel electrophoresis and quantified with a PhosphorImager. Similar experiments were performed with 4–12 distinct preparations of cell extracts.
Fig. 3.
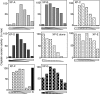
In vitro complementation of differentiated macrophage extracts with XP extracts. Extracts from differentiated HL60 cells and NER-deficient extracts from the seven complementation groups of XP were mixed in various proportions and incubated with a plasmid containing a single cisplatin cross-link as described for Fig. 2_C_. In each panel, the amount of XP extract decreases from left to right as the amount of macrophage extract increases, keeping the total amount of proteins constant. A mix resulting in proficient excision of the cisplatin lesion indicates that differentiated HL60 cells do not lack the corresponding XP enzyme. Because XP-E extracts have an intrinsic activity, they were assayed with and without macrophage extract. N, naïve HL60 extract, included for comparison.
Fig. 4.
Purification of an activity complementing differentiated macrophage extracts. (A) Overall purification scheme. (B) Elution of 15 ml of Manley extract on a DEAE-Sepharose column in a 0–400 mM sodium chloride gradient, pH 6.3 (diagonal line). Solid line, protein concentration; bars, cisplatin excision activity of differentiated THP1 extracts complemented with a portion of the relevant fractions (the background activity in differentiated THP1 extracts was subtracted to better visualize the active fractions). (C) Further purification of the active fractions on a butyl-Sepharose column in a 640–0 mM ammonium sulfate gradient (diagonal line). The activity of the resulting fractions was measured as above (bars). (D) Further purification of the active fractions with a sephacryl S-200 gel filtration column, analyzed by SDS/PAGE and silver staining. Lanes denote: D, DEAE fraction #21; B, butyl fraction #13; 9–14, S-200 fractions number 9 through 14, which contained the bulk of the complementing activity.
Fig. 5.
Confirmation of the involvement of E1. (A and B) Various amounts of ubiquitin-activating enzyme E1 purified from rabbit (triangles) or recombinant His-tagged human E1 (circles) were mixed with differentiated THP1 extract and assayed for cisplatin excision. The activity of a repair-proficient HeLa extract mixed with E1 buffer is shown for comparison (diamond). (C and D) Cisplatin excision was measured with various amounts of extracts from ts85 cells, a thermolabile E1 mutant, grown at 33°C and exposed (open triangles) or not (filled triangles) at 39°C for 16 h. The parental cell line FM3A was similarly exposed to 39°C (squares) to exclude a nonspecific effect of the temperature. Average of three experiments is shown. Error bars are SD. (E) Complementation of differentiated THP1 extract with purified TFIIH. Average of two experiments is shown. Error bars are SEM.
Fig. 6.
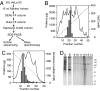
Differences in E1 phosphorylation. (A) Western blot of various batches of HL60 and THP1 extracts, naïve or differentiated, probed with an anti-E1 polyclonal antibody. (B) One microgram of human recombinant E1 was preincubated for 30 min with or without 0.6 mU of potato acid phosphatase before being mixed with THP1-differentiated cell extract and assayed for cisplatin excision. Together: E1 was preincubated alone, and the phosphatase was added together with the extract. (C) E1 was immunoprecipitated from naïve (Upper) or differentiated (Lower) THP1 cells, submitted to 2D gel electrophoresis and analyzed by Western blotting with antiphosphoserine antibodies. Numbers 0 through 4 indicate the number of phosphates on E1. (D) Quantification of the above, together with a similar experiment. Results are displayed as changes between naïve and differentiated extracts in the contribution of the various phosphorylated forms to the total amount of E1.
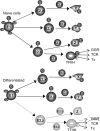
Fig. 7.
Model of NER control by E1 phosphorylation. In naïve cells, E1 is phosphorylated (P) on four sites, leading to ubiquitination (Ub) of an NER enzyme, most likely TFIIH, among others proteins (X, Y, Z). E1 phosphorylation is decreased in differentiated cells, which impairs its association with the E2 enzyme required to ubiquitinate TFIIH (E2.z). A lack in TFIIH ubiquitination decreases its activity in global genomic repair (GGR), but not in transcription-coupled repair (TCR), or in mediating RNA PolII transcription (Tx). The model predicts that ubiquitination of other proteins (e.g., Z) may also be deficient if it relies on the same E2, or on an E2 affected by the same phosphorylation site in E1.
Similar articles
- Cellular ubiquitination and proteasomal functions positively modulate mammalian nucleotide excision repair.
Wang QE, Wani MA, Chen J, Zhu Q, Wani G, El-Mahdy MA, Wani AA. Wang QE, et al. Mol Carcinog. 2005 Jan;42(1):53-64. doi: 10.1002/mc.20065. Mol Carcinog. 2005. PMID: 15547920 - DNA repair in differentiated cells: some new answers to old questions.
Nouspikel T. Nouspikel T. Neuroscience. 2007 Apr 14;145(4):1213-21. doi: 10.1016/j.neuroscience.2006.07.006. Epub 2006 Aug 22. Neuroscience. 2007. PMID: 16920273 Review. - XPB and XPD helicases in TFIIH orchestrate DNA duplex opening and damage verification to coordinate repair with transcription and cell cycle via CAK kinase.
Fuss JO, Tainer JA. Fuss JO, et al. DNA Repair (Amst). 2011 Jul 15;10(7):697-713. doi: 10.1016/j.dnarep.2011.04.028. Epub 2011 May 14. DNA Repair (Amst). 2011. PMID: 21571596 Free PMC article. Review. - Mutations in Cockayne Syndrome-Associated Genes (Csa and Csb) Predispose to Cisplatin-Induced Hearing Loss in Mice.
Rainey RN, Ng SY, Llamas J, van der Horst GT, Segil N. Rainey RN, et al. J Neurosci. 2016 Apr 27;36(17):4758-70. doi: 10.1523/JNEUROSCI.3890-15.2016. J Neurosci. 2016. PMID: 27122034 Free PMC article. - Comparative study of nucleotide excision repair defects between XPD-mutated fibroblasts derived from trichothiodystrophy and xeroderma pigmentosum patients.
Nishiwaki T, Kobayashi N, Iwamoto T, Yamamoto A, Sugiura S, Liu YC, Sarasin A, Okahashi Y, Hirano M, Ueno S, Mori T. Nishiwaki T, et al. DNA Repair (Amst). 2008 Dec 1;7(12):1990-8. doi: 10.1016/j.dnarep.2008.08.009. Epub 2008 Oct 10. DNA Repair (Amst). 2008. PMID: 18817897
Cited by
- HOXC9 directly regulates distinct sets of genes to coordinate diverse cellular processes during neuronal differentiation.
Wang X, Choi JH, Ding J, Yang L, Ngoka LC, Lee EJ, Zha Y, Mao L, Jin B, Ren M, Cowell J, Huang S, Shi H, Cui H, Ding HF. Wang X, et al. BMC Genomics. 2013 Nov 25;14:830. doi: 10.1186/1471-2164-14-830. BMC Genomics. 2013. PMID: 24274069 Free PMC article. - Age-related changes in the proteostasis network in the brain of the naked mole-rat: Implications promoting healthy longevity.
Triplett JC, Tramutola A, Swomley A, Kirk J, Grimes K, Lewis K, Orr M, Rodriguez K, Cai J, Klein JB, Perluigi M, Buffenstein R, Butterfield DA. Triplett JC, et al. Biochim Biophys Acta. 2015 Oct;1852(10 Pt A):2213-24. doi: 10.1016/j.bbadis.2015.08.002. Epub 2015 Aug 4. Biochim Biophys Acta. 2015. PMID: 26248058 Free PMC article. - Balancing self-renewal against genome preservation in stem cells: How do they manage to have the cake and eat it too?
Tsai RY. Tsai RY. Cell Mol Life Sci. 2016 May;73(9):1803-23. doi: 10.1007/s00018-016-2152-y. Epub 2016 Feb 17. Cell Mol Life Sci. 2016. PMID: 26886024 Free PMC article. Review. - Terminally differentiated muscle cells are defective in base excision DNA repair and hypersensitive to oxygen injury.
Narciso L, Fortini P, Pajalunga D, Franchitto A, Liu P, Degan P, Frechet M, Demple B, Crescenzi M, Dogliotti E. Narciso L, et al. Proc Natl Acad Sci U S A. 2007 Oct 23;104(43):17010-5. doi: 10.1073/pnas.0701743104. Epub 2007 Oct 16. Proc Natl Acad Sci U S A. 2007. PMID: 17940040 Free PMC article. - A proteomic analysis of C-reactive protein stimulated THP-1 monocytes.
Eisenhardt SU, Habersberger J, Oliva K, Lancaster GI, Ayhan M, Woollard KJ, Bannasch H, Rice GE, Peter K. Eisenhardt SU, et al. Proteome Sci. 2011 Jan 10;9(1):1. doi: 10.1186/1477-5956-9-1. Proteome Sci. 2011. PMID: 21219634 Free PMC article.
References
- Nouspikel T, Hanawalt PC. DNA Repair. 2001;1:59–75. - PubMed
- Hahn GM, King D, Yang SJ. Nat New Biol. 1971;230:242–244. - PubMed
- Ho L, Hanawalt PC. Mutat Res DNA Repair. 1991;255:124–141. - PubMed
- Tofilon PJ, Meyn RE. Radiat Res. 1988;116:217–237. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials