The RNA helicase p68 is a novel androgen receptor coactivator involved in splicing and is overexpressed in prostate cancer - PubMed (original) (raw)
. 2008 Oct 1;68(19):7938-46.
doi: 10.1158/0008-5472.CAN-08-0932.
Anne Coulson, Caroline Dalgliesh, Prabhakar Rajan, Samantha M Nicol, Stewart Fleming, Rakesh Heer, Luke Gaughan, Hing Y Leung, David J Elliott, Frances V Fuller-Pace, Craig N Robson
Affiliations
- PMID: 18829551
- PMCID: PMC2561211
- DOI: 10.1158/0008-5472.CAN-08-0932
The RNA helicase p68 is a novel androgen receptor coactivator involved in splicing and is overexpressed in prostate cancer
Emma L Clark et al. Cancer Res. 2008.
Abstract
The androgen receptor (AR) is a member of the nuclear steroid hormone receptor family and is thought to play an important role in the development of both androgen-dependent and androgen-independent prostatic malignancy. Elucidating roles by which cofactors regulate AR transcriptional activity may provide therapeutic advancement for prostate cancer (PCa). The DEAD box RNA helicase p68 (Ddx5) was identified as a novel AR-interacting protein by yeast two-hybrid screening, and we sought to examine the involvement of p68 in AR signaling and PCa. The p68-AR interaction was verified by colocalization of overexpressed protein by immunofluorescence and confirmed in vivo by coimmunoprecipitation in the PCa LNCaP cell line. Chromatin immunoprecipitation in the same cell line showed AR and p68 recruitment to the promoter region of the androgen-responsive prostate-specific antigen (PSA) gene. Luciferase reporter, minigene splicing assays, and RNA interference (RNAi) were used to examine a functional role of p68 in AR-regulated gene expression, whereby p68 targeted RNAi reduced AR-regulated PSA expression, and p68 enhanced AR-regulated repression of CD44 splicing (P = 0.008). Tyrosine phosphorylation of p68 was found to enhance coactivation of ligand-dependent transcription of AR-regulated luciferase reporters independent of ATP-binding. Finally, we observe increased frequency and expression of p68 in PCa compared with benign tissue using a comprehensive prostate tissue microarray (P = 0.003; P = 0.008). These findings implicate p68 as a novel AR transcriptional coactivator that is significantly overexpressed in PCa with a possible role in progression to hormone-refractory disease.
Figures
Figure 1
p68 is a nuclear protein and interacts with the AR. A. Cropped immunoblot images of p68 in nuclear (N) and cytosolic (C) cell lysates obtained from LNCaP, COS-7 and HEK 293 cells probed sequentially with p68, TATA Binding Protein (TBP) and α-tubulin antibody. B. Immunofluorescence images of HEK 293 cells transfected with ARWT-GFP or p68WT-YFP constructs in the presence of 10 nM R1881. Cells were nuclei stained with DAPI prior to confocal microscope imaging. C. Cropped immunoblot images of LNCaP nuclear lysates co-immunoprecipitated (IP) with either AR rabbit or p68 goat polyclonal antibody (+/- R1881 10nM), probed sequentially with p68 and AR mouse monoclonal antibodies. p68 co-immunoprecipitated AR +/- RNase treatment (100μg/ml), was in the presence of R1881 10nM. Control (Con) lanes contain LNCaP nuclear lysate and Protein G Sepharose (PGS) only.
Figure 2
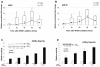
p68 co-occupies the active PSA promoter at ARE regions and enhances AR transcriptional activity. QPCR analysis (n=3) of ChIP assays (n=3) normalised to GAPDH levels (+/- SE) demonstrate p68 (bar) and AR (line) recruitment to A. ARE I promoter and B. ARE III enhancer regions of the PSA gene over a time course of 100 mins after 10 nM R1881 treatment. COS-7 cells were co-transiently transfected in triplicate with 0.1 μg of p(ARE)3 luciferase reporter, 0.1 μg pcDNA3-AR, 0.1 μg of pCMV-β-gal and increasing concentrations of C. pcDNA3-p68WT and D. pcDNA3-p68GLT constructs (+/- 10nM R1881). Luciferase activity was corrected for the corresponding β-gal activity to give a relative activity. The range of plasmid levels (+, ++, +++ and ++++) corresponds to 25, 50, 75 and 100ng respectively. Data shows relative activity from at least three separate luciferase assay experiments (+/- SE).
Figure 3
Co-activation of AR is enhanced by c-Abl mediated tyrosine phosphorylation of p68 at Y593. A. COS-7 cells were co-transiently transfected in triplicate with 0.1 μg of p(ARE)3 luciferase reporter, 0.1 μg pcDNA3-AR, 0.1 μg of pCMV-β-gal and increasing concentrations of pcDNA3-p68WT (bar) and pcDNA3-p68Y593F (line) constructs (+ 10 nM R1881). B. Cropped immunoblot images of LNCaP nuclear lysates (+/- 10 nM R1881) co-immunoprecipitated (IP) with p68 goat polyclonal antibody probed sequentially with P-Tyr-100, c-Abl and p68 mouse monoclonal antibodies. Nuclear lysates of LNCaP cells transfected with p68Y593F mutant (full media + 10 nM R1881) IP with c-Abl mouse monoclonal antibody and sequentially probed with myc (p68Y593F) or c-Abl mouse monoclonal antibody. C. (ARE)3 luciferase reporter plus 0.1 μg pcDNA3-AR and increasing concentrations of pcDNA3-p68WT, pcDNA3-p68Y593F and/or pcDNA3-c-AblWT (+ 10 nM R1881). D. (ARE)3 luciferase reporter plus 0.1 μg pcDNA3-AR and increasing concentrations of pcDNA3-p68WT +/- 10 μM Imatinib (+ 10 nM R1881). Luciferase activity was corrected for the corresponding β-gal activity to give a relative activity. The range of plasmid levels (+, ++, +++ and ++++) corresponds to 25, 50, 75 and 100ng respectively. Data is shown relative to AR activity alone from at least three separate luciferase assay experiments (+/- SE).
Figure 4
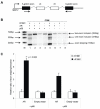
Silencing p68 protein expression by RNAi reduces AR and PSA mRNA and protein levels in LNCaP cells. A. Cropped immunoblot images of LNCaP lysates transfected with p68 and non-silencing (NS) targeted RNAi compared to control (C - non-transfected) cells (+ 10 nM R1881), probed sequentially with p68, AR, PSA and α-tubulin antibodies. N.B. NS and control protein levels are similar demonstrating no off target RNAi affects on protein expression. QPCR analysis of B. p68 C. PSA and D. AR mRNA expression in p68 targeted RNAi cells compared to non-silencing controls. QPCR data was normalised to GAPDH levels and fold change calculated to NS mRNA levels (set as 1). Figure shows the results from three independent RNAi experiments and at least three separate QPCR experiments (+/- SE).
Figure 5
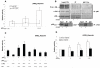
p68 enhances AR-dependent repression of a CD44 variable exon minigene. A. Minigene CDM contains variable exons v4 and v5 cloned downstream of steroid-responsive MMTV promoter. Arrows show location of primers used for RT-PCR. B. HEK 293 cells cultured in steroid-depleted media prior to transfection with the CDM minigene, pcDNA3-AR and pcDNA3-p68WT (+/- 10 nM R1881). Arrows indicate RT-PCR two exon inclusion (393bp), one exon inclusion (278bp) and exon skipped (163bp) products. Image is representative of at least three independent experiments. Marker (M): 1Kb plus DNA ladder (Invitrogen). C. Densitometric assessment from gel images to obtain mean relative ratios of exon skipped/inclusion product (+/- SE).
Figure 6
p68 protein expression levels in clinical prostate biopsies. A. Frequency of p68 immunostaining expression in PCa (66/147; 45%, Gleason Grade [GG] 3, 4 and 5) compared with BPH (11/44; 25%) biopsies. Fisher’s exact test was used for statistical significance. B. Representative images of p68 immunostaining in BPH, GG3, GG4 and GG5 PCa biopsy cores. Magnification, x 5. C. Increasing p68 immunostaining expression with increasing GG of PCa. Kruskall-Wallis test was used for statistical significance. D. Average p68 immunostaining expression in PCa compared with BPH. Mann-Whitney test was used for statistical significance.
Similar articles
- p68/DdX5 supports β-catenin & RNAP II during androgen receptor mediated transcription in prostate cancer.
Clark EL, Hadjimichael C, Temperley R, Barnard A, Fuller-Pace FV, Robson CN. Clark EL, et al. PLoS One. 2013;8(1):e54150. doi: 10.1371/journal.pone.0054150. Epub 2013 Jan 17. PLoS One. 2013. PMID: 23349811 Free PMC article. - Coupling transcription to RNA processing via the p68 DEAD box RNA helicase androgen receptor co-activator in prostate cancer.
Clark EL, Fuller-Pace FV, Elliott DJ, Robson CN. Clark EL, et al. Biochem Soc Trans. 2008 Jun;36(Pt 3):546-7. doi: 10.1042/BST0360546. Biochem Soc Trans. 2008. PMID: 18482004 Review. - Investigation of androgen receptor-dependent alternative splicing has identified a unique subtype of lethal prostate cancer.
Seltzer S, Giannopoulos PN, Bismar TA, Trifiro M, Paliouras M. Seltzer S, et al. Asian J Androl. 2023 May-Jun;25(3):296-308. doi: 10.4103/aja202263. Asian J Androl. 2023. PMID: 36259569 Free PMC article. - The RNA-binding and adaptor protein Sam68 modulates signal-dependent splicing and transcriptional activity of the androgen receptor.
Rajan P, Gaughan L, Dalgliesh C, El-Sherif A, Robson CN, Leung HY, Elliott DJ. Rajan P, et al. J Pathol. 2008 May;215(1):67-77. doi: 10.1002/path.2324. J Pathol. 2008. PMID: 18273831 - The role of DEAD-box RNA helicase p68 (DDX5) in the development and treatment of breast cancer.
Hashemi V, Masjedi A, Hazhir-Karzar B, Tanomand A, Shotorbani SS, Hojjat-Farsangi M, Ghalamfarsa G, Azizi G, Anvari E, Baradaran B, Jadidi-Niaragh F. Hashemi V, et al. J Cell Physiol. 2019 May;234(5):5478-5487. doi: 10.1002/jcp.26912. Epub 2018 Nov 11. J Cell Physiol. 2019. PMID: 30417346 Review.
Cited by
- RNA Helicases as Shadow Modulators of Cell Cycle Progression.
Sergeeva O, Zatsepin T. Sergeeva O, et al. Int J Mol Sci. 2021 Mar 15;22(6):2984. doi: 10.3390/ijms22062984. Int J Mol Sci. 2021. PMID: 33804185 Free PMC article. Review. - The DEAD-box RNA helicase DDX5 acts as a positive regulator of Japanese encephalitis virus replication by binding to viral 3' UTR.
Li C, Ge LL, Li PP, Wang Y, Sun MX, Huang L, Ishag H, Di DD, Shen ZQ, Fan WX, Mao X. Li C, et al. Antiviral Res. 2013 Nov;100(2):487-99. doi: 10.1016/j.antiviral.2013.09.002. Epub 2013 Sep 12. Antiviral Res. 2013. PMID: 24035833 Free PMC article. - RNA helicase p68 (DDX5) regulates tau exon 10 splicing by modulating a stem-loop structure at the 5' splice site.
Kar A, Fushimi K, Zhou X, Ray P, Shi C, Chen X, Liu Z, Chen S, Wu JY. Kar A, et al. Mol Cell Biol. 2011 May;31(9):1812-21. doi: 10.1128/MCB.01149-10. Epub 2011 Feb 22. Mol Cell Biol. 2011. PMID: 21343338 Free PMC article. - RNA splicing and splicing regulator changes in prostate cancer pathology.
Munkley J, Livermore K, Rajan P, Elliott DJ. Munkley J, et al. Hum Genet. 2017 Sep;136(9):1143-1154. doi: 10.1007/s00439-017-1792-9. Epub 2017 Apr 5. Hum Genet. 2017. PMID: 28382513 Free PMC article. Review. - Role of the DEAD-box RNA helicase DDX5 (p68) in cancer DNA repair, immune suppression, cancer metabolic control, virus infection promotion, and human microbiome (microbiota) negative influence.
Li F, Ling X, Chakraborty S, Fountzilas C, Wang J, Jamroze A, Liu X, Kalinski P, Tang DG. Li F, et al. J Exp Clin Cancer Res. 2023 Aug 19;42(1):213. doi: 10.1186/s13046-023-02787-x. J Exp Clin Cancer Res. 2023. PMID: 37596619 Free PMC article. Review.
References
- Ferlay J, Autier P, Boniol M, Heanue M, Colombet M, Boyle P. Estimates of the cancer incidence and mortality in Europe in 2006. Ann Oncol. 2007;18:581–92. - PubMed
- Gann PH. Interpreting recent trends in prostate cancer incidence and mortality. Epidemiology. 1997;8:117–20. - PubMed
- Feldman BJ, Feldman D. The development of androgen-independent prostate cancer. Nature Review Cancer. 2001;1:34–45. - PubMed
- Chen CD, Welsbie DS, Tran C, et al. Molecular determinants of resistance to antiandrogen therapy. Nature Medicine. 2004;10:33–9. - PubMed
- Chmelar R, Buchanan G, Need EF, Tilley W, Greenberg NM. Androgen receptor coregulators and their involvement in the development and progression of prostate cancer. International Journal of Cancer. 2007;120:719–33. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- G0100100(64424)/MRC_/Medical Research Council/United Kingdom
- G0100100/MRC_/Medical Research Council/United Kingdom
- G0500482/MRC_/Medical Research Council/United Kingdom
- G0100100/64424/DH_/Department of Health/United Kingdom
- 06-0705/AICR_/Worldwide Cancer Research/United Kingdom
- 063389/WT_/Wellcome Trust/United Kingdom
- CRUK_/Cancer Research UK/United Kingdom
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous