Integrated analysis of germline and somatic variants in ovarian cancer - PubMed (original) (raw)
Kimberly J Johnson 2, Charles Lu 1, Michael D McLellan 3, Mark D M Leiserson 4, Michael C Wendl 5, Qunyuan Zhang 6, Daniel C Koboldt 3, Mingchao Xie 3, Cyriac Kandoth 3, Joshua F McMichael 3, Matthew A Wyczalkowski 3, David E Larson 6, Heather K Schmidt 3, Christopher A Miller 3, Robert S Fulton 6, Paul T Spellman 7, Elaine R Mardis 8, Todd E Druley 9, Timothy A Graubert 10, Paul J Goodfellow 11, Benjamin J Raphael 4, Richard K Wilson 8, Li Ding 12
Affiliations
- PMID: 24448499
- PMCID: PMC4025965
- DOI: 10.1038/ncomms4156
Integrated analysis of germline and somatic variants in ovarian cancer
Krishna L Kanchi et al. Nat Commun. 2014.
Abstract
We report the first large-scale exome-wide analysis of the combined germline-somatic landscape in ovarian cancer. Here we analyse germline and somatic alterations in 429 ovarian carcinoma cases and 557 controls. We identify 3,635 high confidence, rare truncation and 22,953 missense variants with predicted functional impact. We find germline truncation variants and large deletions across Fanconi pathway genes in 20% of cases. Enrichment of rare truncations is shown in BRCA1, BRCA2 and PALB2. In addition, we observe germline truncation variants in genes not previously associated with ovarian cancer susceptibility (NF1, MAP3K4, CDKN2B and MLL3). Evidence for loss of heterozygosity was found in 100 and 76% of cases with germline BRCA1 and BRCA2 truncations, respectively. Germline-somatic interaction analysis combined with extensive bioinformatics annotation identifies 222 candidate functional germline truncation and missense variants, including two pathogenic BRCA1 and 1 TP53 deleterious variants. Finally, integrated analyses of germline and somatic variants identify significantly altered pathways, including the Fanconi, MAPK and MLL pathways.
Conflict of interest statement
COMPETING FINANCIAL INTERESTS
The authors declare no competing financial interests.
Figures
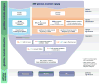
Figure 1. Overview of the integrated analysis of germline and somatic variants in 429 TCGA serous ovarian cases
A total of 27,280 somatic mutations were identified, including 6 SMGs (blue shaded area). germline variants included a total of 3 BRCA1 large-scale deletions, Following filtering of variants with >1% MAF in the population, TCGA ovarian cancer cases, and WHISP controls, a total of 3,635 truncation variants, and 22,953 missense variants (17,348 in expressed genes) remained for TCGA cases. For WHISP controls a total of 10,443 truncation and 30,335 missense variants (in expressed genes) remained. After applying the burden test using WHISP exome sequence data, a total of 6 and 24 genes were significantly enriched for truncation events and missense variants, respectively (orange shaded area). The germline/somatic interaction analysis (purple shaded area) that retained variants in expressed genes in ovarian cancer that met 2 out of 5 criteria identified a total of 237 candidate germline susceptibility variants. The pathway analysis identified three significant pathways involved in ovarian cancer pathogenesis, Fanconi, MAPK, and MLL.
Figure 2. Germline copy number variants in BRCA1
Shown are three germline copy number deletion variants affecting BRCA1 in three ovarian tumor pairs. Normal samples appear above the corresponding tumor samples. Red lines indicate normalized copy number segments based on a minimum of eight probes and blue dots indicate individual probe intensities from Affymetrix 6.0 SNP arrays within the region.
Figure 3. Lolliplots showing the distribution of germline truncation variants and somatic mutations
Somatic mutations in BRCA1, BRCA2, PALB2, CHEK2, BRIP1, BLM, MAP3K15, and PTPRH are shown in blue and germline truncation variants are in orange. Two known pathogenic BRCA1 germline missense variants are also shown (G1788V and R1699W).
Figure 4. Loss of heterozygosity analysis in tumor samples
(a) Scatter plot displaying variant allele frequencies for all germline truncation variants in normal and tumor samples. Truncation variants in BRCA1 and BRCA2 are highlighted in red and blue, respectively. (b) Scatter plot displaying variant allele frequencies for germline missense variants from cancer genes in normal and tumor samples. Germline missense variants in BRCA1 and BRCA2 are highlighted in red and blue, respectively. (c) VAFs for the 32 samples showing LOH truncation in BRCA1, (d) VAFs for 25 samples showing LOH in BRCA2, (e) VAFs in ATM, BLM, BRIP1, CHEK2, ERCC2, FANCA, and PALB2. Overall, 100% (32/32) and 76% (19/25) of respective germline BRCA1 and BRCA2 truncation variants showed increased VAFs in the tumor. All germline truncation variants in BRIP1 and CHEK2 also showed increased VAFs in corresponding tumors.
Figure 5. Significant pathways and subnetworks in ovarian cancer
(a) Oncoprint of genes with germline truncation variants and somatic mutations found in the Fanconi subnetwork identified as significant by HotNet. Genes in the iRefIndex database 58 are underlined. (b) The age distribution for patients with or without germline alterations in Fanconi genes (genes include: Panel a). The Horizontal red line indicates the median age of the group and the blue whiskers represent the age of the individual sample. (c) Oncoprint of genes with germline truncation variants and somatic mutations found in the MAPK subnetwork identified as significant by HotNet. Additional genes in the MAPK pathway with somatic mutations and/or germline truncation variants are included. (d) Oncoprint of genes with germline truncation variants and somatic mutations found in a subnetwork including MLL, MLL3, and SETD1A identified as significant by HotNet. Additional chromatin modifiers with somatic mutations and/or germline truncation variants are included.
Figure 5. Significant pathways and subnetworks in ovarian cancer
(a) Oncoprint of genes with germline truncation variants and somatic mutations found in the Fanconi subnetwork identified as significant by HotNet. Genes in the iRefIndex database 58 are underlined. (b) The age distribution for patients with or without germline alterations in Fanconi genes (genes include: Panel a). The Horizontal red line indicates the median age of the group and the blue whiskers represent the age of the individual sample. (c) Oncoprint of genes with germline truncation variants and somatic mutations found in the MAPK subnetwork identified as significant by HotNet. Additional genes in the MAPK pathway with somatic mutations and/or germline truncation variants are included. (d) Oncoprint of genes with germline truncation variants and somatic mutations found in a subnetwork including MLL, MLL3, and SETD1A identified as significant by HotNet. Additional chromatin modifiers with somatic mutations and/or germline truncation variants are included.
References
- Howlader NNA, Krapcho M, Neyman N, Aminou R, Altekruse SF, Kosary CL, Ruhl J, Tatalovich Z, Cho H, Mariotto A, Eisner MP, Lewis DR, Chen HS, Feuer EJ, Cronin KA, editors. SEER Cancer Statistics Review, 1975–2009 (Vintage 2009 Populations) National Cancer Institute; Bethesda, MD: Apr, 2012. http://seer.cancer.gov/csr/1975_2009_pops09/, based on November 2011 SEER data submission, posted to the SEER web site.
- Weissman SM, Weiss SM, Newlin AC. Genetic testing by cancer site: ovary. Cancer J. 2012;18:320–7. -PubMed
Publication types
MeSH terms
Grants and funding
- R01HG005690/HG/NHGRI NIH HHS/United States
- R01 HG005690/HG/NHGRI NIH HHS/United States
- U54HG003079/HG/NHGRI NIH HHS/United States
- R01 CA180006/CA/NCI NIH HHS/United States
- U54 HG003079/HG/NHGRI NIH HHS/United States
- R01CA180006/CA/NCI NIH HHS/United States
- U01 HG006517/HG/NHGRI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous