DNA Methylation Reprogramming during Mammalian Development - PubMed (original) (raw)
Review
DNA Methylation Reprogramming during Mammalian Development
Yang Zeng et al. Genes (Basel). 2019.
Abstract
DNA methylation (5-methylcytosine, 5mC) is a major form of DNA modification in the mammalian genome that plays critical roles in chromatin structure and gene expression. In general, DNA methylation is stably maintained in somatic tissues. However, DNA methylation patterns and levels show dynamic changes during development. Specifically, the genome undergoes two waves of global demethylation and remethylation for the purpose of producing the next generation. The first wave occurs in the germline, initiated with the erasure of global methylation in primordial germ cells (PGCs) and completed with the establishment of sex-specific methylation patterns during later stages of germ cell development. The second wave occurs after fertilization, including the erasure of most methylation marks inherited from the gametes and the subsequent establishment of the embryonic methylation pattern. The two waves of DNA methylation reprogramming involve both distinct and shared mechanisms. In this review article, we provide an overview of the key reprogramming events, focusing on the important players in these processes, including DNA methyltransferases (DNMTs) and ten-eleven translocation (TET) family of 5mC dioxygenases.
Keywords: DNA methylation; DNMTs; TETs; embryogenesis; germ cells.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Figure 1
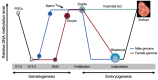
Dynamic changes in DNA methylation during mammalian development. Schematically shown are the two waves of global DNA demethylation and remethylation in the life cycle (adapted from [6]). Primordial germ cells (PGCs) initially have high levels of DNA methylation. Global demethylation occurs during PGC expansion and migration. At later stages of germ cell development (before birth in male and after birth in female), de novo methylation results in the establishment of sex-specific germ cell methylation patterns, including methylation marks at imprinted loci. Shortly after fertilization, the methylation marks inherited from the gametes are erased again (except those at imprinted loci and some retrotransposons), with the paternal genome undergoing active demethylation and the maternal genome undergoing passive demethylation. Upon implantation, a wave of de novo methylation establishes the initial embryonic methylation pattern.
Figure 2
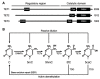
DNA methylation machinery. (A) The protein domains in DNA methyltransferases (DNMTs) and ubiquitin−like with plant homeodomain (PHD) and really interesting new gene (RING) finger domains 1 (UHRF1) are shown (the sizes/amino acid numbers refer to mouse proteins). The DNMT1 and DNMT3 families of proteins share conserved catalytic motifs (I–X) in the C-terminal catalytic domains (DNMT3L lacks catalytic activity because some essential motifs are missing or mutated) but have distinct N-terminal regulatory regions. PBD, proliferating cell nuclear antigen (PCNA)-binding domain; NLS, nuclear localization signal; RFTS, replication foci targeting sequence; CXXC, a cysteine-rich zinc finger domain; BAH, bromo-adjacent homology domain; (GK)n, glycine/lysine repeats; PWWP, proline-tryptophan-tryptophan-proline domain; ADD, ATRX-DNMT3-DNMT3L domain; UBL, ubiquitin-like domain; TTD, tandem Tudor domain; PHD, plant homeodomain; SRA, Su(var)3-9, Enhancer of zeste, and Trithorax (SET)- and RING-associated domain; RING, really interesting new gene domain. (B) De novo and maintenance methylation activities. The de novo methyltransferases (DNMT3A, DNMT3B, and DNMT3C), in complex with their accessory factor DNMT3L, methylate unmethylated CpG sites to establish methylation patterns. The maintenance methyltransferase DNMT1, in complex with its accessory factor UHRF1, methylates hemi-methylated CpG sites at each round of DNA replication to maintain methylation patterns.
Figure 3
Ten-eleven translocation (TET) proteins and relevant DNA demethylation pathways. (A) The protein domains in TET proteins (TET1, TET2 and TET3) are shown (the sizes/amino acid numbers refer to mouse proteins). Their C-terminal catalytic domains contain two characteristic sequence features, a cysteine-rich region (Cys) and a double-stranded β helix (DSBH) fold. Their N-terminal regulatory regions are less conserved, with TET1 and TET3 containing a CXXC zinc finger domain. (B) DNA demethylation pathways involving TETs. TET proteins initiate DNA demethylation by oxidizing 5-methylcytosine (5mC) to 5-hydroxymethylcytosine (5hmC), which can be further oxidized to 5-formylcytosine (5fC) and 5-carboxylcytosine (5caC). 5fC and 5caC can be excised by thymine DNA glycosylase (TDG). The residual abasic site can then be repaired by the base excision repair (BER) pathway to complete ‘active’ demethylation. 5mC, 5hmC, 5fC, and 5caC can also be removed through DNA replication-coupled ‘passive’ dilution.
Figure 4
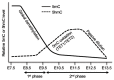
DNA demethylation in primordial germ cells (PGCs). PGCs undergo DNA demethylation in two phases. The first phase is mainly a passive process due to repression of important components of the DNA methylation machinery, resulting in global demethylation. The second phase, which affects specific loci including imprinted genes, is initiated by TET1- and TET2-mediated 5mC oxidation, followed by passive dilution of oxidized derivatives.
Figure 5
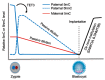
DNA demethylation and remethylation during early embryogenesis. Shortly after fertilization, TET3-mediated 5mC oxidation occurs in the paternal genome and the oxidized derivatives are subsequently removed through passive dilution during preimplantation development. The maternal genome mainly undergoes passive demethylation during preimplantation development. Upon implantation, a wave of de novo methylation establishes the embryonic methylation pattern.
Similar articles
- DNA methylation reprogramming of genomic imprints in the mammalian germline: A TET-centric view.
Caldwell BA, Bartolomei MS. Caldwell BA, et al. Andrology. 2023 Jul;11(5):884-890. doi: 10.1111/andr.13303. Epub 2022 Oct 2. Andrology. 2023. PMID: 36150101 Review. - DNA methylation dynamics during epigenetic reprogramming of medaka embryo.
Wang X, Bhandari RK. Wang X, et al. Epigenetics. 2019 Jun;14(6):611-622. doi: 10.1080/15592294.2019.1605816. Epub 2019 Apr 22. Epigenetics. 2019. PMID: 31010368 Free PMC article. - Role of Tet1 in erasure of genomic imprinting.
Yamaguchi S, Shen L, Liu Y, Sendler D, Zhang Y. Yamaguchi S, et al. Nature. 2013 Dec 19;504(7480):460-4. doi: 10.1038/nature12805. Epub 2013 Dec 1. Nature. 2013. PMID: 24291790 Free PMC article. - Reprogramming DNA methylation in the mammalian life cycle: building and breaking epigenetic barriers.
Seisenberger S, Peat JR, Hore TA, Santos F, Dean W, Reik W. Seisenberger S, et al. Philos Trans R Soc Lond B Biol Sci. 2013 Jan 5;368(1609):20110330. doi: 10.1098/rstb.2011.0330. Philos Trans R Soc Lond B Biol Sci. 2013. PMID: 23166394 Free PMC article. Review. - Promoter DNA methylation couples genome-defence mechanisms to epigenetic reprogramming in the mouse germline.
Hackett JA, Reddington JP, Nestor CE, Dunican DS, Branco MR, Reichmann J, Reik W, Surani MA, Adams IR, Meehan RR. Hackett JA, et al. Development. 2012 Oct;139(19):3623-32. doi: 10.1242/dev.081661. Development. 2012. PMID: 22949617 Free PMC article.
Cited by
- Global analysis of DNA methylation changes during experimented lingual carcinogenesis.
Liu H, Yue W, Shao S, Sun J, Yang Y, Dai X. Liu H, et al. Hua Xi Kou Qiang Yi Xue Za Zhi. 2024 Jun 1;42(3):319-328. doi: 10.7518/hxkq.2024.2023416. Hua Xi Kou Qiang Yi Xue Za Zhi. 2024. PMID: 39049651 Free PMC article. Chinese, English. - A DNA Methylation Perspective on Infertility.
Shacfe G, Turko R, Syed HH, Masoud I, Tahmaz Y, Samhan LM, Alkattan K, Shafqat A, Yaqinuddin A. Shacfe G, et al. Genes (Basel). 2023 Nov 27;14(12):2132. doi: 10.3390/genes14122132. Genes (Basel). 2023. PMID: 38136954 Free PMC article. Review. - Genetic and epigenetic regulation of growth, reproduction, disease resistance and stress responses in aquaculture.
Liu Z, Zhou T, Gao D. Liu Z, et al. Front Genet. 2022 Nov 2;13:994471. doi: 10.3389/fgene.2022.994471. eCollection 2022. Front Genet. 2022. PMID: 36406125 Free PMC article. Review. - Bioinformatics of nanopore sequencing.
Makałowski W, Shabardina V. Makałowski W, et al. J Hum Genet. 2020 Jan;65(1):61-67. doi: 10.1038/s10038-019-0659-4. Epub 2019 Aug 26. J Hum Genet. 2020. PMID: 31451715 Review. - DNA Methylation and Schizophrenia: Current Literature and Future Perspective.
Magwai T, Shangase KB, Oginga FO, Chiliza B, Mpofana T, Xulu KR. Magwai T, et al. Cells. 2021 Oct 26;10(11):2890. doi: 10.3390/cells10112890. Cells. 2021. PMID: 34831111 Free PMC article. Review.
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources