U1 small nuclear RNA and spliceosomal introns in Euglena gracilis - PubMed (original) (raw)
U1 small nuclear RNA and spliceosomal introns in Euglena gracilis
D G Breckenridge et al. Proc Natl Acad Sci U S A. 1999.
Abstract
In the flagellated protozoon Euglena gracilis, characterized nuclear genes harbor atypical introns that usually are flanked by short repeats, adopt complex secondary structures in pre-mRNA, and do not obey the GT-AG rule of conventional cis-spliced introns. In the nuclear fibrillarin gene of E. gracilis, we have identified three spliceosomal-type introns that have GT-AG consensus borders. Furthermore, we have isolated a small RNA from E. gracilis and propose, on the basis of primary and secondary structure comparisons, that it is a homolog of U1 small nuclear RNA, an essential component of the cis-spliceosome in higher eukaryotes. Conserved sequences at the 5' splice sites of the fibrillarin introns can potentially base pair with Euglena U1 small nuclear RNA. Our observations demonstrate that spliceosomal GT-AG cis-splicing occurs in Euglena, in addition to the nonconventional cis-splicing and spliced leader trans-splicing previously recognized in this early diverging unicellular eukaryote.
Figures
Figure 1
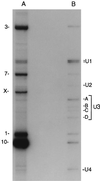
Immunoprecipitation of TMG-capped Euglena small RNAs. Samples were 3′-end-labeled with [5′-32P]pCp and RNA ligase and were resolved in a 10% polyacrylamide/7 M urea sequencing gel. (A) Total RNA. The most abundant labeled bands in this sample [1 (163 nt), 3 (350 nt), 7 (235 nt), and 10 (164 nt)] correspond to components of the fragmented large subunit rRNA (33). Band X (181 nt) was generated by ligation of large subunit rRNA species 12 to the 3′-end of large subunit rRNA species 14 (33). (B) Anti-TMG immunoprecipitate. Chemical sequencing revealed that the mAb enriched for U1 (see Results and Discussion), U2, and U4 (identified by sequence analysis and comparison with homologous sequences; M.N.S., unpublished results) snRNAs as well as four bands (A–D) containing variant forms of U3 small nucleolar RNA (34).
Figure 2
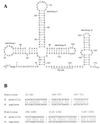
(A) Primary sequence and secondary structure of E. gracilis U1 snRNA. Am, _O_2′-methyladenosine; Um, _O_2′-methyluridine. (B) Alignment of regions of E. gracilis and Homo sapiens (42) U1 snRNAs. Only regions that can be aligned unambiguously are shown (position numbers are based on the E. gracilis sequence). Identities between the two sequences are indicated (|).
Figure 3
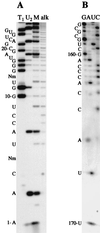
Autoradiograms of representative sequencing gels (20% acrylamide/7 M urea). (A) Enzymatic sequencing of 5′-end-labeled E. gracilis U1 snRNA. T1, RNase T1 (cleaves after G); U2, RNase U2 (cleaves after A); M, RNase Phy M (cleaves after A>U); alk, alkali ladder; Nm, _O_2′-methylnucleoside. (B) Chemical sequencing of the long version (170 nt) of E. gracilis U1 snRNA. Minor bands are the result of cross-contamination with the short version (169 nt) of this RNA species.
Figure 4
(A) Schematic representation of a genomic PCR product corresponding to a segment of the E. gracilis fibrillarin gene and comparison with the cDNA sequence. (B) Primary sequence of the PCR product. Positions of the three introns (lower case) are indicated with the conserved GT and AG motifs at the 5′- and 3′-splice junctions underlined. Positions corresponding to the PCR primer binding sites are presented in bold.
Figure 5
Potential interactions between the 5′-terminal region of U1 snRNA and the 5′-splice sites of the three fibrillarin pre-mRNA introns and between the 5′-terminal region of U1 snRNA and the 5′-splice site of the SL RNA (17, 18). The cap at the 5′-end of U1 snRNA is not shown. The exon–intron boundaries are indicated by a slash (/), with the conserved GU at the 5′-end of each intron underlined.
Similar articles
- Order of removal of conventional and nonconventional introns from nuclear transcripts of Euglena gracilis.
Gumińska N, Płecha M, Zakryś B, Milanowski R. Gumińska N, et al. PLoS Genet. 2018 Oct 26;14(10):e1007761. doi: 10.1371/journal.pgen.1007761. eCollection 2018 Oct. PLoS Genet. 2018. PMID: 30365503 Free PMC article. - Spliceosomal introns in conserved sequences of U1 and U5 small nuclear RNA genes in yeast Rhodotorula hasegawae.
Takahashi Y, Tani T, Ohshima Y. Takahashi Y, et al. J Biochem. 1996 Sep;120(3):677-83. doi: 10.1093/oxfordjournals.jbchem.a021465. J Biochem. 1996. PMID: 8902636 - Trans-splicing and cis-splicing in the colourless Euglenoid, Entosiphon sulcatum.
Ebel C, Frantz C, Paulus F, Imbault P. Ebel C, et al. Curr Genet. 1999 Jun;35(5):542-50. doi: 10.1007/s002940050451. Curr Genet. 1999. PMID: 10369962 - snRNAs as the catalysts of pre-mRNA splicing.
Valadkhan S. Valadkhan S. Curr Opin Chem Biol. 2005 Dec;9(6):603-8. doi: 10.1016/j.cbpa.2005.10.008. Epub 2005 Oct 20. Curr Opin Chem Biol. 2005. PMID: 16242989 Review. - RNA splicing in lower eukaryotes.
Woolford JL Jr, Peebles CL. Woolford JL Jr, et al. Curr Opin Genet Dev. 1992 Oct;2(5):712-9. doi: 10.1016/s0959-437x(05)80131-5. Curr Opin Genet Dev. 1992. PMID: 1333856 Review.
Cited by
- A possible role for short introns in the acquisition of stroma-targeting peptides in the flagellate Euglena gracilis.
Vesteg M, Vacula R, Steiner JM, Mateásiková B, Löffelhardt W, Brejová B, Krajcovic J. Vesteg M, et al. DNA Res. 2010 Aug;17(4):223-31. doi: 10.1093/dnares/dsq015. Epub 2010 Jun 29. DNA Res. 2010. PMID: 20587589 Free PMC article. - U3 snoRNA genes are multi-copy and frequently linked to U5 snRNA genes in Euglena gracilis.
Charette JM, Gray MW. Charette JM, et al. BMC Genomics. 2009 Nov 16;10:528. doi: 10.1186/1471-2164-10-528. BMC Genomics. 2009. PMID: 19917113 Free PMC article. - Genomic characterization of Neoparamoeba pemaquidensis (Amoebozoa) and its kinetoplastid endosymbiont.
Tanifuji G, Kim E, Onodera NT, Gibeault R, Dlutek M, Cawthorn RJ, Fiala I, Lukes J, Greenwood SJ, Archibald JM. Tanifuji G, et al. Eukaryot Cell. 2011 Aug;10(8):1143-6. doi: 10.1128/EC.05027-11. Epub 2011 Jun 10. Eukaryot Cell. 2011. PMID: 21666073 Free PMC article. - Viral and cellular mRNA capping: past and prospects.
Furuichi Y, Shatkin AJ. Furuichi Y, et al. Adv Virus Res. 2000;55:135-84. doi: 10.1016/s0065-3527(00)55003-9. Adv Virus Res. 2000. PMID: 11050942 Free PMC article. Review.
References
- Breathnach R, Chambon P. Annu Rev Biochem. 1981;50:349–383. - PubMed
- Moore M J, Query C C, Sharp P A. In: The RNA World. Gesteland R F, Atkins J F, editors. Plainview, New York: Cold Spring Harbor Lab. Press; 1993. pp. 303–357.
- Lamond A I. BioEssays. 1993;15:595–603. - PubMed
- Lührmann R, Kastner B, Bach M. Biochim Biophys Acta. 1990;1087:265–292. - PubMed
- Baserga S J, Steitz J A. In: The RNA World. Gesteland R F, Atkins J F, editors. Plainview, NY: Cold Spring Harbor Lab. Press; 1993. pp. 359–381.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous