lncRNAdb: a reference database for long noncoding RNAs (original) (raw)
Abstract
Large numbers of long RNAs with little or no protein-coding potential [long noncoding RNAs (lncRNAs)] are being identified in eukaryotes. In parallel, increasing data describing the expression profiles, molecular features and functions of individual lncRNAs in a variety of systems are accumulating. To enable the systematic compilation and updating of this information, we have developed a database (lncRNAdb) containing a comprehensive list of lncRNAs that have been shown to have, or to be associated with, biological functions in eukaryotes, as well as messenger RNAs that have regulatory roles. Each entry contains referenced information about the RNA, including sequences, structural information, genomic context, expression, subcellular localization, conservation, functional evidence and other relevant information. lncRNAdb can be searched by querying published RNA names and aliases, sequences, species and associated protein-coding genes, as well as terms contained in the annotations, such as the tissues in which the transcripts are expressed and associated diseases. In addition, lncRNAdb is linked to the UCSC Genome Browser for visualization and Noncoding RNA Expression Database (NRED) for expression information from a variety of sources. lncRNAdb provides a platform for the ongoing collation of the literature pertaining to lncRNAs and their association with other genomic elements. lncRNAdb can be accessed at: http://www.lncrnadb.org/.
INTRODUCTION
The eukaryotic transcriptome is enormous, and comprises not only a large set of protein-coding messenger RNAs, but also large numbers of non-protein coding transcripts that have structural, regulatory or unknown functions (1). Noncoding RNAs include intergenic transcripts as well as a complex array of RNAs that overlap protein-coding loci on both strands (2–4), including promoter-associated RNAs (4), intronic RNAs (5), convergent and bi-directional transcripts (3), noncoding alternatively spliced isoforms of protein-coding genes (6) and mRNAs that also have regulatory roles as untranslated RNAs (7,8). While some of these RNAs are processed to form small RNA species such as microRNAs (miRNAs) and small nucleolar RNAs (snoRNAs), many can have diverse roles as primary or spliced long noncoding RNAs (lncRNAs) (9). In structural terms, lncRNAs range in size from approximately 100 to 100 000 bases, can be spliced or unspliced, polyadenylated or non-polyadenylated, nuclear or cytoplasmic, and are usually transcribed by RNA polymerase II and/or III (10).
The combination of different experimental methodologies to study the transcriptome of several species has resulted in a continuous discovery of novel transcripts (11), with the FANTOM project alone cataloguing more than 30 000 putative lncRNA transcripts in mouse tissues by full-length cDNA cloning (2). This provides a challenge for their molecular and functional characterization, as well as for their cataloguing. Although the subset of lncRNAs characterized to date corresponds to a small fraction of the long noncoding transcripts in multicellular organisms, growing interest in the field has resulted in a rapid increase in the characterization of individual lncRNAs in a variety of systems.
A decade ago less than a dozen lncRNAs were recognized and catalogued in all eukaryotes (12), but following the dramatic increase in their numbers, a recent catalogue listed approximately 40 functionally characterized lncRNAs in mammals alone (13). This list contained lncRNAs whose functions had been assessed either by loss- or gain-of-function experiments, and did not include many that have other types of evidence for functionality. For example, several studies have documented the cell type-specific and/or dynamic expression of hundreds of long ncRNAs in various developmental systems, such as embryonic stem cell differentiation (14) and oligodendrogenesis (15), finding in each case different subsets of differentially expressed lncRNAs.
These studies have indicated that lncRNAs comprise a class of bona fide gene products that have been largely unaccounted for in public databases such as RefSeq (16), UniGene (17) and the Mammalian Gene Collection (18). While there are databases cataloguing ncRNAs, these have been predominantly focused on well-validated classes of small RNAs and contain limited data pertaining to lncRNAs. Recently, a database with diverse information on imprinted RNAs was made available, which is largely comprised of small RNAs, such as piRNAs (piwi-interacting RNAs), but also contains imprinted lncRNAs, and is restricted to mammals (19). Our previous database of noncoding RNAs, RNAdb (20), was also limited to mammals, and included all classes of regulatory RNAs, such as snoRNAs, piRNAs and miRNAs. Due to the rapid expansion in each of these classes, they are now independently curated in dedicated databases, including miRbase (21), piRNAbank (22) and Sno/scaRNAbase (23). Consequently, there is a need for a dedicated database of lncRNAs that includes not only mammalian lncRNAs, but detailed annotations of lncRNAs from all eukaryotic species.
AIMS OF THE DATABASE
lncRNAdb provides a central repository of known lncRNAs in eukaryotic cells (including those derived from viruses), their aliases and published characteristics. A well-collated library will greatly facilitate research of these poorly annotated genes. It also aims to decrease instances of replication and unknown identity by the inclusion of aliases such as p15AS, which was reported as a novel antisense RNA (24), but appears to be an unspliced isoform of a previously described ncRNA called ANRIL (25), and Gomafu (26), which is also known as MIAT (27) and RNCR2 (28).
The centralization of lncRNAdb enables integration with other resources including the UCSC Genome Browser (29) and the Noncoding RNA Expression Database (NRED) (30), which provide insight into genomic context and expression data. This ensures that researchers interested in lncRNAs can conveniently find a wide range of information regarding genes of interest from a single location.
Currently, the characteristics and functions of most lncRNAs are still unexplored, but it is expected that the number of studied RNAs will rapidly increase by in silico, in vitro and in vivo characterization. By providing a simple interface accessible to public users, we aim to provide a tool for the scientific community that will allow existing entries to be updated, modified to improve accuracy, and new entries to be added by users. Following verification of new published data by the curators, the information will become available in lncRNAdb.
DATABASE STRUCTURE
lncRNAdb is available online at: http://www.lncrnadb.org. Users can search the database by lncRNA name, nucleotide sequence string, species, annotation status or through a full-text search with results being displayed for online perusal and available as a tab delimited file download.
Annotated entries include one or more literature references, annotations across a series of categories and a list of species in which the lncRNA is observed. Literature references and genomic coordinates are hyperlinked to PubMed and the UCSC Genome Browser, respectively. Other biological components related to the annotated lncRNAs, such as genomically-associated genes or interacting proteins, are also listed and briefly described in a separate table that provides links to the PubMed reference.
The lncRNAdb database also links to NRED, our online expression analysis application for ncRNAs in mouse and human (30). This extends the database through access to relative expression levels of both the lncRNAs and their contextually related transcripts in various public microarray experiments, such as NCode data and the GNF atlas (31), as well as the Allen Brain Atlas (32), which includes mouse brain in situ expression data for over 800 expressed lncRNAs (33).
The application architecture consists of a Microsoft asp.NET 3.5 presentation layer (c#), c# 4.0 data model and application layer, and MySQL persistent storage.
QUERYING THE DATABASE
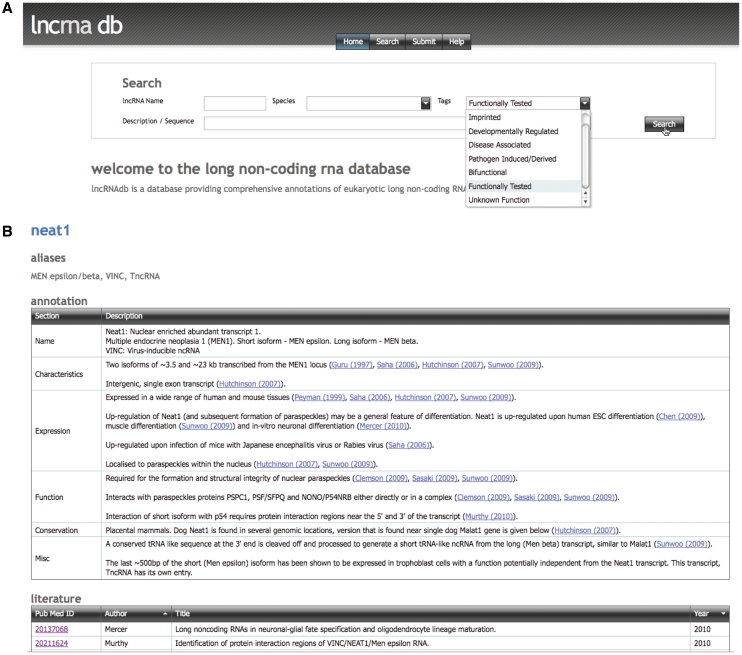
Querying the database is a matter of entering one or more search criteria (Figure 1A); a full or partial lncRNA name (or alias), a search string to interrogate each ncRNA’s annotations or a species in which the lncRNA is known, or any combination of those. lncRNAdb will return a list of matching entries whose full detail can be viewed by clicking on the lncRNA name.
Figure 1.
Representative screenshots of lncRNAdb showing (A) the search bar and (B) a lncRNA catalogued in lncRNAdb. Figure 1A shows the search fields available for querying the database and some of the pre-made descriptors available. Figure 1B depicts part of the annotation and references for the Neat1 transcript.
To download the database values for the entire list as a file of tab delimited text, users can click on the ‘Export Results’ button under the search results.
lncRNAdb CONTENT
Although lncRNAs have been defined previously as transcripts >200 nt (9), this was an arbitrary definition largely based on a convenient biochemical cutoff in RNA isolation protocols and the fact that it excluded most known small RNAs. In an effort to make a clear distinction to small RNA species and to create a more biologically meaningful definition of lncRNAs, we consider lncRNAs as noncoding RNAs that may have a function as either primary or spliced transcripts, which is independent of processing into known classes of small RNAs, such as miRNAs, piRNAs and snoRNAs, while also excluding structural RNAs from classical housekeeping families (tRNAs, snoRNAs, spliceosomal RNAs, etc). Existing databases and archives, including Sno/scaRNAbase (23), miRBase (21), tRNAdb-CE (34) and piRNAbank (22), already represent such ncRNAs. However, some lncRNAs that are host genes for small RNA species (35,36), but may also have roles as regulatory lncRNAs, have been included in the annotations. An example is the GAS5 lncRNA, which is a repressor of the glucocorticoid receptor but also encodes several intronic snoRNAs (37).
The database currently contains over 150 lncRNAs identified from the literature in around 60 different species. Each entry contains a comprehensive range of available information about the RNA, including sequences, structural information, genomic context, expression, subcellular localization, conservation, functional evidence and relevant ‘miscellaneous’ information (see e.g. in Figure 1B). As expected, most (∼75%) of catalogued lncRNAs are from mammals, for which more transcriptomic data is available and which have been more intensively studied, but lncRNAs from vertebrates to single-celled eukaryotes have been included.
Among the entries in lncRNAdb, approximately 100 have functions directly tested by in vitro and/or in vivo experiments. For quick reference, these have been listed in Supplementary Table S1, highlighting the wide array of functional mechanisms and processes affected by lncRNAs. This list is not exhaustive, but it catalogues functional lncRNAs found not only in mammals but also lncRNAs tested in diverse eukaryotic species, such as meiRNA (38) in yeast, frq antisense RNAs in Neurospora crassa (39), rncs-1 in Caenorhabditis elegans (40), IPS1 (41) in plants, Xlsirts in frogs (42) and bereft in Drosophila (43). In addition to RNAs expressed in normal physiological states, the database contains information about RNAs expressed in disease, and even lncRNAs derived from viruses and expressed in eukaryotic cells during infection (see below). Examples of lncRNA categories that can be used to limit queries are listed below.
Imprinted lncRNAs
LncRNAs are prevalent in imprinted regions where they can function to control imprinting and the expression of other genes from the locus, such as Air (44) and Kncq1ot1 (45). Some imprinted lncRNAs are host genes for small RNAs, such as Bsr (46), where the lncRNA host has no known function; or has an independent function, e.g. the putative tumour suppressor Meg3/Gtl2 (47).
Disease associated lncRNAs
Underscoring their importance in cellular functions, a growing number of lncRNAs have been implicated in a variety of diseases, including cancer. These are described in the database and include putative or confirmed cancer-associated lncRNAs, such as NDM29 (48), HOTAIR, which regulates metastatic progression (49), and H19, which has been described as both an oncogene (50) and a tumour suppressor (51). We also include lncRNAs that have been implicated in neurological functions and diseases, including BACE1AS (52) that shows increased expression in Alzheimer’s disease, and the Drosophila hsr-ω gene, which is induced by a variety of stresses and which has been shown to greatly increase protein polyglutamine-induced toxicity and neurodegeneration (53).
In addition, the observation that a substantial fraction of the genotypic variation underlying complex phenotypic traits occurs in noncoding regions, many of which are transcribed into discrete lncRNAs, has led to the appreciation that lncRNAs may play a central role in the molecular etiology of complex diseases (13). We have catalogued RNAs associated with these loci in lncRNAdb, such as ANRIL, a well-characterized lncRNA located in the complex genetic susceptibility locus INK4b/ARF/INK4a implicated in coronary artery disease, type 2 diabetes, periodontitis and cancer (24,54,55). We opted to also include uncharacterized lncRNAs linked to disease-susceptibility loci, as cataloguing these transcripts may facilitate their recognition as candidates for functional studies in normal and pathological conditions. Examples include AK023948, which is located in a susceptibility locus to papillary thyroid tumour (56), and LOC285194, which is located in a copy number alteration and loss-of-heterozygosity region in osteosarcoma (57).
Pathogen-induced or derived lncRNAs
Some lncRNAs are produced and modulated by pathogens or host cells during infection. These have been mostly omitted from other noncoding RNA catalogues, but incorporated in lncRNAdb. These include eukaryotic parasite transcripts such as the Pinci1 ncRNA family in the fungal plant-pathogen Phytophthora infestans, which are specifically upregulated during infection (58), and mammalian lncRNAs that are induced during viral infection (59), such as Neat1/VINC in the mouse brain (60), or produced in infected cells by oncogenic viruses, such as human herpesviruses (61). Indeed, the accumulating examples of lncRNAs encoded by viruses and expressed in eukaryotic cells have been annotated in lncRNAdb, because they can regulate cell function and are relevant to disease etiology. For instance, β2.7 is a ∼2.7 kb ncRNA encoded in the herpesvirus HCMV genome that is rapidly accumulated upon infection and has a fundamental role in preventing metabolic dysfunction and apoptosis of the host cell (62).
Bifunctional RNAs
An emerging class of genes is those encoding bifunctional RNAs, which can have multiple independent roles, such as acting as a regulatory lncRNA or being translated into a protein. Examples include well-known lncRNAs such as the co-activator SRA transcript, isoforms of which also encode a protein (8,63), and known protein coding genes, such as p53, whose transcripts also act as regulatory RNAs (7). In some cases, splicing isoforms are known to encode a protein, whereas specific splicing isoforms encode regulatory lncRNAs, such as the LXRB/LXRBSV isoform pair (6).
lncRNAs of unknown function
lncRNAdb includes RNAs that are well described in the literature but whose functions have yet to be identified. These regions are transcribed into lncRNAs that have been characterized to some extent at the structural and/or expression level, including dozens of lncRNAs that show tissue specificity and dynamic expression during development (64), as well as cellular localization, suggestive of undiscovered functionality. Likewise, the database also includes transcripts such as PHO5 antisense lncRNA in yeast (65) where it is not yet established whether the functional role is conveyed by the lncRNA or its transcription.
FUTURE DIRECTIONS
The data in lncRNAdb will be extended through manual curation by the authors and submissions to the site by other researchers on an ongoing basis. The lowering cost and improved depths of RNA-sequencing methodologies is already enabling transcriptomics studies for alternative model- and non-model species, and it is expected that the identification and characterization of lncRNAs will follow, which may substantially increase the representation of non-mammalian species in the database. New functionality will be included with integration of the publicly available successor to NRED (NRED2) early in 2011, giving access to not only additional microarray expression data but also transcriptomic RNA-seq data for contextually related long noncoding and coding genes, tools for visualization of the lncRNA and cross-referencing of experimental expression profiles. Finally, the absence of a centralized database of lncRNAs is reflected by the ad hoc naming of newly identified members, which are often not self-consistent and potentially confusing. With the number of functional lncRNAs likely to be very large, it will be important to establish a standardized nomenclature to minimize confusion and allow this emerging field to be as accessible as possible to all biologists.
SUPPLEMENTARY DATA
Supplementary Data are available at NAR Online.
FUNDING
Australian Research Council/University of Queensland co-sponsored Federation Fellowship (FF0561986; to J.S.M.); National Health and Medical Research Council of Australia Career Development Award (CDA631542; to M.E.D.); Queensland Government Department of Employment, Economic Development and Innovation Smart Futures Fellowship (to M.E.D.); Australian Research Council Postgraduate Awards (to P.P.A., M.B.C. and D.G.A). Funding for open access charge: The University of Queensland.
Conflict of interest statement. None declared.
REFERENCES
- 1.Amaral PP, Dinger ME, Mercer TR, Mattick JS. The eukaryotic genome as an RNA machine. Science. 2008;319:1787–1789. doi: 10.1126/science.1155472. [DOI] [PubMed] [Google Scholar]
- 2.Carninci P, Kasukawa T, Katayama S, Gough J, Frith MC, Maeda N, Oyama R, Ravasi T, Lenhard B, Wells C, et al. The transcriptional landscape of the mammalian genome. Science. 2005;309:1559–1563. doi: 10.1126/science.1112014. [DOI] [PubMed] [Google Scholar]
- 3.Katayama S, Tomaru Y, Kasukawa T, Waki K, Nakanishi M, Nakamura M, Nishida H, Yap CC, Suzuki M, Kawai J, et al. Antisense transcription in the mammalian transcriptome. Science. 2005;309:1564–1566. doi: 10.1126/science.1112009. [DOI] [PubMed] [Google Scholar]
- 4.Kapranov P, Cheng J, Dike S, Nix DA, Duttagupta R, Willingham AT, Stadler PF, Hertel J, Hackermuller J, Hofacker IL, et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science. 2007;316:1484–1488. doi: 10.1126/science.1138341. [DOI] [PubMed] [Google Scholar]
- 5.Nakaya HI, Amaral PP, Louro R, Lopes A, Fachel AA, Moreira YB, El-Jundi TA, da Silva AM, Reis EM, Verjovski-Almeida S. Genome mapping and expression analyses of human intronic noncoding RNAs reveal tissue-specific patterns and enrichment in genes related to regulation of transcription. Genome Biol. 2007;8:R43. doi: 10.1186/gb-2007-8-3-r43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Hashimoto K, Ishida E, Matsumoto S, Shibusawa N, Okada S, Monden T, Satoh T, Yamada M, Mori M. A liver X receptor (LXR)-beta alternative splicing variant (LXRBSV) acts as an RNA co-activator of LXR-beta. Biochem. Biophys. Res. Commun. 2009;390:1260–1265. doi: 10.1016/j.bbrc.2009.10.132. [DOI] [PubMed] [Google Scholar]
- 7.Candeias MM, Malbert-Colas L, Powell DJ, Daskalogianni C, Maslon MM, Naski N, Bourougaa K, Calvo F, Fahraeus R. p53 mRNA controls p53 activity by managing Mdm2 functions. Nat. Cell Biol. 2008;10:1098–1105. doi: 10.1038/ncb1770. [DOI] [PubMed] [Google Scholar]
- 8.Chooniedass-Kothari S, Hamedani MK, Auge C, Wang X, Carascossa S, Yan Y, Cooper C, Vincett D, Myal Y, Jalaguier S, et al. The steroid receptor RNA activator protein is recruited to promoter regions and acts as a transcriptional repressor. FEBS Lett. 2010;584:2218–2224. doi: 10.1016/j.febslet.2010.04.022. [DOI] [PubMed] [Google Scholar]
- 9.Mercer TR, Dinger ME, Mattick JS. Long noncoding RNAs: insights into function. Nat. Rev. Genet. 2009;10:155–159. doi: 10.1038/nrg2521. [DOI] [PubMed] [Google Scholar]
- 10.Dinger ME, Amaral PP, Mercer TR, Mattick JS. Pervasive transcription of the eukaryotic genome: functional indices and conceptual implications. Brief. Funct. Genomic Proteomic. 2009;8:407–423. doi: 10.1093/bfgp/elp038. [DOI] [PubMed] [Google Scholar]
- 11.Huttenhofer A, Vogel J. Experimental approaches to identify non-coding RNAs. Nucleic Acids Res. 2006;34:635–646. doi: 10.1093/nar/gkj469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Erdmann VA, Szymanski M, Hochberg A, de Groot N, Barciszewski J. Collection of mRNA-like non-coding RNAs. Nucleic Acids Res. 1999;27:192–195. doi: 10.1093/nar/27.1.192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Mattick JS. The genetic signatures of noncoding RNAs. PLoS Genet. 2009;5:e1000459. doi: 10.1371/journal.pgen.1000459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Dinger ME, Amaral PP, Mercer TR, Pang KC, Bruce SJ, Gardiner BB, Askarian-Amiri ME, Ru K, Solda G, Simons C, et al. Long noncoding RNAs in mouse embryonic stem cell pluripotency and differentiation. Genome Res. 2008;18:1433–1445. doi: 10.1101/gr.078378.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Mercer TR, Qureshi IA, Gokhan S, Dinger ME, Li G, Mattick JS, Mehler MF. Long noncoding RNAs in neuronal-glial fate specification and oligodendrocyte lineage maturation. BMC Neurosci. 2010;11:14. doi: 10.1186/1471-2202-11-14. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Pruitt KD, Tatusova T, Maglott DR. NCBI reference sequences (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res. 2007;35:D61–D65. doi: 10.1093/nar/gkl842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Sayers EW, Barrett T, Benson DA, Bolton E, Bryant SH, Canese K, Chetvernin V, Church DM, Dicuccio M, Federhen S, et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2010;38:D5–D6. doi: 10.1093/nar/gkp967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Gerhard DS, Wagner L, Feingold EA, Shenmen CM, Grouse LH, Schuler G, Klein SL, Old S, Rasooly R, Good P, et al. The status, quality, and expansion of the NIH full-length cDNA project: the Mammalian Gene Collection (MGC) Genome Res. 2004;14:2121–2127. doi: 10.1101/gr.2596504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Zhang Y, Guan DG, Yang JH, Shao P, Zhou H, Qu LH. ncRNAimprint: a comprehensive database of mammalian imprinted noncoding RNAs. RNA. 2010;16:1889–1901. doi: 10.1261/rna.2226910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Pang KC, Stephen S, Dinger ME, Engstrom PG, Lenhard B, Mattick JS. RNAdb 2.0–an expanded database of mammalian non-coding RNAs. Nucleic Acids Res. 2007;35:D178–D182. doi: 10.1093/nar/gkl926. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ. miRBase: tools for microRNA genomics. Nucleic Acids Res. 2008;36:D154–D158. doi: 10.1093/nar/gkm952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Sai Lakshmi S, Agrawal S. piRNABank: a web resource on classified and clustered Piwi-interacting RNAs. Nucleic Acids Res. 2008;36:D173–D177. doi: 10.1093/nar/gkm696. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Xie J, Zhang M, Zhou T, Hua X, Tang L, Wu W. Sno/scaRNAbase: a curated database for small nucleolar RNAs and cajal body-specific RNAs. Nucleic Acids Res. 2007;35:D183–D187. doi: 10.1093/nar/gkl873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Yu W, Gius D, Onyango P, Muldoon-Jacobs K, Karp J, Feinberg AP, Cui H. Epigenetic silencing of tumour suppressor gene p15 by its antisense RNA. Nature. 2008;451:202–206. doi: 10.1038/nature06468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Pasmant E, Laurendeau I, Heron D, Vidaud M, Vidaud D, Bieche I. Characterization of a germ-line deletion, including the entire INK4/ARF locus, in a melanoma-neural system tumor family: identification of ANRIL, an antisense noncoding RNA whose expression coclusters with ARF. Cancer Res. 2007;67:3963–3969. doi: 10.1158/0008-5472.CAN-06-2004. [DOI] [PubMed] [Google Scholar]
- 26.Sone M, Hayashi T, Tarui H, Agata K, Takeichi M, Nakagawa S. The mRNA-like noncoding RNA Gomafu constitutes a novel nuclear domain in a subset of neurons. J. Cell Sci. 2007;120:2498–2506. doi: 10.1242/jcs.009357. [DOI] [PubMed] [Google Scholar]
- 27.Ishii N, Ozaki K, Sato H, Mizuno H, Saito S, Takahashi A, Miyamoto Y, Ikegawa S, Kamatani N, Hori M, et al. Identification of a novel non-coding RNA, MIAT, that confers risk of myocardial infarction. J. Hum. Genet. 2006;51:1087–1099. doi: 10.1007/s10038-006-0070-9. [DOI] [PubMed] [Google Scholar]
- 28.Blackshaw S, Harpavat S, Trimarchi J, Cai L, Huang H, Kuo WP, Weber G, Lee K, Fraioli RE, Cho SH, et al. Genomic analysis of mouse retinal development. PLoS Biol. 2004;2:E247. doi: 10.1371/journal.pbio.0020247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Rhead B, Karolchik D, Kuhn RM, Hinrichs AS, Zweig AS, Fujita PA, Diekhans M, Smith KE, Rosenbloom KR, Raney BJ, et al. The UCSC Genome Browser database: update 2010. Nucleic Acids Res. 2010;38:D613–D619. doi: 10.1093/nar/gkp939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Dinger ME, Pang KC, Mercer TR, Crowe ML, Grimmond SM, Mattick JS. NRED: a database of long noncoding RNA expression. Nucleic Acids Res. 2009;37:D122–D126. doi: 10.1093/nar/gkn617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Su AI, Wiltshire T, Batalov S, Lapp H, Ching KA, Block D, Zhang J, Soden R, Hayakawa M, Kreiman G, et al. A gene atlas of the mouse and human protein-encoding transcriptomes. Proc. Natl Acad. Sci. USA. 2004;101:6062–6067. doi: 10.1073/pnas.0400782101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lein ES, Hawrylycz MJ, Ao N, Ayres M, Bensinger A, Bernard A, Boe AF, Boguski MS, Brockway KS, Byrnes EJ, et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature. 2007;445:168–176. doi: 10.1038/nature05453. [DOI] [PubMed] [Google Scholar]
- 33.Mercer TR, Dinger ME, Sunkin SM, Mehler MF, Mattick JS. Specific expression of long noncoding RNAs in the mouse brain. Proc. Natl Acad. Sci. USA. 2008;105:716–721. doi: 10.1073/pnas.0706729105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Abe T, Ikemura T, Ohara Y, Uehara H, Kinouchi M, Kanaya S, Yamada Y, Muto A, Inokuchi H. tRNADB-CE: tRNA gene database curated manually by experts. Nucleic Acids Res. 2009;37:D163–D168. doi: 10.1093/nar/gkn692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Tycowski KT, Shu MD, Steitz JA. A mammalian gene with introns instead of exons generating stable RNA products. Nature. 1996;379:464–466. doi: 10.1038/379464a0. [DOI] [PubMed] [Google Scholar]
- 36.Bortolin ML, Kiss T. Human U19 intron-encoded snoRNA is processed from a long primary transcript that possesses little potential for protein coding. RNA. 1998;4:445–454. [PMC free article] [PubMed] [Google Scholar]
- 37.Kino T, Hurt DE, Ichijo T, Nader N, Chrousos GP. Noncoding RNA gas5 is a growth arrest- and starvation-associated repressor of the glucocorticoid receptor. Sci. Signal. 2010;3:ra8. doi: 10.1126/scisignal.2000568. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Watanabe Y, Yamamoto M. S. pombe mei2+ encodes an RNA-binding protein essential for premeiotic DNA synthesis and meiosis I, which cooperates with a novel RNA species meiRNA. Cell. 1994;78:487–498. doi: 10.1016/0092-8674(94)90426-x. [DOI] [PubMed] [Google Scholar]
- 39.Kramer C, Loros JJ, Dunlap JC, Crosthwaite SK. Role for antisense RNA in regulating circadian clock function in Neurospora crassa. Nature. 2003;421:948–952. doi: 10.1038/nature01427. [DOI] [PubMed] [Google Scholar]
- 40.Hellwig S, Bass BL. A starvation-induced noncoding RNA modulates expression of Dicer-regulated genes. Proc. Natl Acad. Sci. USA. 2008;105:12897–12902. doi: 10.1073/pnas.0805118105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Franco-Zorrilla JM, Valli A, Todesco M, Mateos I, Puga MI, Rubio-Somoza I, Leyva A, Weigel D, Garcia JA, Paz-Ares J. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat. Genet. 2007;39:1033–1037. doi: 10.1038/ng2079. [DOI] [PubMed] [Google Scholar]
- 42.Kloc M, Wilk K, Vargas D, Shirato Y, Bilinski S, Etkin LD. Potential structural role of non-coding and coding RNAs in the organization of the cytoskeleton at the vegetal cortex of Xenopus oocytes. Development. 2005;132:3445–3457. doi: 10.1242/dev.01919. [DOI] [PubMed] [Google Scholar]
- 43.Hardiman KE, Brewster R, Khan SM, Deo M, Bodmer R. The bereft gene, a potential target of the neural selector gene cut, contributes to bristle morphogenesis. Genetics. 2002;161:231–247. doi: 10.1093/genetics/161.1.231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Sleutels F, Zwart R, Barlow DP. The non-coding Air RNA is required for silencing autosomal imprinted genes. Nature. 2002;415:810–813. doi: 10.1038/415810a. [DOI] [PubMed] [Google Scholar]
- 45.Pandey RR, Mondal T, Mohammad F, Enroth S, Redrup L, Komorowski J, Nagano T, Mancini-Dinardo D, Kanduri C. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol. Cell. 2008;32:232–246. doi: 10.1016/j.molcel.2008.08.022. [DOI] [PubMed] [Google Scholar]
- 46.Royo H, Basyuk E, Marty V, Marques M, Bertrand E, Cavaille J. Bsr, a nuclear-retained RNA with monoallelic expression. Mol. Biol. Cell. 2007;18:2817–2827. doi: 10.1091/mbc.E06-10-0920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Zhang X, Gejman R, Mahta A, Zhong Y, Rice KA, Zhou Y, Cheunsuchon P, Louis DN, Klibanski A. Maternally expressed gene 3, an imprinted noncoding RNA gene, is associated with meningioma pathogenesis and progression. Cancer Res. 2010;70:2350–2358. doi: 10.1158/0008-5472.CAN-09-3885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Castelnuovo M, Massone S, Tasso R, Fiorino G, Gatti M, Robello M, Gatta E, Berger A, Strub K, Florio T, et al. An Alu-like RNA promotes cell differentiation and reduces malignancy of human neuroblastoma cells. Faseb J. 2010;24:4033–4046. doi: 10.1096/fj.10-157032. [DOI] [PubMed] [Google Scholar]
- 49.Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature. 2010;464:1071–1076. doi: 10.1038/nature08975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Matouk IJ, DeGroot N, Mezan S, Ayesh S, Abu-lail R, Hochberg A, Galun E. The H19 non-coding RNA is essential for human tumor growth. PLoS ONE. 2007;2:e845. doi: 10.1371/journal.pone.0000845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Yoshimizu T, Miroglio A, Ripoche MA, Gabory A, Vernucci M, Riccio A, Colnot S, Godard C, Terris B, Jammes H, et al. The H19 locus acts in vivo as a tumor suppressor. Proc. Natl Acad. Sci. USA. 2008;105:12417–12422. doi: 10.1073/pnas.0801540105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Faghihi MA, Modarresi F, Khalil AM, Wood DE, Sahagan BG, Morgan TE, Finch CE, St. Laurent G, 3rd, Kenny PJ, Wahlestedt C. Expression of a noncoding RNA is elevated in Alzheimer’s disease and drives rapid feed-forward regulation of beta-secretase. Nat. Med. 2008;14:723–730. doi: 10.1038/nm1784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Sengupta S, Lakhotia SC. Altered expressions of the noncoding hsromega gene enhances poly-Q-induced neurotoxicity in Drosophila. RNA Biol. 2006;3:28–35. doi: 10.4161/rna.3.1.2559. [DOI] [PubMed] [Google Scholar]
- 54.Holdt LM, Beutner F, Scholz M, Gielen S, Gabel G, Bergert H, Schuler G, Thiery J, Teupser D. ANRIL expression is associated with atherosclerosis risk at chromosome 9p21. Arterioscler. Thromb. Vasc. Biol. 2010;30:620–627. doi: 10.1161/ATVBAHA.109.196832. [DOI] [PubMed] [Google Scholar]
- 55.Yap KL, Li S, Munoz-Cabello AM, Raguz S, Zeng L, Mujtaba S, Gil J, Walsh MJ, Zhou MM. Molecular interplay of the noncoding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol. Cell. 2010;38:662–674. doi: 10.1016/j.molcel.2010.03.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.He H, Nagy R, Liyanarachchi S, Jiao H, Li W, Suster S, Kere J, de la Chapelle A. A susceptibility locus for papillary thyroid carcinoma on chromosome 8q24. Cancer Res. 2009;69:625–631. doi: 10.1158/0008-5472.CAN-08-1071. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Pasic I, Shlien A, Durbin AD, Stavropoulos DJ, Baskin B, Ray PN, Novokmet A, Malkin D. Recurrent focal copy-number changes and loss of heterozygosity implicate two noncoding RNAs and one tumor suppressor gene at chromosome 3q13.31 in osteosarcoma. Cancer Res. 2010;70:160–171. doi: 10.1158/0008-5472.CAN-09-1902. [DOI] [PubMed] [Google Scholar]
- 58.Avrova AO, Whisson SC, Pritchard L, Venter E, De Luca S, Hein I, Birch PR. A novel non-protein-coding infection-specific gene family is clustered throughout the genome of Phytophthora infestans. Microbiology. 2007;153:747–759. doi: 10.1099/mic.0.2006/002220-0. [DOI] [PubMed] [Google Scholar]
- 59.Mrazek J, Kreutmayer SB, Grasser FA, Polacek N, Huttenhofer A. Subtractive hybridization identifies novel differentially expressed ncRNA species in EBV-infected human B cells. Nucleic Acids Res. 2007;35:e73. doi: 10.1093/nar/gkm244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Saha S, Murthy S, Rangarajan PN. Identification and characterization of a virus-inducible non-coding RNA in mouse brain. J. Gen. Virol. 2006;87:1991–1995. doi: 10.1099/vir.0.81768-0. [DOI] [PubMed] [Google Scholar]
- 61.Swaminathan S. Noncoding RNAs produced by oncogenic human herpesviruses. J. Cell Physiol. 2008;216:321–326. doi: 10.1002/jcp.21480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Reeves MB, Davies AA, McSharry BP, Wilkinson GW, Sinclair JH. Complex I binding by a virally encoded RNA regulates mitochondria-induced cell death. Science. 2007;316:1345–1348. doi: 10.1126/science.1142984. [DOI] [PubMed] [Google Scholar]
- 63.Foulds CE, Tsimelzon A, Long W, Le A, Tsai SY, Tsai MJ, O’Malley BW. Research resource: expression profiling reveals unexpected targets and functions of the human steroid receptor RNA activator (SRA) gene. Mol. Endocrinol. 2010;24:1090–1105. doi: 10.1210/me.2009-0427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Amaral PP, Mattick JS. Noncoding RNA in development. Mamm. Genome. 2008;19:454–492. doi: 10.1007/s00335-008-9136-7. [DOI] [PubMed] [Google Scholar]
- 65.Uhler JP, Hertel C, Svejstrup JQ. A role for noncoding transcription in activation of the yeast PHO5 gene. Proc. Natl Acad. Sci. USA. 2007;104:8011–8016. doi: 10.1073/pnas.0702431104. [DOI] [PMC free article] [PubMed] [Google Scholar]