Epigenetic modifications in cardiovascular disease (original) (raw)
Introduction
The term “epigenetics” refers to heritable changes in gene expression, which are not a result of changes in the DNA sequence, but rather due to alterations related to the packaging (thereby altering DNA accessibility) and/or translation of genetic information [[6](/article/10.1007/s00395-012-0245-9#ref-CR6 "Bird A (2007) Perceptions of epigenetics. Nature 447:396–398. doi: 10.1038/nature05913
")\]. These changes are mediated through complex environment/genome interactions. Even though epigenetic variability of genetic information is part of normal development and differentiation it also underlies exogenous stimuli, e.g., smoking or drug abuse, and as such may reflect the role of these factors on the development of diseases. Important alterations encompassing epigenetic changes are DNA methylation and RNA-based mechanisms including those controlled by microRNAs (miRNA) and long intergenic non-coding RNAs (linc-RNA). These modifications result in a variable expression of identical genetic information based on the surrounding conditions leading to enhanced expression or silencing of genes. The study of these alterations represents a fascinating avenue to further elucidate the underlying mechanisms of cardiovascular disease. This review provides an introduction to epigenetic mechanisms such as DNA methylation, histone modifications and RNA-based modifications as well as an outline of how these changes pertain to the development of cardiovascular disease.DNA methylation
In the mammalian genome the most widely appreciated epigenetic modification involves the covalent binding of a methyl group, termed DNA methylation, to the 5′-position of cytosine residues in the so-called CpG (Cytosine preceding Guanosine) dinucleotides (see Fig. 1) [[7](/article/10.1007/s00395-012-0245-9#ref-CR7 "Bird A (1986) CpG-rich islands and the function of DNA methylation. Nature 321:209–213. doi: 10.1038/321209a0
")\]. CpG-islands differ from the remainder of the genomic pattern in that they are Cytosine-Guanosine-rich (CpG-rich) and predominantly non-methylated \[[16](/article/10.1007/s00395-012-0245-9#ref-CR16 "Deaton AM, Bird A (2011) CpG islands and the regulation of transcription. Genes Dev 25(10):1010–1022. doi:
10.1002/bies.201000057
")\]. CpG-islands have a length of approximately 200–300 base pairs with a Cytosine + Guanosine (C + G) content of about 50% and an observed CpG/expected CpG in excess of 0.6\. \[[22](/article/10.1007/s00395-012-0245-9#ref-CR22 "Gardiner-Garden M, Frommer M (1987) CpG islands in vertebrate genomes. J Mol Biol 196:261–282. doi:
10.1016/0022-2836(87)90689-9
")\]. However, more recent analyses have shown that regions of DNA with a length of more than 500 base pairs, including a G + C content of at least 55% and observed CpG/expected CpG of 0.65 were more likely to be associated with the 5′ regions of genes \[[71](/article/10.1007/s00395-012-0245-9#ref-CR71 "Takai D, Jones PA (2002) Comprehensive analysis of CpG islands in human chromosomes 21 and 22. Proc Natl Acad Sci USA 99:3740–3745. doi:
10.1073/pnas.052410099
")\]. This definition excluded most _Alu_\-repetitive elements. Most CpG-Islands are sites of transcription initiation. Due to the mutagenic properties of methylcytosine vertebrate genomes are methylated predominantly at the dinucleotide CpG and are CpG-deficient \[[8](/article/10.1007/s00395-012-0245-9#ref-CR8 "Bird AP (1980) DNA methylation and the frequency of CpG in animal DNA. Nucleic Acids Res 8:1499–1504. doi:
10.1093/nar/8.7.1499
")\]. A small fraction of cytosines (around 4%) are physiologically methylated in the mammalian genome \[[21](/article/10.1007/s00395-012-0245-9#ref-CR21 "Gama-Sosa MA, Midgett RM, Slagel VA, Githens S, Kuo KC, Gehrke CW, Ehrlich M (1983) Tissue-specific differences in DNA methylation in various mammals. Biochim Biophys Acta 740:212–219. doi:
10.1016/0167-4781(83)90079-9
"), [53](/article/10.1007/s00395-012-0245-9#ref-CR53 "Mund C, Brueckner B, Lyko F (2006) Reactivation of epigenetically silenced genes by DNA methyltransferase inhibitors: basic concepts and clinical applications. Epigenetics 1(1):7–13. doi:
10.4161/epi.1.1.2375
")\].Fig. 1
DNA methylation. Chromosome, histones and the DNA double-helix including a CpG-rich DNA sequence is shown, where the coupling of methyl groups is catalyzed by methyltransferases (DNMT). These changes may also occur in response to exogenous stimuli, e.g., smoking, cocaine abuse, arterial hypertension (examples are given in the main text). DNMT-3a and -3b drive “de novo” methylation of genes, while DNMT-1 is associated with maintaining the methylation state. Demethylation occurs after a reduction in DNMT-1 activity. Recent evidence suggests that demethylation may also be an active process (see [[14](/article/10.1007/s00395-012-0245-9#ref-CR14 "Chen L, Hahn H, Wu G, Chen CH, Liron T, Schechtman D, Cavallaro G, Banci L, Guo Y, Bolli R, Dorn GW 2nd, Mochly-Rosen D (2001) Opposing cardioprotective actions and parallel hypertrophic effects of delta PKC and epsilon PKC. Proc Natl Acad Sci USA 98:11114–11119. doi: 10.1073/pnas.191369098
")\])DNA methylation plays an essential role in several epigenetic phenomena including genomic imprinting, X-chromosome inactivation and retro-element silencing [[42](/article/10.1007/s00395-012-0245-9#ref-CR42 "Li E, Beard C, Jaenisch R (1993) Role for DNA methylation in genomic imprinting. Nature 366:362–365. doi: 10.1038/366362a0
"), [59](/article/10.1007/s00395-012-0245-9#ref-CR59 "Panning B, Jaenisch R (1996) DNA hypomethylation can activate Xist expression and silence X-linked genes. Genes Dev 10:1991–2002. doi:
10.1101/gad.10.16.1991
"), [79](/article/10.1007/s00395-012-0245-9#ref-CR79 "Walsh CP, Chaillet JR, Bestor TH (1998) Transcription of IAP endogenous retroviruses is constrained by cytosine methylation. Nat Genet 20:116–117. doi:
10.1038/2413
")\]. CpG dinucleotides are unequally distributed across the human genome. Since vast stretches of sequences are deficient for CpGs and are interspersed by CpG clusters, these are also defined as “CpG islands” (CpG-rich areas). They are mainly associated with promoter regions of genes (approximately 70% of gene promoters) and mostly remain unmethylated \[[22](/article/10.1007/s00395-012-0245-9#ref-CR22 "Gardiner-Garden M, Frommer M (1987) CpG islands in vertebrate genomes. J Mol Biol 196:261–282. doi:
10.1016/0022-2836(87)90689-9
"), [53](/article/10.1007/s00395-012-0245-9#ref-CR53 "Mund C, Brueckner B, Lyko F (2006) Reactivation of epigenetically silenced genes by DNA methyltransferase inhibitors: basic concepts and clinical applications. Epigenetics 1(1):7–13. doi:
10.4161/epi.1.1.2375
"), [67](/article/10.1007/s00395-012-0245-9#ref-CR67 "Saxonov S, Berg P, Brutlag DL (2006) A genome-wide analysis of CpG dinucleotides in the human genome distinguishes two distinct classes of promoters. Proc Natl Acad Sci USA 103(5):1412–1417. doi:
10.1073/pnas.0510310103
"), [71](/article/10.1007/s00395-012-0245-9#ref-CR71 "Takai D, Jones PA (2002) Comprehensive analysis of CpG islands in human chromosomes 21 and 22. Proc Natl Acad Sci USA 99:3740–3745. doi:
10.1073/pnas.052410099
")\]. Cytosine methylation is enzymatically driven by a transfer of a methyl group from the methyl donor _S_\-adenosylmethionine to the carbon-5 position of cytosine \[[38](/article/10.1007/s00395-012-0245-9#ref-CR38 "Laird PW (2003) The power and the promise of DNA methylation markers. Nat Rev Cancer 3(4):253–266. doi:
10.1038/nrc1045
")\]. DNA methylation is, as currently known, catalyzed by at least three different DNA methyltransferases (DNMTs): DNMT1, DNMT3a and DNMT3b \[[11](/article/10.1007/s00395-012-0245-9#ref-CR11 "Broadbent HM, Peden JF, Lorkowski S, Goel A, Ongen H, Green F, Clarke R, Collins R, Franzosi MG, Tognoni G, Seedorf U, Rust S, Eriksson P, Hamsten A, Farrall M, Watkins H, PROCARDIS Consortium (2008) Susceptibility to coronary artery disease and diabetes is encoded by distinct, tightly linked SNPs in the ANRIL locus on chromosome 9p. Hum Mol Genet 17:806–814. doi:
10.1093/hmg/ddm352
")\]. DNMT1 represents the main enzyme in mammals. It serves a maintenance function (referred to as maintenance methylation), since it is responsible for postreplicative (during mitotic cell division) restoration of hemi-methylated sites to full methylation \[[38](/article/10.1007/s00395-012-0245-9#ref-CR38 "Laird PW (2003) The power and the promise of DNA methylation markers. Nat Rev Cancer 3(4):253–266. doi:
10.1038/nrc1045
")\]. Demethylation may occur after a reduction in DNMT-1 activity. Recent evidence suggests that demethylation may also be an active process \[[5](/article/10.1007/s00395-012-0245-9#ref-CR5 "Bhutani N, Burns DM, Blau HM (2011) DNA demethylation dynamics. Cell 146(6):866–872. doi:
10.1016/j.cell.2011.08.042
")\]. However, this has not been proven in the cardiovascular system. DNMT3a and DNMT3b are implicated in de novo methylation (see Fig. [1](/article/10.1007/s00395-012-0245-9#Fig1)) \[[80](/article/10.1007/s00395-012-0245-9#ref-CR80 "Weber M, Hellmann I, Stadler MB, Ramos L, Paabo S, Rebhan M, Schubeler D (2007) Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome. Nat Genet 39:457–466. doi:
10.1038/ng1990
")\].Methylation of cytosine in CpG islands is associated with transcriptional repression by impeding the binding of transcription factors to _cis_-DNA binding elements present in the promoter regions of genes [[80](/article/10.1007/s00395-012-0245-9#ref-CR80 "Weber M, Hellmann I, Stadler MB, Ramos L, Paabo S, Rebhan M, Schubeler D (2007) Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome. Nat Genet 39:457–466. doi: 10.1038/ng1990
")\]. Recently, a family of methyl-CpG binding proteins has also been recognized that specifically bind to methylation marks, thereby contributing to transcriptional repression by recruiting histone-modifying proteins. These include the MBD protein family (MBD1, MBD2, MBD4 and MeCP2), Kaiso and Kaiso-like proteins, and SRA domain proteins (e.g., SUVH9 and SUVH2) \[[17](/article/10.1007/s00395-012-0245-9#ref-CR17 "Defossez PA, Stancheva I (2011) Biological functions of methyl-CpG-binding proteins. Prog Mol Biol Transl Sci 101:377–398. doi:
10.1016/B978-0-12-387685-0.00012-3
")\].Histone modifications
DNA within eukaryotic cells is condensed in chromatin. The nucleosome is the core unit of chromatin and is composed of an octamer of four different histones (H3, H4, H2A, and H2B), around which DNA is wound with a length of approximately 140–150 base pairs (see Fig. 2) [[37](/article/10.1007/s00395-012-0245-9#ref-CR37 "Kouzarides T (2007) Chromatin modifications and their function. Cell 128(4):693–705. doi: 10.1016/j.cell.2007.02.005
")\]. Histones are characterized by their large number and diversity of modified residues \[[37](/article/10.1007/s00395-012-0245-9#ref-CR37 "Kouzarides T (2007) Chromatin modifications and their function. Cell 128(4):693–705. doi:
10.1016/j.cell.2007.02.005
")\]. Currently, at least eight different types of modifications are known. These modifications are catalyzed by distinct enzymes. These include enzymes for methylation, acetylation, phosphorylation, sumoylation, ubiquitination, ADP-ribosylation, deimination and proline isomerisation \[[37](/article/10.1007/s00395-012-0245-9#ref-CR37 "Kouzarides T (2007) Chromatin modifications and their function. Cell 128(4):693–705. doi:
10.1016/j.cell.2007.02.005
")\]. Histone modifications serve the role of allotting the genome into “active” or euchromatin, in which DNA is accessible for transcription, and “inactive” or heterochromatin, in which DNA is inaccessible for transcription \[[37](/article/10.1007/s00395-012-0245-9#ref-CR37 "Kouzarides T (2007) Chromatin modifications and their function. Cell 128(4):693–705. doi:
10.1016/j.cell.2007.02.005
")\].Fig. 2
Histone modifications. Within the nucleus DNA is packaged as chromatin in the nucleosome. It is composed of an octamer of four different histones (H3, H4, H2A, H2B). Histones display a large number of modified residues. Histone acetylation, methylation and phosphorylation are shown as examples of histone modifications. These modifications are catalyzed by distinct enzymes. Histone modifications divide the genome into “active” or euchromatin, in which DNA is accessible for transcription, and “inactive” or heterochromatin, in which DNA is inaccessible for transcription. Examples of how histone modifications may be altered in cardiovascular (patho)physiology are shown. K lysine, S serine
RNA-based mechanisms
Recent high-throughput analyses of the transcriptome have revealed that the genome transcribes ~90% of genomic DNA, of which only 1–2% encode for proteins, while the majority are transcribed as non-coding RNAs [[18](/article/10.1007/s00395-012-0245-9#ref-CR18 "The ENCODE Project Consortium (2004) The ENCODE (ENCyclopedia Of DNA Elements) Project. Science 306:636–640. doi: 10.1126/science.1105136
")\]. There is mounting evidence for the role of non-coding RNAs in the regulation of development \[[23](/article/10.1007/s00395-012-0245-9#ref-CR23 "Golding MC, Magri LS, Zhang L, Lalone SA, Higgins MJ, Mann MR (2011) Depletion of Kcnq1ot1 non-coding RNA does not affect imprinting maintenance in stem cells. Development 138(17):3667–3678. doi:
10.1242/dev.057778
")\], in response to environmental stress \[[20](/article/10.1007/s00395-012-0245-9#ref-CR20 "Ferguson LR (2011) RNA silencing: mechanism, biology and responses to environmental stress. Mutat Res 714(1–2):93–94. doi:
10.1016/j.mrfmmm.2011.07.007
")\] and disease initiation/progression \[[81](/article/10.1007/s00395-012-0245-9#ref-CR81 "Yang F, Zhang L, Huo XS, Yuan JH, Xu D, Yuan SX, Zhu N, Zhou WP, Yang GS, Wang YZ, Shang JL, Gao CF, Zhang FR, Wang F, Sun SH (2011) Long non-coding RNA high expressed in hepatocellular carcinoma (lncRNA-HEIH) facilitates tumor growth through enhancer of zeste homolog 2. Hepatology (in press). doi:
10.1002/hep.24563
")\]. Current research efforts are therefore aimed at elucidating the role of non-coding RNAs with respect to physiological functions and diseases.Non-coding RNAs are classified as infrastructural (small nuclear and nucleolar RNAs, ribosomal RNAs) and regulatory RNAs (microRNAs, long non-coding RNAs, Piwi-interacting RNAs, small interfering RNAs). We here will focus on the role of two regulatory RNAs—microRNAs and long non-coding RNAs—in the development of cardiovascular disease.
MicroRNAs
In the early 1990s Lee and co-workers were the first to describe a function for a specific miRNA, lin-4, during postembryonic development in the nematode Caenorhabditis elegans [[39](/article/10.1007/s00395-012-0245-9#ref-CR39 "Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75:843–854. doi: 10.1016/0092-8674(93)90529-Y
")\]. Since then, an array of miRNAs have been discovered and analyzed. Currently, 851 different miRNAs have been identified in humans, 793 in mice and 698 in rats according to the MicroCosm Targets web resource \[[49](/article/10.1007/s00395-012-0245-9#ref-CR49 "MicroCosm Targets (2010) Version 5. EMBL–EBI database [online].
http://www.ebi.ac.uk/enright-srv/microcosm/htdocs/targets/v5/
")\] (formerly miRBase Targets) developed by the Enright laboratory at the European Bioinformatics Institute, Cambridge, UK. However, the exact number of miRNAs within different species is currently unknown.MiRNAs lead to the repression of target genes through the post-transcriptional degradation of messenger-RNA and/or translational inhibition of protein expression [[4](/article/10.1007/s00395-012-0245-9#ref-CR4 "Bauersachs J, Thum T (2011) Biogenesis and regulation of cardiovascular microRNAs. Circ Res 109(3):334–347. doi: 10.1161/CIRCRESAHA.110.228676
")\]. Similar to mRNAs, primary miRNAs (pri-miRNAs) have a 5′ 7-methyl guanylate cap and 3′ polyadenylated tail \[[12](/article/10.1007/s00395-012-0245-9#ref-CR12 "Cai X, Hagedorn CH, Cullen BR (2004) Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA 10:1957–1966. doi:
10.1261/rna.7135204
"), [40](/article/10.1007/s00395-012-0245-9#ref-CR40 "Lee Y, Kim M, Han J, Yeom KH, Lee S, Baek SH, Kim VN (2004) MicroRNA genes are transcribed by RNA polymerase II. EMBO J 23:4051–4060. doi:
10.1038/sj.emboj.7600385
")\]. After transcription of the pri-miRNAs by RNA polymerase II, the pri-miRNA, Drosha and the RNA-binding protein DGCR8 complex is processed into a hairpin structure, termed the precursor miRNA \[[26](/article/10.1007/s00395-012-0245-9#ref-CR26 "Gregory RI, Yan KP, Amuthan G, Chendrimada T, Doratotaj B, Cooch N, Shiekhattar R (2004) The microprocessor complex mediates the genesis of microRNAs. Nature 432:235–240. doi:
10.1038/nature03120
"), [40](/article/10.1007/s00395-012-0245-9#ref-CR40 "Lee Y, Kim M, Han J, Yeom KH, Lee S, Baek SH, Kim VN (2004) MicroRNA genes are transcribed by RNA polymerase II. EMBO J 23:4051–4060. doi:
10.1038/sj.emboj.7600385
"), [41](/article/10.1007/s00395-012-0245-9#ref-CR41 "Lee Y, Ahn C, Han J, Choi H, Kim J, Yim J, Lee J, Provost P, Rådmark O, Kim S, Kim VN (2003) The nuclear RNase III Drosha initiates microRNA processing. Nature 425:415–419. doi:
10.1038/nature01957
")\]. Via binding to exportin 5 and Ran-GTP the precursor miRNA is transported into the cytoplasm, where it is cleaved by Dicer, and processed into a double-stranded product consisting of 22 nucleotides. This mature miRNA consists of a guide strand and a passenger strand. The miRNA guide strand is incorporated into the RNA-induced silencing complex (RISC) while the passenger strand is degraded. The RISC–miRNA complex specifically targets mRNAs and leads to negative regulation of protein synthesis or mRNA degradation \[[1](/article/10.1007/s00395-012-0245-9#ref-CR1 "Ambros V (2004) The functions of animal microRNAs. Nature 431:350–355. doi:
10.1038/nature02871
"), [3](/article/10.1007/s00395-012-0245-9#ref-CR3 "Bartel DP (2003) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297. doi:
10.1016/S0092-8674(04)00045-5
"), [77](/article/10.1007/s00395-012-0245-9#ref-CR77 "Valencia-Sanchez MA, Liu J, Hannon GJ, Parker R (2006) Control of translation and mRNA degradation by miRNAs and siRNAs. Genes Dev 20:515–524. doi:
10.1101/gad.1399806
")\]. Using a ribosome profiling strategy, it was recently shown that miRNAs predominantly act though destabilization of target mRNAs, which subsequently leads to reduced protein output \[[27](/article/10.1007/s00395-012-0245-9#ref-CR27 "Guo H, Ingolia NT, Weissman JS, Bartel DP (2010) Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature 466:835–840. doi:
10.1038/nature09267
")\]. These results show that destabilization of target mRNAs in addition to translational inhibition may also be a mechanism resulting in impaired protein production.Currently, several groups have elucidated epigenetic silencing of certain genes encoding microRNAs in the cancer field, thus fundamentally impacting on the expression of genetic information [[2](/article/10.1007/s00395-012-0245-9#ref-CR2 "Balaguer F, Link A, Lozano JJ, Cuatrecasas M, Nagasaka T, Boland CR, Goel A (2010) Epigenetic silencing of miR-137 is an early event in colorectal carcinogenesis. Cancer Res 70(16):6609–6618. doi: 10.1158/0008-5472.CAN-10-0622
"), [65](/article/10.1007/s00395-012-0245-9#ref-CR65 "Saito Y, Liang G, Egger G, Friedman JM, Chuang JC, Coetzee GA, Jones PA (2006) Specific activation of microRNA-127 with downregulation of the proto-oncogene BCL6 by chromatin-modifying drugs in human cancer cells. Cancer Cell 9(6):435–443. doi:
10.1016/j.ccr.2006.04.020
")\]. The aim of the present review was to underline the potential importance of this mechanism in cardiovascular disease. We therefore discussed several important manuscripts dealing with epigenetic silencing of microRNAs.Long non-coding RNAs
By definition non-coding transcripts with a length >200 nucleotides are considered as long non-coding RNAs (lncRNA) [[63](/article/10.1007/s00395-012-0245-9#ref-CR63 "Ponting CP, Oliver PL, Reik W (2009) Evolution and functions of long noncoding RNAs. Cell 136:629–641. doi: 10.1016/j.cell.2009.02.006
")\]. Depending on their position with regard to protein coding genes lncRNA can be categorized as: (1) sense or (2) antisense, (3) bidirectional, (4) intronic or (5) intergenic \[[63](/article/10.1007/s00395-012-0245-9#ref-CR63 "Ponting CP, Oliver PL, Reik W (2009) Evolution and functions of long noncoding RNAs. Cell 136:629–641. doi:
10.1016/j.cell.2009.02.006
")\]. LncRNAs are generally characterized by nuclear localization, low level of expression and sequence conservation and may be polyadenylated \[[36](/article/10.1007/s00395-012-0245-9#ref-CR36 "Kaneda R, Takada S, Yamashita Y, Choi YL, Nonaka-Sarukawa M, Soda M, Misawa Y, Isomura T, Shimada K, Mano H (2009) Genome-wide histone methylation profile for heart failure. Genes Cells 14(1):69–77. doi:
10.1111/j.1365-2443.2008.01252.x
")\]. It was recently shown that long intergenic non-coding RNAs (lincRNAs) significantly impact on the development of human diseases \[[13](/article/10.1007/s00395-012-0245-9#ref-CR13 "Calin GA, Liu CG, Ferracin M, Hyslop T, Spizzo R, Sevignani C, Fabbri M, Cimmino A, Lee EJ, Wojcik SE, Shimizu M, Tili E, Rossi S, Taccioli C, Pichiorri F, Liu X, Zupo S, Herlea V, Gramantieri L, Lanza G, Alder H, Rassenti L, Volinia S, Schmittgen TD, Kipps TJ, Negrini M, Croce CM (2007) Ultraconserved regions encoding ncRNAs are altered in human leukemias and carcinomas. Cancer Cell 12:215–229. doi:
10.1016/j.ccr.2007.07.027
"), [33](/article/10.1007/s00395-012-0245-9#ref-CR33 "Huarte M, Rinn JL (2010) Large non-coding RNAs: missing links in cancer? Hum Mol Genet 19(R2): R152–R161. doi:
10.1093/hmg/ddq353
"), [83](/article/10.1007/s00395-012-0245-9#ref-CR83 "Yu W, Gius D, Onyango P, Muldoon-Jacobs K, Karp J, Feinberg AP, Cui H (2008) Epigenetic silencing of tumour suppressor gene p15 by its antisense RNA. Nature 451:202–206. doi:
10.1038/nature06468
")\]. LincRNAs control gene expression by direct recruitment of histone modifying enzymes to chromatin, regulate dosage compensation, imprinting and developmental gene expression by establishing chromatin domains in an allele- and cell-type specific manner \[[63](/article/10.1007/s00395-012-0245-9#ref-CR63 "Ponting CP, Oliver PL, Reik W (2009) Evolution and functions of long noncoding RNAs. Cell 136:629–641. doi:
10.1016/j.cell.2009.02.006
")\]. LincRNAs are characterized by trimethylation of lysine 4 of histone H3 (H3K4me3) at their promoter and trimethylation of lysine 36 of histone H3 (H3K36me3) along the transcribed region \[[35](/article/10.1007/s00395-012-0245-9#ref-CR35 "Kaikkonen MU, Lam MT, Glass CK (2011) Non-coding RNAs as regulators of gene expression and epigenetics. Cardiovasc Res 90(3):430–440. doi:
10.1093/cvr/cvr097
")\]. Contrary to most lncRNAs, lincRNAs are higher conserved between different species \[[35](/article/10.1007/s00395-012-0245-9#ref-CR35 "Kaikkonen MU, Lam MT, Glass CK (2011) Non-coding RNAs as regulators of gene expression and epigenetics. Cardiovasc Res 90(3):430–440. doi:
10.1093/cvr/cvr097
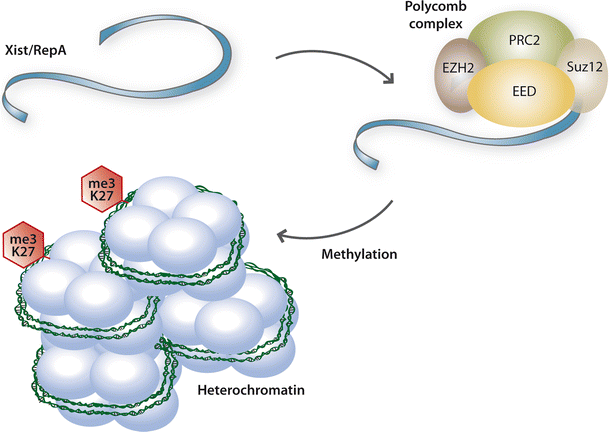
")\]. Long non-coding RNAs play a critical role in the regulation of imprinting, exemplified by the lincRNA _Air,_ and X-chromosome inactivation by _Xist_ (X-inactive specific transcript) (see Fig. [3](/article/10.1007/s00395-012-0245-9#Fig3)) \[[63](/article/10.1007/s00395-012-0245-9#ref-CR63 "Ponting CP, Oliver PL, Reik W (2009) Evolution and functions of long noncoding RNAs. Cell 136:629–641. doi:
10.1016/j.cell.2009.02.006
")\]. _Xist_ through its co-factor _RepA_ associates with the polycomb complex and thereby initiates epigenetic silencing during X-chromosome inactivation \[[84](/article/10.1007/s00395-012-0245-9#ref-CR84 "Zhao J, Sun BK, Erwin JA, Song JJ, Lee JT (2008) Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 322(5902):750–756. doi:
10.1126/science.1163045
")\]. _Air_ targets repressive histone-modifying activities through molecular interaction with specific chromatin domains to epigenetically silence transcription \[[55](/article/10.1007/s00395-012-0245-9#ref-CR55 "Nagano T, Mitchell JA, Sanz LA, Pauler FM, Ferguson-Smith AC, Feil R, Fraser P (2008) The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science 322:1717–1720. doi:
10.1126/science.1163802
")\]. This illustrates a first example of how a lincRNA may impact on epigenetic changes.Fig. 3
Long non-coding RNAs. Long non-coding RNAs (non-coding transcripts with a length >200 nucleotides) comprise a recently identified class of chromatin-modifiers. These RNAs play a critical role in the regulation of several important regulatory mechanisms including X-chromosome inactivation. During this process Xist (X-inactive-specific transcript) via its co-factor RepA associates with the polycomb complex and thereby initiates epigenetic silencing. A role for long non-coding RNAs in the cardiovascular system is highly likely, but so far unexplored. Ezh2 enhancer of zest homolog 2, PRC2 polycomb repressive complex 2, Suz12 suppressor of zest 12, EED embryonic ectodermal development
Epigenetic impact on cardiovascular development
Mathiyalagan and co-workers analyzed epigenetic modifications on the specific gene expression during physiological cardiovascular development. Increased ANP and BNP gene expression in the left ventricle (LV) are associated with increased histone acetylation and dimethylation. In addition, increased α-MHC gene expression in the LV as compared to the right ventricle (RV) was associated with H3K4 methylation. Chromatin immunopurification confirmed that the histone acetyltransferase p300 is recruited and increased on the active ANP and BNP genes in the left ventricle. The authors concluded that cardiac chambers are thus epigenetically distinguishable [[46](/article/10.1007/s00395-012-0245-9#ref-CR46 "Mathiyalagan P, Chang L, Du XJ, El-Osta A (2010) Cardiac ventricular chambers are epigenetically distinguishable. Cell Cycle 9(3):612–617. http://dx.doi.org/10.4161/cc.9.3.10612
")\].Another example of epigenetic importance shows that the loss of histone deacetylases 3 in neural crest results in perinatal lethality and cardiovascular abnormalities. This is associated with an impairment in aortic arch artery smooth muscle differentiation and a downregulation of the Notch ligand Jagged1 resulting in abnormalities of the cardiac outflow tract [[68](/article/10.1007/s00395-012-0245-9#ref-CR68 "Singh N, Trivedi CM, Lu M, Mullican SE, Lazar MA, Epstein JA (2011) Histone deacetylase 3 regulates smooth muscle differentiation in neural crest cells and development of the cardiac outflow Tract. Circ Res. doi: 10.1038/nrm3198
")\].Recently, the role of epigenetic modifications in the regulation of vascular development has also come into focus. Histone deacetylase 7 was shown to influence vascular development through an alteration of the extracellular matrix [[45](/article/10.1007/s00395-012-0245-9#ref-CR45 "Margariti A, Zampetaki A, Xiao Q, Zhou B, Karamariti E, Martin D, Yin X, Mayr M, Li H, Zhang Z, De Falco E, Hu Y, Cockerill G, Xu Q, Zeng L (2010) Histone deacetylase 7 controls endothelial cell growth through modulation of beta-catenin. Circ Res 106:1202–1211. doi: 10.1161/CIRCRESAHA.109.213165
"), [50](/article/10.1007/s00395-012-0245-9#ref-CR50 "Mottet D, Bellahcene A, Pirotte S, Waltregny D, Deroanne C, Lamour V, Lidereau R, Castronovo V (2007) Histone deacetylase 7 silencing alters endothelial cell migration, a key step in angiogenesis. Circ Res 101:1237–1246. doi:
10.1161/CIRCRESAHA.107.149377
")\]. The flow-dependent regulation of gene expression during vasculogenesis was shown to be influenced by HDACs modulating in vitro experiments involving shear stress \[[34](/article/10.1007/s00395-012-0245-9#ref-CR34 "Illi B, Nanni S, Scopece A, Farsetti A, Biglioli P, Capogrossi MC, Gaetano C (2003) Shear stress-mediated chromatin remodeling provides molecular basis for flow-dependent regulation of gene expression. Circ Res 25 93(2):155–161. doi:
10.1161/01.RES.0000080933.82105.29
")\]. In addition, histone acetylation is crucial for vascular gene expression in endothelial and smooth muscle cells (SMC) \[[47](/article/10.1007/s00395-012-0245-9#ref-CR47 "McDonald OG, Owens GK (2007) Programming smooth muscle plasticity with chromatin dynamics. Circ Res 100:1428–1441. doi:
10.1161/01.RES.0000266448.30370.a0
"), [69](/article/10.1007/s00395-012-0245-9#ref-CR69 "Spin JM, Quertermous T, Tsao PS (2010) Chromatin remodeling pathways in smooth muscle cell differentiation, and evidence for an integral role for p300. PLoS One 5(12):e14301. doi:
10.1371/journal.pone.0014301
")\]. The histone acetyltransferase p300 significantly impacts on the differentiation of vascular SMCs underlining that chromatin remodeling is important for SMC phenotypic switching \[[69](/article/10.1007/s00395-012-0245-9#ref-CR69 "Spin JM, Quertermous T, Tsao PS (2010) Chromatin remodeling pathways in smooth muscle cell differentiation, and evidence for an integral role for p300. PLoS One 5(12):e14301. doi:
10.1371/journal.pone.0014301
")\].DNA methylation and cardiovascular disease
Phenotypic alterations during adulthood may be explained by exposure to adverse environmental conditions in utero. Ground-breaking work elucidating this mechanism in humans came from Heijmans and co-workers [[29](/article/10.1007/s00395-012-0245-9#ref-CR29 "Heijmans BT, Tobi EW, Stein AD, Putter H, Blauw GJ, Susser ES, Slagboom PE, Lumey LH (2008) Persistent epigenetic differences associated with prenatal exposure to famine in humans. Proc Natl Acad Sci USA 105:17046–17049. doi: 10.1073/pnas.0806560105
")\], who studied individuals prenatally exposed to famine during the Dutch Hunger Winter in 1944/45\. Compared to siblings of the same sex these individuals displayed an altered genetic methylation pattern. DNA methylation of the imprinted insulin-like growth factor II (_IGF2_) gene was shown to be lower. The prevalence of obesity and coronary heart disease in these individuals was higher than that of adults born before or conceived after that period \[[58](/article/10.1007/s00395-012-0245-9#ref-CR58 "Painter RC, Roseboom TJ, Bleker OP (2005) Prenatal exposure to the Dutch famine and disease in later life: an overview. Reprod Toxicol 20:345–352. doi:
10.1016/j.reprotox.2005.04.005
")\]. Low birth weight was shown to be an independent risk factor for coronary heart disease \[[25](/article/10.1007/s00395-012-0245-9#ref-CR25 "Godfrey KM, Barker DJ (2001) Fetal programming and adult health. Pub Health Nutr 4:611–624. doi:
10.1079/PHN2001145
")\]. These findings are corroborated by experimental studies, in which the critical impact of maternal diet during pregnancy on the pattern of DNA methylation of genes pertaining to blood pressure control was shown \[[9](/article/10.1007/s00395-012-0245-9#ref-CR9 "Bogdarina I, Welham S, King PJ, Burns SP, Clark AJ (2007) Epigenetic modification of the reninangiotensin system in the fetal programming of hypertension. Circ Res 100:520–526. doi:
10.1161/01.RES.0000258855.60637.58
")\].The prenatal exposure to tobacco severely alters the methylation pattern of specific genes and may increase the risk of cardiovascular disease later in the child’s life. While the methylation of the DNA repetitive element AluYb8 is generally significantly associated with prenatal tobacco exposure, differences in smoking-related effects on LINE1 methylation are dependent upon a lack in a functional detoxifying enzyme glutathione _S_-transferase P (GSTP1) [[10](/article/10.1007/s00395-012-0245-9#ref-CR10 "Breton CV, Byun HM, Wenten M, Pan F, Yang A, Gilliland FD (2009) Prenatal tobacco smoke exposure affects global and gene-specific DNA methylation. Am J Respir Crit Care Med 180(5):462–467. doi: 10.1164/rccm.200901-0135OC
")\]. Thus, variants in detoxification genes may modulate the effects of in utero exposure through epigenetic mechanisms.In patients with end-stage heart failure differential DNA methylation was shown to correlate with the gene expression of angiogenic factors, namely the PECAM1, ARHGAP24 and AMOTL2 gene [[51](/article/10.1007/s00395-012-0245-9#ref-CR51 "Movassagh M, Choy MK, Goddard M, Bennett MR, Down TA, Foo RS (2010) Differential DNA methylation correlates with differential expression of angiogenic factors in human heart failure. PLoS One 5(1):e8564. doi: 10.1371/journal.pone.0008564
")\].Zhang and co-workers [[85](/article/10.1007/s00395-012-0245-9#ref-CR85 "Zhang H, Darwanto A, Linkhart TA, Sowers LC, Zhang L (2007) Maternal cocaine administration causes an epigenetic modification of protein kinase Cepsilon gene expression in fetal rat heart. Mol Pharmacol 71(5):1319–1328. doi: 10.1124/mol.106.032011
")\] demonstrated the detrimental effect of prenatal cocaine exposure regarding the methylation status of specific genes. Cocaine exposure induces a decrease in cardiac PKC-epsilon expression through methylation of CpGs in the activator protein 1 (AP-1) binding site of the PKC-epsilon gene. Previous in vitro and in vivo studies demonstrated that PKC-epsilon has a critical cardioprotective role during cardiac ischemia and reperfusion injury \[[14](/article/10.1007/s00395-012-0245-9#ref-CR14 "Chen L, Hahn H, Wu G, Chen CH, Liron T, Schechtman D, Cavallaro G, Banci L, Guo Y, Bolli R, Dorn GW 2nd, Mochly-Rosen D (2001) Opposing cardioprotective actions and parallel hypertrophic effects of delta PKC and epsilon PKC. Proc Natl Acad Sci USA 98:11114–11119. doi:
10.1073/pnas.191369098
"), [24](/article/10.1007/s00395-012-0245-9#ref-CR24 "Gray MO, Zhou HZ, Schafhalter-Zoppoth I, Zhu P, Mochly-Rosen D, Messing RO (2004) Preservation of base-line hemodynamic function and loss of inducible cardioprotection in adult mice lacking protein kinase Cε. J Biol Chem 279:3596–3604. doi:
10.1074/jbc.M311459200
"), [32](/article/10.1007/s00395-012-0245-9#ref-CR32 "Heusch G, Boengler K, Schulz R (2010) Cardioprotection: nitric oxide, protein kinases, and mitochondria. Circulation 118(19):1915–1919. doi:
10.1161/CIRCULATIONAHA.108.805242
"), [54](/article/10.1007/s00395-012-0245-9#ref-CR54 "Murriel CL, Mochly-Rosen D (2003) Opposing roles of delta and epsilon PKC in cardiac ischemia and reperfusion: targeting the apoptotic machinery. Arch Biochem Biophys 420:246–254. doi:
10.1016/j.abb.2003.08.038
")\]. The results of this study indicate that prenatal cocaine abuse might alter the cardiac response to ischemia reperfusion injury through methylation of the AP-1 binding site and subsequent repression of the PKC-epsilon gene. The PKC-epsilon gene is also targeted during chronic prenatal hypoxia \[[61](/article/10.1007/s00395-012-0245-9#ref-CR61 "Patterson AJ, Chen M, Xue Q, Xiao D, Zhang L (2010) Chronic prenatal hypoxia induces epigenetic programming of PKC{epsilon} gene repression in rat hearts. Circ Res 107(3):365–373. doi:
10.1161/CIRCRESAHA.110.221259
")\]. Chronic prenatal hypoxia leads to a decrease in PKC-epsilon expression due to methylation of the specificity protein 1 (SP1) binding sites in the PKC-epsilon promoter in fetal hearts. Methylation of the SP1 site is gender-specific with more males being affected. This difference is most likely due to the higher abundance of estrogen receptor alpha and beta isoforms found in females compared to males, since both estrogen receptor alpha and beta interact with the SP1 binding site in the fetal heart.Histone methylation and cardiovascular disease
To further address the epigenetic response to cardiac ischemia/reperfusion Das and co-workers [[15](/article/10.1007/s00395-012-0245-9#ref-CR15 "Das M, Das S, Lekli I, Das DK (2011) Caveolin induces cardioprotection through epigenetic regulation. J Cell Mol Med (in press). doi: 10.1111/j.1582-4934.2011.01372.x
")\] analyzed caveolin knockout mice. Caveolins are an integral part of lipid rafts, a sub-compartment of the plasma membrane. Recent work suggests an important role for caveolins during cardiac ischemia reperfusion injury \[[75](/article/10.1007/s00395-012-0245-9#ref-CR75 "Tsutsumi YM, Horikawa YT, Jennings MM, Kidd MW, Niesman IR, Yokoyama U, Head BP, Hagiwara Y, Ishikawa Y, Miyanohara A, Patel PM, Insel PA, Patel HH, Roth DM (2008) Cardiac-specific overexpression of caveolin-3 induces endogenous cardiac protection by mimicking ischemic preconditioning. Circulation 118(19):1979–1988. doi:
10.1161/CIRCULATIONAHA.108.788331
")\]. Histone methylation was shown to be increased and associated with increased histone methyl-transferase G9a protein levels and histone decaetylase activity in caveolin knockout mice following ischemia reperfusion injury. In addition, the translocation of Foxo-3a to the nucleus and expression of sirtuin-1 was reduced in caveolin knockout mice. The cardioprotective function of caveolins was further confirmed by reduced ventricular function, increased cardiomyocyte apoptosis, increased expression of Janus kinase and Bax and decreased expression of phospho-AMPK, phospho-AKT and Bcl-2 in caveolin knockout mice. These findings underline the importance of caveolins for cardioprotection and their association with epigenetic mechanisms during cardiac ischemia reperfusion injury.Depending on the methylation of specific lysine or arginine residues histone methylation can either be associated with transcriptional repression or activation [[44](/article/10.1007/s00395-012-0245-9#ref-CR44 "Litt MD, Simpson M, Gaszner M, Allis CD, Felsenfeld G (2001) Correlation between histone lysine methylation and developmental changes at the chicken betaglobin locus. Science 293(5539):2453–2455. doi: 10.1126/science.1064413
"), [66](/article/10.1007/s00395-012-0245-9#ref-CR66 "Santos-Rosa H, Schneider R, Bannister AJ, Sherriff J, Bernstein BE, Emre NC, Schreiber SL, Mellor J, Kouzarides T (2002) Active genes are tri-methylated at K4 of histone H3. Nature 419(6905):407–411. doi:
10.1038/nature01080
")\]. A genome-wide histone methylation profile revealed a differential marking of trimethylation of H3K4 and H3K9 (H3K4me3 and H3K9me3) in cardiomyocytes during development of heart failure in both animal models and human \[[36](/article/10.1007/s00395-012-0245-9#ref-CR36 "Kaneda R, Takada S, Yamashita Y, Choi YL, Nonaka-Sarukawa M, Soda M, Misawa Y, Isomura T, Shimada K, Mano H (2009) Genome-wide histone methylation profile for heart failure. Genes Cells 14(1):69–77. doi:
10.1111/j.1365-2443.2008.01252.x
")\]. Histone H3 lysine 4 methylation (H3K4me) marks transcriptional activation \[[66](/article/10.1007/s00395-012-0245-9#ref-CR66 "Santos-Rosa H, Schneider R, Bannister AJ, Sherriff J, Bernstein BE, Emre NC, Schreiber SL, Mellor J, Kouzarides T (2002) Active genes are tri-methylated at K4 of histone H3. Nature 419(6905):407–411. doi:
10.1038/nature01080
")\]. Stein and co-workers \[[70](/article/10.1007/s00395-012-0245-9#ref-CR70 "Stein AB, Jones TA, Herron TJ, Patel SR, Day SM, Noujaim SF, Milstein ML, Klos M, Furspan PB, Jalife J, Dressler GR (2011) Loss of H3K4 methylation destabilizes gene expression patterns and physiological functions in adult murine cardiomyocytes. J Clin Invest 121(7):2641–2650. doi:
10.1172/JCI44641
")\] recently analyzed the effect of reducing histone H3 lysine 4 methylation on differentiated cardiomyocytes. PAX interacting protein 1 (PTIP) protein, an essential co-factor of the H3K4me complex, was deleted through a cardiac-specific, inducible knockout model. This resulted in reduced H3K4me impacting on downstream signaling cascades in cardiomyocytes. The gene encoding for Kv channel-interacting protein 2 (_Kcnip2_) was shown to be down-regulated by trimethylation of lysine 4 of histone H3 (H3K4me3). Regulation of Kcnip2 resulted in several impaired cellular functions, e.g., sodium and L-type calcium current, action potential upstroke velocity as well as enhanced action potential duration. PTIP mutant cardiomyocytes were more sensitive to premature ventricular complexes upon stimulation with isoproterenol and caffeine. The results of this study clearly underline the significance of histone methylation for the physiological function of cardiomyocytes.To elucidate the role of H3K9 methylation on the development of cardiac hypertrophy and failure Zhang et al. [[86](/article/10.1007/s00395-012-0245-9#ref-CR86 "Zhang QJ, Chen HZ, Wang L, Liu DP, Hill JA, Liu ZP (2011) The histone trimethyllysine demethylase JMJD2A promotes cardiac hypertrophy in response to hypertrophic stimuli in mice. J Clin Invest 121(6):2447–2456. doi: 10.1172/JCI46277
")\] studied the involvement of the histone trimethyllysine demethylase JMJD2A. JMJD2 proteins are lysine trimethyl-specific histone demethylases that catalyze the demethylation of trimethylated H3K9 (H3K9me3) and H3K36 (H3K36me3) \[[86](/article/10.1007/s00395-012-0245-9#ref-CR86 "Zhang QJ, Chen HZ, Wang L, Liu DP, Hill JA, Liu ZP (2011) The histone trimethyllysine demethylase JMJD2A promotes cardiac hypertrophy in response to hypertrophic stimuli in mice. J Clin Invest 121(6):2447–2456. doi:
10.1172/JCI46277
")\]. JMJD2A was shown to mediate cardiac hypertrophy through regulation of its target _FHL1_ (four-and-a-half LIM domains 1). Zhang and co-workers thus provide a novel mechanistic explanation for the epigenetic regulation of cardiac hypertrophy.Recently, an epigenetic silencing of the endothelial nitric oxide synthase (eNOS) promotor through methylation of the lysine residue 27 on histone 3 (H3K27me3) was shown to control angiogenesis [[57](/article/10.1007/s00395-012-0245-9#ref-CR57 "Ohtani K, Vlachojannis GJ, Koyanagi M, Boeckel JN, Urbich C, Farcas R, Bonig H, Marquez VE, Zeiher AM, Dimmeler S (2011) Epigenetic regulation of endothelial lineage committed genes in pro-angiogenic hematopoietic and endothelial progenitor cells. Circ Res doi. doi: 10.1161/CIRCRESAHA.111.247304
")\]. During hypoxia eNOS expression was increased through reduction of repressive H3K27me3\. This reduction was associated with an increase of the histone demethylase Jmjd3 (see Fig. [2](/article/10.1007/s00395-012-0245-9#Fig2)).Movassagh and co-workers [[52](/article/10.1007/s00395-012-0245-9#ref-CR52 "Movassagh M, Choy MK, Knowles DA, Cordeddu L, Haider S, Down T, Siggens L, Vujic A, Simeoni I, Penkett C, Goddard M, Lio P, Bennett MR, Foo RS (2011) Distinct epigenomic features in end-stage failing human hearts. Circulation. doi: 10.1161/CIRCULATIONAHA.111.040071
")\] found the epigenomic pattern in patients with end-stage heart failure to be highly altered compared to control patients. Specifically, differential DNA methylation and trimethylation of lysine 36 of histone 3 (H3K36me3) was found to be influenced in RNA transcripts encoded by the DUX4 locus \[[68](/article/10.1007/s00395-012-0245-9#ref-CR68 "Singh N, Trivedi CM, Lu M, Mullican SE, Lazar MA, Epstein JA (2011) Histone deacetylase 3 regulates smooth muscle differentiation in neural crest cells and development of the cardiac outflow Tract. Circ Res. doi:
10.1038/nrm3198
")\], which codes for a double homeobox transcription factor and has been implicated to play a role in facioscapulohumeral muscular dystrophy \[[19](/article/10.1007/s00395-012-0245-9#ref-CR19 "Dixit M, Ansseau E, Tassin A, Winokur S, Shi R, Qian H, Sauvage S, Matteotti C, van Acker AM, Leo O, Figlewicz D, Barro M, Laoudj-Chenivesse D, Belayew A, Coppee F, Chen YW (2007) DUX4, a candidate gene of facioscapulohumeral muscular dystrophy, encodes a transcriptional activator of PITX1. Proc Natl Acad Sci USA 104:18157–18162. doi:
10.1073/pnas.0708659104
"), [62](/article/10.1007/s00395-012-0245-9#ref-CR62 "Pearson CE (2010) FSHD: a repeat contraction disease finally ready to expand (our understanding of its pathogenesis). PLoS Genet 6:e1001180. doi:
10.1371/journal.pgen.1001180
")\].MicroRNAs and cardiovascular disease
MicroRNAs fundamentally are involved in and have impact on cardiovascular disease. In addition, microRNA has been shown to control cardiovascular differentiation [[56](/article/10.1007/s00395-012-0245-9#ref-CR56 "Ohtani K, Dimmeler S (2011) Control of cardiovascular differentiation by microRNAs. Basic Res Cardiol 106(1):5–11. doi: 10.1007/s00395-010-0139-7
")\]. An array of studies elucidated a disease-specific role for selected miRNAs (for instance, miR-21 and miR-29 in cardiac fibrosis) \[[73](/article/10.1007/s00395-012-0245-9#ref-CR73 "Thum T, Gross C, Fiedler J, Fischer T, Kissler S, Bussen M, Galuppo P, Just S, Rottbauer W, Frantz S, Castoldi M, Soutschek J, Koteliansky V, Rosenwald A, Basson MA, Licht JD, Pena JT, Rouhanifard SH, Muckenthaler MU, Tuschl T, Martin GR, Bauersachs J, Engelhardt S (2008) MicroRNA-21 contributes to myocardial disease by stimulating MAP kinase signalling in fibroblasts. Nature 456:980–984. doi:
10.1038/nature07511
"), [78](/article/10.1007/s00395-012-0245-9#ref-CR78 "van Rooij E, Sutherland LB, Thatcher JE, DiMaio JM, Naseem RH, Marshall WS, Hill JA, Olson EN (2008) Dysregulation of microRNAs after myocardial infarction reveals a role of miR-29 in cardiac fibrosis. Proc Natl Acad Sci USA 105:13027–13032. doi:
10.1073/pnas.0805038105
")\]. Moreover, microRNA signatures in total peripheral blood have been identified as biomarkers of myocardial infarction \[[48](/article/10.1007/s00395-012-0245-9#ref-CR48 "Meder B, Keller A, Vogel B, Haas J, Sedaghat-Hamedani F, Kayvanpour E, Just S, Borries A, Rudloff J, Leidinger P, Meese E, Katus HA, Rottbauer W (2011) MicroRNA signatures in total peripheral blood as novel biomarkers for acute myocardial infarction. Basic Res Cardiol 106(1):13–23. doi:
10.1002/ijc.26419
")\]. Deregulation of miRNAs results in a profound disturbance of downstream gene networks and signaling cascades culminating in disease initiation and/or progression \[[74](/article/10.1007/s00395-012-0245-9#ref-CR74 "Thum T, Galuppo P, Wolf C, Fiedler J, Kneitz S, van Laake LW, Doevendans PA, Mummery CL, Borlak J, Haverich A, Gross C, Engelhardt S, Ertl G, Bauersachs J (2007) MicroRNAs in the human heart: a clue to fetal gene reprogramming in heart failure. Circulation 116:258–267. doi:
10.1161/CIRCULATIONAHA.107.687947
")\]. The role of miRNAs in the initiation of cardiovascular disease has previously been reviewed (see \[[28](/article/10.1007/s00395-012-0245-9#ref-CR28 "Hartmann D, Thum T (2011) MicroRNAs and vascular (dys)function. Vascul Pharmacol (in press). doi:
10.1016/j.vph.2011.07.005
"), [76](/article/10.1007/s00395-012-0245-9#ref-CR76 "Urbich C, Kuehbacher A, Dimmeler S (2008) Role of microRNAs in vascular diseases, inflammation, and angiogenesis. Cardiovasc Res 79(4):581–588. doi:
10.1093/cvr/cvn156
")\]). Thus, the present review focuses on the epigenetic regulation of microRNAs during pathophysiological processes. We here outline DNA methylation of specific miRNA genes and its impact on the expression levels of miRNAs. This mechanism represents a fascinating novel regulation of microRNA expression, which has not been studied in cardiovascular disease so far. We therefore shortly mention and discuss cancer-specific epigenetic regulation of miRNA genes to spark interest in this new field.Several studies have addressed the effect of tumor-specific alterations in DNA methylation of miRNA genes on the expression of mature miRNAs. Saito and co-workers [[65](/article/10.1007/s00395-012-0245-9#ref-CR65 "Saito Y, Liang G, Egger G, Friedman JM, Chuang JC, Coetzee GA, Jones PA (2006) Specific activation of microRNA-127 with downregulation of the proto-oncogene BCL6 by chromatin-modifying drugs in human cancer cells. Cancer Cell 9(6):435–443. doi: 10.1016/j.ccr.2006.04.020
")\] evaluated the specific miRNA signature of T24 human bladder cancer cells after simultaneous treatment with the DNA methyltransferase inhibitor 5-aza-2′-deoxycytidine (DAC) and the histone deacetylase (HDAC) inhibitor 4-phenylbutyric acid (PBA). Genome-wide expression analysis revealed that miR-127 was strongly up-regulated following treatment with the chromatin-modifying drugs. Importantly, the corresponding gene was found to be embedded in a CpG island. MiR-127 is physiologically expressed as part of a miRNA cluster with miR-136, -431, -432, and -433 in normal tissues and cultured fibroblasts. In primary tumors and various cancer cell lines it is down-regulated. Simultaneous treatment with the chromatin-modifying drugs resulted in an induction of miR-127\. The results of this study suggest that it is subject to epigenetic silencing. Its role in cardiovascular disease, e.g., fibrosis, remains to be elucidated.The epigenetic regulation of miR-137 and its contribution to colorectal carcinogenesis was shown using several colorectal cancer cell lines and colorectal tissues [[2](/article/10.1007/s00395-012-0245-9#ref-CR2 "Balaguer F, Link A, Lozano JJ, Cuatrecasas M, Nagasaka T, Boland CR, Goel A (2010) Epigenetic silencing of miR-137 is an early event in colorectal carcinogenesis. Cancer Res 70(16):6609–6618. doi: 10.1158/0008-5472.CAN-10-0622
")\]. The promoter region of miR-137 was identified to be extensively methylated in cancer cell lines, which was reversible following DAC treatment. The level of methylation was significantly higher in colorectal cancer tissues compared with their corresponding histologically normal mucosa. The expression of miR-137 in cancer tissue was significantly down-regulated. Over-expression of miR-137 in cancer cell lines resulted in a significant reduction of cell proliferation. Lysine (K)-specific demethylase 1A (_LSD1_) was identified as a target of miR-137 in colorectal cancer cell lines using luciferase gene reporter assays. Thus, in colorectal cancer miR-137 might act as a tumor suppressor. During colon cancerogenesis it is frequently silenced by promoter hypermethylation.Currently, the epigenetic regulation of microRNA expression through methylation of CpG islands in the promoter region of genes encoding for specific microRNAs has not been evaluated in the setting of cardiovascular disease. The results of the aforementioned studies suggest that this mechanism may also play a pivotal role in the initiation of cardiovascular complications. Future studies are warranted to address this phenomenon.
Long non-coding RNA and cardiovascular disease
The characterization of lncRNAs pertaining to the development of cardiovascular disease is still in its infancy. Nevertheless, the elucidation of “cardiovascular” lncRNAs holds enormous future promise given their regulatory function in health and disease and there soon will be more insight in this emerging topic.
In clear cell renal carcinoma, a natural antisense long non-coding transcript (aHIF) that is complementary to the 3′-untranslated region of hypoxia-inducible factor 1a (HIF-1a) messenger RNA has been discovered [[72](/article/10.1007/s00395-012-0245-9#ref-CR72 "Thrash-Bingham CA, Tartof KD (1999) aHIF: a natural antisense transcript overexpressed in human renal cancer and during hypoxia. J Natl Cancer Inst 91:143–151. doi: 10.1093/jnci/91.2.143
")\]. It modulates the stability of HIF-1a mRNA, thereby impacting on the regulation of angiogenesis.Robb and co-workers [[64](/article/10.1007/s00395-012-0245-9#ref-CR64 "Robb GB, Carson AR, Tai SC, Fish JE, Singh S, Yamada T, Scherer SW, Nakabayashi K, Marsden PA (2004) Post-transcriptional regulation of endothelial nitric-oxide synthase by an overlapping antisense mRNA transcript. J Biol Chem 279(36):37982–37996. doi: 10.1074/jbc.M400271200
")\] discovered an antisense transcript to endothelial nitric oxide synthase (eNOS), termed sONE. RNA interference-mediated inhibition of sONE expression in vascular smooth muscle cells increased eNOS expression. Overexpression of sONE in endothelial cells reduced eNOS expression. The antisense lncRNA sONE might thus contribute to the post-transcriptional regulation of endothelial cell-specific gene expression.Recently, an antisense long non-coding transcript to tyrosine kinase containing immunoglobulin and epidermal growth factor homology domain-1 (tie-1), tie-1AS, was uncovered [[43](/article/10.1007/s00395-012-0245-9#ref-CR43 "Li K, Blum Y, Verma A, Liu Z, Pramanik K, Leigh NR, Chun CZ, Samant GV, Zhao B, Garnaas MK, Horswill MA, Stanhope SA, North PE, Miao RQ, Wilkinson GA, Affolter M, Ramchandran R (2010) A noncoding antisense RNA in tie-1 locus regulates tie-1 function in vivo. Blood 115(1):133–139. doi: 10.1182/blood-2009-09-242180
")\]. The tie-1AS lncRNA was shown to target the tie-1 mRNA in vivo, thereby regulating tie-1 transcript levels. This resulted in abnormalities in endothelial cell contact junctions suggesting a role in the maintenance of vascular homeostasis.Single nucleotide polymorphisms (SNPs) on chromosome 9p21.3 near the INK4/ARF (CDKN2a/b) locus correlate with susceptibility to cardiovascular disease [[30](/article/10.1007/s00395-012-0245-9#ref-CR30 "Helgadottir A, Thorleifsson G, Manolescu A, Gretarsdottir S, Blondal T, Jonasdottir A, Jonasdottir A, Sigurdsson A, Baker A, Palsson A, Masson G, Gudbjartsson DF, Magnusson KP, Andersen K, Levey AI, Backman VM, Matthiasdottir S, Jonsdottir T, Palsson S, Einarsdottir H, Gunnarsdottir S, Gylfason A, Vaccarino V, Hooper WC, Reilly MP, Granger CB, Austin H, Rader DJ, Shah SH, Quyyumi AA, Gulcher JR, Thorgeirsson G, Thorsteinsdottir U, Kong A, Stefansson K (2007) A common variant on chromosome 9p21 affects the risk of myocardial infarction. Science 316:1491–1493. doi: 10.1126/science.1142842
"), [31](/article/10.1007/s00395-012-0245-9#ref-CR31 "Helgadottir A, Thorleifsson G, Magnusson KP, Grétarsdottir S, Steinthorsdottir V, Manolescu A, Jones GT, Rinkel GJ, Blankensteijn JD, Ronkainen A, Jääskeläinen JE, Kyo Y, Lenk GM, Sakalihasan N, Kostulas K, Gottsäter A, Flex A, Stefansson H, Hansen T, Andersen G, Weinsheimer S, Borch-Johnsen K, Jorgensen T, Shah SH, Quyyumi AA, Granger CB, Reilly MP, Austin H, Levey AI, Vaccarino V, Palsdottir E, Walters GB, Jonsdottir T, Snorradottir S, Magnusdottir D, Gudmundsson G, Ferrell RE, Sveinbjornsdottir S, Hernesniemi J, Niemelä M, Limet R, Andersen K, Sigurdsson G, Benediktsson R, Verhoeven EL, Teijink JA, Grobbee DE, Rader DJ, Collier DA, Pedersen O, Pola R, Hillert J, Lindblad B, Valdimarsson EM, Magnadottir HB, Wijmenga C, Tromp G, Baas AF, Ruigrok YM, van Rij AM, Kuivaniemi H, Powell JT, Matthiasson SE, Gulcher JR, Thorgeirsson G, Kong A, Thorsteinsdottir U, Stefansson K (2008) The same sequence variant on 9p21 associates with myocardial infarction, abdominal aortic aneurysm and intracranial aneurysm. Nat Genet 40:217–224. doi:
10.1038/ng.72
")\]. The SNPs most strongly associated with cardiovascular disease are located \~120 kb from the nearest coding gene within a long non-coding RNA (ncRNA) known as ANRIL (CDKN2BAS) \[[11](/article/10.1007/s00395-012-0245-9#ref-CR11 "Broadbent HM, Peden JF, Lorkowski S, Goel A, Ongen H, Green F, Clarke R, Collins R, Franzosi MG, Tognoni G, Seedorf U, Rust S, Eriksson P, Hamsten A, Farrall M, Watkins H, PROCARDIS Consortium (2008) Susceptibility to coronary artery disease and diabetes is encoded by distinct, tightly linked SNPs in the ANRIL locus on chromosome 9p. Hum Mol Genet 17:806–814. doi:
10.1093/hmg/ddm352
")\]. ANRIL is an antisense non-coding RNA in the INK4 Locus, which was first discovered in a genetic analysis of familial melanoma patients with neural system tumors \[[60](/article/10.1007/s00395-012-0245-9#ref-CR60 "Pasmant E, Laurendeau I, Héron D, Vidaud M, Vidaud D, Bièche I (2007) Characterization of a germ-line deletion, including the entire INK4/ARF locus, in a melanoma-neural system tumor family: identification of ANRIL, an antisense noncoding RNA whose expression coclusters with ARF. Cancer Res 67:3963–3969. doi:
10.1158/0008-5472.CAN-06-2004
")\]. It recruits polycomb repressive complexes to induce epigenetic transcriptional repression \[[82](/article/10.1007/s00395-012-0245-9#ref-CR82 "Yap KL, Li S, Muñoz-Cabello AM, Raguz S, Zeng L, Mujtaba S, Gil J, Walsh MJ, Zhou MM (2010) Molecular interplay of the noncoding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol Cell 38(5):662–674. doi:
10.1016/j.molcel.2010.03.021
")\]. The identification of altered expression patterns and function of individual lincRNAs will be a fascinating future avenue to explore in the setting of numerous cardiovascular diseases.Conclusions
Epigenetic modifications such as DNA methylation, histone modifications and RNA-based mechanisms represent the molecular substrate for detrimental environmental stimuli and may lead to disease initiation including cardiovascular disease. Epigenetic changes may be targeted and influenced in the future by pharmacological and/or genetic interventions. For instance, the demethylating agent 5-aza-2′-deoxycytidine has been used in several experimental studies [[61](/article/10.1007/s00395-012-0245-9#ref-CR61 "Patterson AJ, Chen M, Xue Q, Xiao D, Zhang L (2010) Chronic prenatal hypoxia induces epigenetic programming of PKC{epsilon} gene repression in rat hearts. Circ Res 107(3):365–373. doi: 10.1161/CIRCRESAHA.110.221259
"), [65](/article/10.1007/s00395-012-0245-9#ref-CR65 "Saito Y, Liang G, Egger G, Friedman JM, Chuang JC, Coetzee GA, Jones PA (2006) Specific activation of microRNA-127 with downregulation of the proto-oncogene BCL6 by chromatin-modifying drugs in human cancer cells. Cancer Cell 9(6):435–443. doi:
10.1016/j.ccr.2006.04.020
")\].Additional studies are clearly needed to further elucidate how epigenetic phenomena impact on the development of cardiovascular disease. The number of studies on the role of long non-coding RNAs and microRNAs, for which methylated CpG islands have been identified, is presently rather limited. However, studies pertaining to various forms of cancer already suggest the major impact of these processes on cellular homeostasis and the development of disease. Manipulating the expression of long non-coding RNAs and “methylated” microRNAs may thus represent an interesting future therapeutic direction.
References
- Ambros V (2004) The functions of animal microRNAs. Nature 431:350–355. doi:10.1038/nature02871
Article PubMed CAS Google Scholar - Balaguer F, Link A, Lozano JJ, Cuatrecasas M, Nagasaka T, Boland CR, Goel A (2010) Epigenetic silencing of miR-137 is an early event in colorectal carcinogenesis. Cancer Res 70(16):6609–6618. doi:10.1158/0008-5472.CAN-10-0622
Article PubMed CAS Google Scholar - Bartel DP (2003) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297. doi:10.1016/S0092-8674(04)00045-5
Article Google Scholar - Bauersachs J, Thum T (2011) Biogenesis and regulation of cardiovascular microRNAs. Circ Res 109(3):334–347. doi:10.1161/CIRCRESAHA.110.228676
Article PubMed CAS Google Scholar - Bhutani N, Burns DM, Blau HM (2011) DNA demethylation dynamics. Cell 146(6):866–872. doi:10.1016/j.cell.2011.08.042
Article PubMed CAS Google Scholar - Bird A (2007) Perceptions of epigenetics. Nature 447:396–398. doi:10.1038/nature05913
Article PubMed CAS Google Scholar - Bird A (1986) CpG-rich islands and the function of DNA methylation. Nature 321:209–213. doi:10.1038/321209a0
Article PubMed CAS Google Scholar - Bird AP (1980) DNA methylation and the frequency of CpG in animal DNA. Nucleic Acids Res 8:1499–1504. doi:10.1093/nar/8.7.1499
Article PubMed CAS Google Scholar - Bogdarina I, Welham S, King PJ, Burns SP, Clark AJ (2007) Epigenetic modification of the reninangiotensin system in the fetal programming of hypertension. Circ Res 100:520–526. doi:10.1161/01.RES.0000258855.60637.58
Article PubMed CAS Google Scholar - Breton CV, Byun HM, Wenten M, Pan F, Yang A, Gilliland FD (2009) Prenatal tobacco smoke exposure affects global and gene-specific DNA methylation. Am J Respir Crit Care Med 180(5):462–467. doi:10.1164/rccm.200901-0135OC
Article PubMed CAS Google Scholar - Broadbent HM, Peden JF, Lorkowski S, Goel A, Ongen H, Green F, Clarke R, Collins R, Franzosi MG, Tognoni G, Seedorf U, Rust S, Eriksson P, Hamsten A, Farrall M, Watkins H, PROCARDIS Consortium (2008) Susceptibility to coronary artery disease and diabetes is encoded by distinct, tightly linked SNPs in the ANRIL locus on chromosome 9p. Hum Mol Genet 17:806–814. doi:10.1093/hmg/ddm352
Article PubMed CAS Google Scholar - Cai X, Hagedorn CH, Cullen BR (2004) Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA 10:1957–1966. doi:10.1261/rna.7135204
Article PubMed CAS Google Scholar - Calin GA, Liu CG, Ferracin M, Hyslop T, Spizzo R, Sevignani C, Fabbri M, Cimmino A, Lee EJ, Wojcik SE, Shimizu M, Tili E, Rossi S, Taccioli C, Pichiorri F, Liu X, Zupo S, Herlea V, Gramantieri L, Lanza G, Alder H, Rassenti L, Volinia S, Schmittgen TD, Kipps TJ, Negrini M, Croce CM (2007) Ultraconserved regions encoding ncRNAs are altered in human leukemias and carcinomas. Cancer Cell 12:215–229. doi:10.1016/j.ccr.2007.07.027
Article PubMed CAS Google Scholar - Chen L, Hahn H, Wu G, Chen CH, Liron T, Schechtman D, Cavallaro G, Banci L, Guo Y, Bolli R, Dorn GW 2nd, Mochly-Rosen D (2001) Opposing cardioprotective actions and parallel hypertrophic effects of delta PKC and epsilon PKC. Proc Natl Acad Sci USA 98:11114–11119. doi:10.1073/pnas.191369098
Article PubMed CAS Google Scholar - Das M, Das S, Lekli I, Das DK (2011) Caveolin induces cardioprotection through epigenetic regulation. J Cell Mol Med (in press). doi:10.1111/j.1582-4934.2011.01372.x
- Deaton AM, Bird A (2011) CpG islands and the regulation of transcription. Genes Dev 25(10):1010–1022. doi:10.1002/bies.201000057
Article PubMed CAS Google Scholar - Defossez PA, Stancheva I (2011) Biological functions of methyl-CpG-binding proteins. Prog Mol Biol Transl Sci 101:377–398. doi:10.1016/B978-0-12-387685-0.00012-3
Article PubMed CAS Google Scholar - The ENCODE Project Consortium (2004) The ENCODE (ENCyclopedia Of DNA Elements) Project. Science 306:636–640. doi:10.1126/science.1105136
Article Google Scholar - Dixit M, Ansseau E, Tassin A, Winokur S, Shi R, Qian H, Sauvage S, Matteotti C, van Acker AM, Leo O, Figlewicz D, Barro M, Laoudj-Chenivesse D, Belayew A, Coppee F, Chen YW (2007) DUX4, a candidate gene of facioscapulohumeral muscular dystrophy, encodes a transcriptional activator of PITX1. Proc Natl Acad Sci USA 104:18157–18162. doi:10.1073/pnas.0708659104
Article PubMed CAS Google Scholar - Ferguson LR (2011) RNA silencing: mechanism, biology and responses to environmental stress. Mutat Res 714(1–2):93–94. doi:10.1016/j.mrfmmm.2011.07.007
PubMed CAS Google Scholar - Gama-Sosa MA, Midgett RM, Slagel VA, Githens S, Kuo KC, Gehrke CW, Ehrlich M (1983) Tissue-specific differences in DNA methylation in various mammals. Biochim Biophys Acta 740:212–219. doi:10.1016/0167-4781(83)90079-9
PubMed CAS Google Scholar - Gardiner-Garden M, Frommer M (1987) CpG islands in vertebrate genomes. J Mol Biol 196:261–282. doi:10.1016/0022-2836(87)90689-9
Article PubMed CAS Google Scholar - Golding MC, Magri LS, Zhang L, Lalone SA, Higgins MJ, Mann MR (2011) Depletion of Kcnq1ot1 non-coding RNA does not affect imprinting maintenance in stem cells. Development 138(17):3667–3678. doi:10.1242/dev.057778
Article PubMed CAS Google Scholar - Gray MO, Zhou HZ, Schafhalter-Zoppoth I, Zhu P, Mochly-Rosen D, Messing RO (2004) Preservation of base-line hemodynamic function and loss of inducible cardioprotection in adult mice lacking protein kinase Cε. J Biol Chem 279:3596–3604. doi:10.1074/jbc.M311459200
Article PubMed CAS Google Scholar - Godfrey KM, Barker DJ (2001) Fetal programming and adult health. Pub Health Nutr 4:611–624. doi:10.1079/PHN2001145
Article CAS Google Scholar - Gregory RI, Yan KP, Amuthan G, Chendrimada T, Doratotaj B, Cooch N, Shiekhattar R (2004) The microprocessor complex mediates the genesis of microRNAs. Nature 432:235–240. doi:10.1038/nature03120
Article PubMed CAS Google Scholar - Guo H, Ingolia NT, Weissman JS, Bartel DP (2010) Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature 466:835–840. doi:10.1038/nature09267
Article PubMed CAS Google Scholar - Hartmann D, Thum T (2011) MicroRNAs and vascular (dys)function. Vascul Pharmacol (in press). doi:10.1016/j.vph.2011.07.005
- Heijmans BT, Tobi EW, Stein AD, Putter H, Blauw GJ, Susser ES, Slagboom PE, Lumey LH (2008) Persistent epigenetic differences associated with prenatal exposure to famine in humans. Proc Natl Acad Sci USA 105:17046–17049. doi:10.1073/pnas.0806560105
Article PubMed CAS Google Scholar - Helgadottir A, Thorleifsson G, Manolescu A, Gretarsdottir S, Blondal T, Jonasdottir A, Jonasdottir A, Sigurdsson A, Baker A, Palsson A, Masson G, Gudbjartsson DF, Magnusson KP, Andersen K, Levey AI, Backman VM, Matthiasdottir S, Jonsdottir T, Palsson S, Einarsdottir H, Gunnarsdottir S, Gylfason A, Vaccarino V, Hooper WC, Reilly MP, Granger CB, Austin H, Rader DJ, Shah SH, Quyyumi AA, Gulcher JR, Thorgeirsson G, Thorsteinsdottir U, Kong A, Stefansson K (2007) A common variant on chromosome 9p21 affects the risk of myocardial infarction. Science 316:1491–1493. doi:10.1126/science.1142842
Article PubMed CAS Google Scholar - Helgadottir A, Thorleifsson G, Magnusson KP, Grétarsdottir S, Steinthorsdottir V, Manolescu A, Jones GT, Rinkel GJ, Blankensteijn JD, Ronkainen A, Jääskeläinen JE, Kyo Y, Lenk GM, Sakalihasan N, Kostulas K, Gottsäter A, Flex A, Stefansson H, Hansen T, Andersen G, Weinsheimer S, Borch-Johnsen K, Jorgensen T, Shah SH, Quyyumi AA, Granger CB, Reilly MP, Austin H, Levey AI, Vaccarino V, Palsdottir E, Walters GB, Jonsdottir T, Snorradottir S, Magnusdottir D, Gudmundsson G, Ferrell RE, Sveinbjornsdottir S, Hernesniemi J, Niemelä M, Limet R, Andersen K, Sigurdsson G, Benediktsson R, Verhoeven EL, Teijink JA, Grobbee DE, Rader DJ, Collier DA, Pedersen O, Pola R, Hillert J, Lindblad B, Valdimarsson EM, Magnadottir HB, Wijmenga C, Tromp G, Baas AF, Ruigrok YM, van Rij AM, Kuivaniemi H, Powell JT, Matthiasson SE, Gulcher JR, Thorgeirsson G, Kong A, Thorsteinsdottir U, Stefansson K (2008) The same sequence variant on 9p21 associates with myocardial infarction, abdominal aortic aneurysm and intracranial aneurysm. Nat Genet 40:217–224. doi:10.1038/ng.72
Article PubMed CAS Google Scholar - Heusch G, Boengler K, Schulz R (2010) Cardioprotection: nitric oxide, protein kinases, and mitochondria. Circulation 118(19):1915–1919. doi:10.1161/CIRCULATIONAHA.108.805242
Article Google Scholar - Huarte M, Rinn JL (2010) Large non-coding RNAs: missing links in cancer? Hum Mol Genet 19(R2): R152–R161. doi:10.1093/hmg/ddq353
- Illi B, Nanni S, Scopece A, Farsetti A, Biglioli P, Capogrossi MC, Gaetano C (2003) Shear stress-mediated chromatin remodeling provides molecular basis for flow-dependent regulation of gene expression. Circ Res 25 93(2):155–161. doi:10.1161/01.RES.0000080933.82105.29
Google Scholar - Kaikkonen MU, Lam MT, Glass CK (2011) Non-coding RNAs as regulators of gene expression and epigenetics. Cardiovasc Res 90(3):430–440. doi:10.1093/cvr/cvr097
Article PubMed CAS Google Scholar - Kaneda R, Takada S, Yamashita Y, Choi YL, Nonaka-Sarukawa M, Soda M, Misawa Y, Isomura T, Shimada K, Mano H (2009) Genome-wide histone methylation profile for heart failure. Genes Cells 14(1):69–77. doi:10.1111/j.1365-2443.2008.01252.x
Article PubMed CAS Google Scholar - Kouzarides T (2007) Chromatin modifications and their function. Cell 128(4):693–705. doi:10.1016/j.cell.2007.02.005
Article PubMed CAS Google Scholar - Laird PW (2003) The power and the promise of DNA methylation markers. Nat Rev Cancer 3(4):253–266. doi:10.1038/nrc1045
Article PubMed CAS Google Scholar - Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene _lin_-4 encodes small RNAs with antisense complementarity to _lin_-14. Cell 75:843–854. doi:10.1016/0092-8674(93)90529-Y
Article PubMed CAS Google Scholar - Lee Y, Kim M, Han J, Yeom KH, Lee S, Baek SH, Kim VN (2004) MicroRNA genes are transcribed by RNA polymerase II. EMBO J 23:4051–4060. doi:10.1038/sj.emboj.7600385
Article PubMed CAS Google Scholar - Lee Y, Ahn C, Han J, Choi H, Kim J, Yim J, Lee J, Provost P, Rådmark O, Kim S, Kim VN (2003) The nuclear RNase III Drosha initiates microRNA processing. Nature 425:415–419. doi:10.1038/nature01957
Article PubMed CAS Google Scholar - Li E, Beard C, Jaenisch R (1993) Role for DNA methylation in genomic imprinting. Nature 366:362–365. doi:10.1038/366362a0
Article PubMed CAS Google Scholar - Li K, Blum Y, Verma A, Liu Z, Pramanik K, Leigh NR, Chun CZ, Samant GV, Zhao B, Garnaas MK, Horswill MA, Stanhope SA, North PE, Miao RQ, Wilkinson GA, Affolter M, Ramchandran R (2010) A noncoding antisense RNA in tie-1 locus regulates tie-1 function in vivo. Blood 115(1):133–139. doi:10.1182/blood-2009-09-242180
Article PubMed CAS Google Scholar - Litt MD, Simpson M, Gaszner M, Allis CD, Felsenfeld G (2001) Correlation between histone lysine methylation and developmental changes at the chicken betaglobin locus. Science 293(5539):2453–2455. doi:10.1126/science.1064413
Article PubMed CAS Google Scholar - Margariti A, Zampetaki A, Xiao Q, Zhou B, Karamariti E, Martin D, Yin X, Mayr M, Li H, Zhang Z, De Falco E, Hu Y, Cockerill G, Xu Q, Zeng L (2010) Histone deacetylase 7 controls endothelial cell growth through modulation of beta-catenin. Circ Res 106:1202–1211. doi:10.1161/CIRCRESAHA.109.213165
Article PubMed CAS Google Scholar - Mathiyalagan P, Chang L, Du XJ, El-Osta A (2010) Cardiac ventricular chambers are epigenetically distinguishable. Cell Cycle 9(3):612–617. http://dx.doi.org/10.4161/cc.9.3.10612
Google Scholar - McDonald OG, Owens GK (2007) Programming smooth muscle plasticity with chromatin dynamics. Circ Res 100:1428–1441. doi:10.1161/01.RES.0000266448.30370.a0
Article PubMed CAS Google Scholar - Meder B, Keller A, Vogel B, Haas J, Sedaghat-Hamedani F, Kayvanpour E, Just S, Borries A, Rudloff J, Leidinger P, Meese E, Katus HA, Rottbauer W (2011) MicroRNA signatures in total peripheral blood as novel biomarkers for acute myocardial infarction. Basic Res Cardiol 106(1):13–23. doi:10.1002/ijc.26419
Article PubMed CAS Google Scholar - MicroCosm Targets (2010) Version 5. EMBL–EBI database [online]. http://www.ebi.ac.uk/enright-srv/microcosm/htdocs/targets/v5/
- Mottet D, Bellahcene A, Pirotte S, Waltregny D, Deroanne C, Lamour V, Lidereau R, Castronovo V (2007) Histone deacetylase 7 silencing alters endothelial cell migration, a key step in angiogenesis. Circ Res 101:1237–1246. doi:10.1161/CIRCRESAHA.107.149377
Article PubMed CAS Google Scholar - Movassagh M, Choy MK, Goddard M, Bennett MR, Down TA, Foo RS (2010) Differential DNA methylation correlates with differential expression of angiogenic factors in human heart failure. PLoS One 5(1):e8564. doi:10.1371/journal.pone.0008564
Article PubMed Google Scholar - Movassagh M, Choy MK, Knowles DA, Cordeddu L, Haider S, Down T, Siggens L, Vujic A, Simeoni I, Penkett C, Goddard M, Lio P, Bennett MR, Foo RS (2011) Distinct epigenomic features in end-stage failing human hearts. Circulation. doi:10.1161/CIRCULATIONAHA.111.040071
- Mund C, Brueckner B, Lyko F (2006) Reactivation of epigenetically silenced genes by DNA methyltransferase inhibitors: basic concepts and clinical applications. Epigenetics 1(1):7–13. doi:10.4161/epi.1.1.2375
Article PubMed Google Scholar - Murriel CL, Mochly-Rosen D (2003) Opposing roles of delta and epsilon PKC in cardiac ischemia and reperfusion: targeting the apoptotic machinery. Arch Biochem Biophys 420:246–254. doi:10.1016/j.abb.2003.08.038
Article PubMed CAS Google Scholar - Nagano T, Mitchell JA, Sanz LA, Pauler FM, Ferguson-Smith AC, Feil R, Fraser P (2008) The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science 322:1717–1720. doi:10.1126/science.1163802
Article PubMed CAS Google Scholar - Ohtani K, Dimmeler S (2011) Control of cardiovascular differentiation by microRNAs. Basic Res Cardiol 106(1):5–11. doi:10.1007/s00395-010-0139-7
Article PubMed CAS Google Scholar - Ohtani K, Vlachojannis GJ, Koyanagi M, Boeckel JN, Urbich C, Farcas R, Bonig H, Marquez VE, Zeiher AM, Dimmeler S (2011) Epigenetic regulation of endothelial lineage committed genes in pro-angiogenic hematopoietic and endothelial progenitor cells. Circ Res doi. doi:10.1161/CIRCRESAHA.111.247304
Google Scholar - Painter RC, Roseboom TJ, Bleker OP (2005) Prenatal exposure to the Dutch famine and disease in later life: an overview. Reprod Toxicol 20:345–352. doi:10.1016/j.reprotox.2005.04.005
Article PubMed CAS Google Scholar - Panning B, Jaenisch R (1996) DNA hypomethylation can activate Xist expression and silence X-linked genes. Genes Dev 10:1991–2002. doi:10.1101/gad.10.16.1991
Article PubMed CAS Google Scholar - Pasmant E, Laurendeau I, Héron D, Vidaud M, Vidaud D, Bièche I (2007) Characterization of a germ-line deletion, including the entire INK4/ARF locus, in a melanoma-neural system tumor family: identification of ANRIL, an antisense noncoding RNA whose expression coclusters with ARF. Cancer Res 67:3963–3969. doi:10.1158/0008-5472.CAN-06-2004
Article PubMed CAS Google Scholar - Patterson AJ, Chen M, Xue Q, Xiao D, Zhang L (2010) Chronic prenatal hypoxia induces epigenetic programming of PKC{epsilon} gene repression in rat hearts. Circ Res 107(3):365–373. doi:10.1161/CIRCRESAHA.110.221259
Article PubMed CAS Google Scholar - Pearson CE (2010) FSHD: a repeat contraction disease finally ready to expand (our understanding of its pathogenesis). PLoS Genet 6:e1001180. doi:10.1371/journal.pgen.1001180
Article PubMed Google Scholar - Ponting CP, Oliver PL, Reik W (2009) Evolution and functions of long noncoding RNAs. Cell 136:629–641. doi:10.1016/j.cell.2009.02.006
Article PubMed CAS Google Scholar - Robb GB, Carson AR, Tai SC, Fish JE, Singh S, Yamada T, Scherer SW, Nakabayashi K, Marsden PA (2004) Post-transcriptional regulation of endothelial nitric-oxide synthase by an overlapping antisense mRNA transcript. J Biol Chem 279(36):37982–37996. doi:10.1074/jbc.M400271200
Article PubMed CAS Google Scholar - Saito Y, Liang G, Egger G, Friedman JM, Chuang JC, Coetzee GA, Jones PA (2006) Specific activation of microRNA-127 with downregulation of the proto-oncogene BCL6 by chromatin-modifying drugs in human cancer cells. Cancer Cell 9(6):435–443. doi:10.1016/j.ccr.2006.04.020
Article PubMed CAS Google Scholar - Santos-Rosa H, Schneider R, Bannister AJ, Sherriff J, Bernstein BE, Emre NC, Schreiber SL, Mellor J, Kouzarides T (2002) Active genes are tri-methylated at K4 of histone H3. Nature 419(6905):407–411. doi:10.1038/nature01080
Article PubMed CAS Google Scholar - Saxonov S, Berg P, Brutlag DL (2006) A genome-wide analysis of CpG dinucleotides in the human genome distinguishes two distinct classes of promoters. Proc Natl Acad Sci USA 103(5):1412–1417. doi:10.1073/pnas.0510310103
Article PubMed CAS Google Scholar - Singh N, Trivedi CM, Lu M, Mullican SE, Lazar MA, Epstein JA (2011) Histone deacetylase 3 regulates smooth muscle differentiation in neural crest cells and development of the cardiac outflow Tract. Circ Res. doi:10.1038/nrm3198
- Spin JM, Quertermous T, Tsao PS (2010) Chromatin remodeling pathways in smooth muscle cell differentiation, and evidence for an integral role for p300. PLoS One 5(12):e14301. doi:10.1371/journal.pone.0014301
Article PubMed CAS Google Scholar - Stein AB, Jones TA, Herron TJ, Patel SR, Day SM, Noujaim SF, Milstein ML, Klos M, Furspan PB, Jalife J, Dressler GR (2011) Loss of H3K4 methylation destabilizes gene expression patterns and physiological functions in adult murine cardiomyocytes. J Clin Invest 121(7):2641–2650. doi:10.1172/JCI44641
Article PubMed CAS Google Scholar - Takai D, Jones PA (2002) Comprehensive analysis of CpG islands in human chromosomes 21 and 22. Proc Natl Acad Sci USA 99:3740–3745. doi:10.1073/pnas.052410099
Article PubMed CAS Google Scholar - Thrash-Bingham CA, Tartof KD (1999) aHIF: a natural antisense transcript overexpressed in human renal cancer and during hypoxia. J Natl Cancer Inst 91:143–151. doi:10.1093/jnci/91.2.143
Article PubMed CAS Google Scholar - Thum T, Gross C, Fiedler J, Fischer T, Kissler S, Bussen M, Galuppo P, Just S, Rottbauer W, Frantz S, Castoldi M, Soutschek J, Koteliansky V, Rosenwald A, Basson MA, Licht JD, Pena JT, Rouhanifard SH, Muckenthaler MU, Tuschl T, Martin GR, Bauersachs J, Engelhardt S (2008) MicroRNA-21 contributes to myocardial disease by stimulating MAP kinase signalling in fibroblasts. Nature 456:980–984. doi:10.1038/nature07511
Article PubMed CAS Google Scholar - Thum T, Galuppo P, Wolf C, Fiedler J, Kneitz S, van Laake LW, Doevendans PA, Mummery CL, Borlak J, Haverich A, Gross C, Engelhardt S, Ertl G, Bauersachs J (2007) MicroRNAs in the human heart: a clue to fetal gene reprogramming in heart failure. Circulation 116:258–267. doi:10.1161/CIRCULATIONAHA.107.687947
Article PubMed CAS Google Scholar - Tsutsumi YM, Horikawa YT, Jennings MM, Kidd MW, Niesman IR, Yokoyama U, Head BP, Hagiwara Y, Ishikawa Y, Miyanohara A, Patel PM, Insel PA, Patel HH, Roth DM (2008) Cardiac-specific overexpression of caveolin-3 induces endogenous cardiac protection by mimicking ischemic preconditioning. Circulation 118(19):1979–1988. doi:10.1161/CIRCULATIONAHA.108.788331
Article PubMed CAS Google Scholar - Urbich C, Kuehbacher A, Dimmeler S (2008) Role of microRNAs in vascular diseases, inflammation, and angiogenesis. Cardiovasc Res 79(4):581–588. doi:10.1093/cvr/cvn156
Article PubMed CAS Google Scholar - Valencia-Sanchez MA, Liu J, Hannon GJ, Parker R (2006) Control of translation and mRNA degradation by miRNAs and siRNAs. Genes Dev 20:515–524. doi:10.1101/gad.1399806
Article PubMed CAS Google Scholar - van Rooij E, Sutherland LB, Thatcher JE, DiMaio JM, Naseem RH, Marshall WS, Hill JA, Olson EN (2008) Dysregulation of microRNAs after myocardial infarction reveals a role of miR-29 in cardiac fibrosis. Proc Natl Acad Sci USA 105:13027–13032. doi:10.1073/pnas.0805038105
Article PubMed Google Scholar - Walsh CP, Chaillet JR, Bestor TH (1998) Transcription of IAP endogenous retroviruses is constrained by cytosine methylation. Nat Genet 20:116–117. doi:10.1038/2413
Article PubMed CAS Google Scholar - Weber M, Hellmann I, Stadler MB, Ramos L, Paabo S, Rebhan M, Schubeler D (2007) Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome. Nat Genet 39:457–466. doi:10.1038/ng1990
Article PubMed CAS Google Scholar - Yang F, Zhang L, Huo XS, Yuan JH, Xu D, Yuan SX, Zhu N, Zhou WP, Yang GS, Wang YZ, Shang JL, Gao CF, Zhang FR, Wang F, Sun SH (2011) Long non-coding RNA high expressed in hepatocellular carcinoma (lncRNA-HEIH) facilitates tumor growth through enhancer of zeste homolog 2. Hepatology (in press). doi:10.1002/hep.24563
- Yap KL, Li S, Muñoz-Cabello AM, Raguz S, Zeng L, Mujtaba S, Gil J, Walsh MJ, Zhou MM (2010) Molecular interplay of the noncoding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol Cell 38(5):662–674. doi:10.1016/j.molcel.2010.03.021
Article PubMed CAS Google Scholar - Yu W, Gius D, Onyango P, Muldoon-Jacobs K, Karp J, Feinberg AP, Cui H (2008) Epigenetic silencing of tumour suppressor gene p15 by its antisense RNA. Nature 451:202–206. doi:10.1038/nature06468
Article PubMed CAS Google Scholar - Zhao J, Sun BK, Erwin JA, Song JJ, Lee JT (2008) Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 322(5902):750–756. doi:10.1126/science.1163045
Article PubMed CAS Google Scholar - Zhang H, Darwanto A, Linkhart TA, Sowers LC, Zhang L (2007) Maternal cocaine administration causes an epigenetic modification of protein kinase Cepsilon gene expression in fetal rat heart. Mol Pharmacol 71(5):1319–1328. doi:10.1124/mol.106.032011
Article PubMed CAS Google Scholar - Zhang QJ, Chen HZ, Wang L, Liu DP, Hill JA, Liu ZP (2011) The histone trimethyllysine demethylase JMJD2A promotes cardiac hypertrophy in response to hypertrophic stimuli in mice. J Clin Invest 121(6):2447–2456. doi:10.1172/JCI46277
Article PubMed CAS Google Scholar