Feeder-free culture of human embryonic stem cells in conditioned medium for efficient genetic modification (original) (raw)
- Protocol
- Published: 14 August 2008
- Chris Denning3,
- Elena Matsa3,
- Lorraine E Young3,
- Robert Passier1,2 &
- …
- Christine L Mummery1,2
Nature Protocols volume 3, pages 1435–1443 (2008) Cite this article
- 2684 Accesses
- 72 Citations
- Metrics details
Abstract
Realizing the potential of human embryonic stem cells (hESCs) in research and commercial applications requires generic protocols for culture, expansion and genetic modification that function between multiple lines. Here we describe a feeder-free hESC culture protocol that was tested in 13 independent hESC lines derived in five different laboratories. The procedure is based on Matrigel adaptation in mouse embryonic fiboblast conditioned medium (CM) followed by monolayer culture of hESC. When combined, these techniques provide a robust hESC culture platform, suitable for high-efficiency genetic modification via plasmid transfection (using lipofection or electroporation), siRNA knockdown and viral transduction. In contrast to other available protocols, it does not require optimization for individual lines. hESC transiently expressing ectopic genes are obtained within 9 d and stable transgenic lines within 3 weeks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Photographs of human embryonic stem cell (hESC) at different stages.

The alternative text for this image may have been generated using AI.
Figure 2: Flowchart for experimental procedures.
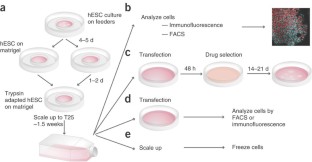
The alternative text for this image may have been generated using AI.
Similar content being viewed by others
References
- Osafune, K. et al. Marked differences in differentiation propensity among human embryonic stem cell lines. Nat. Biotechnol. 26, 313–315 (2008).
Article CAS PubMed Google Scholar - Giudice, A. & Trounson, A. Genetic modification of human embryonic stem cells for derivation of target cells. Cell Stem Cell 2, 422–433 (2008).
Article CAS PubMed Google Scholar - Davis, R.P. et al. Targeting a GFP reporter gene to the MIXL1 locus of human embryonic stem cells identifies human primitive streak-like cells and enables isolation of primitive hematopoietic precursors. Blood 111, 1876–1884 (2008).
Article CAS PubMed Google Scholar - Urbach, A., Schuldiner, M. & Benvenisty, N. Modeling for Lesch-Nyhan disease by gene targeting in human embryonic stem cells. Stem Cells 22, 635–641 (2004).
Article CAS PubMed Google Scholar - Costa, M. et al. A method for genetic modification of human embryonic stem cells using electroporation. Nat. Protoc. 2, 792–796 (2007).
Article CAS PubMed Google Scholar - Zwaka, T.P. & Thomson, J.A. Homologous recombination in human embryonic stem cells. Nat. Biotechnol. 21, 319–321 (2003).
Article CAS PubMed Google Scholar - Draper, J.S. et al. Recurrent gain of chromosomes 17q and 12 in cultured human embryonic stem cells. Nat. Biotechnol. 22, 53–54 (2004).
Article CAS PubMed Google Scholar - Braam, S.R. et al. Improved genetic manipulation of human embryonic stem cells. Nat. Meth. 5, 389–392 (2008).
Article CAS Google Scholar - Liew, C.-G., Draper, J.S., Walsh, J., Moore, H. & Andrews, P.W. Transient and stable transgene expression in human embryonic stem cells. Stem Cells 25, 1521–1528 (2007).
Article CAS PubMed Google Scholar - Dormeyer, W. et al. Plasma membrane proteomics of human embryonic stem cells and human embryonal carcinoma cells. J. Proteome Res. 7, 2936–2951 (2008).
Article CAS PubMed Google Scholar - Ludwig, T.E. et al. Derivation of human embryonic stem cells in defined conditions. Nat. Biotechnol. 24, 185–187 (2006).
Article CAS PubMed Google Scholar - Ng, E.S., Davis, R., Stanley, E.G. & Elefanty, A.G. A protocol describing the use of a recombinant protein-based, animal product-free medium (APEL) for human embryonic stem cell differentiation as spin embryoid bodies. Nat. Protoc. 3, 768–776 (2008).
Article CAS PubMed Google Scholar - Wu, S., Ying, G., Wu, Q. & Capecchi, M.R. A protocol for constructing gene targeting vectors: generating knockout mice for the cadherin family and beyond. Nat. Protoc. 3, 1056–1076 (2008).
Article CAS PubMed Google Scholar - Kameda, T., Smuga-Otto, K. & Thomson, J.A. A severe de novo methylation of episomal vectors by human ES cells. Biochem. Biophys. Res. Commun. 349, 1269–1277 (2006).
Article CAS PubMed Google Scholar - Anderson, D. et al. Transgenic enrichment of cardiomyocytes from human embryonic stem cells. Mol. Ther. 15, 2027–2036 (2007).
Article CAS PubMed Google Scholar - Cowan, C.A. et al. Derivation of embryonic stem-cell lines from human blastocysts. N. Engl. J. Med. 350, 1353–1356 (2004).
Article CAS PubMed Google Scholar - Burridge, P.W. et al. Improved human embryonic stem cell embryoid body homogeneity and cardiomyocyte differentiation from a novel V-96 plate aggregation system highlights interline variability. Stem Cells 25, 929–938 (2007).
Article CAS PubMed Google Scholar - Mitalipova, M. et al. Human embryonic stem cell lines derived from discarded embryos. Stem Cells 21, 521–526 (2003).
Article CAS PubMed Google Scholar - Reubinoff, B.E., Pera, M.F., Fong, C.Y., Trounson, A. & Bongso, A. Embryonic stem cell lines from human blastocysts: somatic differentiation in vitro. Nat. Biotechnol. 18, 399–404 (2000).
Article CAS PubMed Google Scholar - Costa, M. et al. The hESC line Envy expresses high levels of GFP in all differentiated progeny. Nat. Meth. 2, 259–260 (2005).
Article CAS Google Scholar - Michaoska, A.E. Unit 1C.3 Isolation and propagation of mouse embryonic fibroblasts and preparation of mouse embryonic feeder layer cells. Curr. Protoc. Stem Cell Biol. 3, 1C.3.1–1C.3.17 (2007).
Google Scholar - Baker, D.E.C. et al. Adaptation to culture of human embryonic stem cells and oncogenesis in vivo. Nat. Biotechnol. 25, 207–215 (2007).
Article CAS PubMed Google Scholar - Shapiro, H.M. Practical Flow Cytometry (Wiley Interscience, New York, 2003).
Book Google Scholar - Brown, C.M. Fluorescence microscopy—avoiding the pitfalls. J. Cell Sci. 120, 1703–1705 (2007).
Article CAS PubMed Google Scholar
Acknowledgements
We are grateful to D. Ward-van Oostwaard, L. Zeinstra and S. van den Brink for expert technical assistance. We thank Drs Chad Cowan and Douglas Melton for the gift of HUES-1, -5, -7 and -15. This work is/has been supported by the Dutch Program for Tissue Engineering (S.R.B.), European Community's Sixth Framework Programme contract ('HeartRepair') LSHM-CT-2005-018630 (R.P.), the Biotechnology and Biological Sciences Research Council, British Heart Foundation and the University of Nottingham (C.D., E.M. and L.E.Y.).
Author information
Authors and Affiliations
- Hubrecht Institute, Developmental Biology and Stem Cell Research, Uppsalalaan 8, Utrecht, 3584 CT, The Netherlands
Stefan R Braam, Robert Passier & Christine L Mummery - Department of Anatomy and Embryology, Leiden University Medical Centre, Leiden, Postal Zone S-1-P, P.O. Box 9600, Leiden, 2300 RC, The Netherlands
Stefan R Braam, Robert Passier & Christine L Mummery - Wolfson Centre for Stem Cells, Tissue Engineering and Modelling (STEM), Centre for Biomolecular Sciences, University of Nottingham, Nottingham, NG7 2RD, United Kingdom
Chris Denning, Elena Matsa & Lorraine E Young
Authors
- Stefan R Braam
- Chris Denning
- Elena Matsa
- Lorraine E Young
- Robert Passier
- Christine L Mummery
Contributions
S.R.B. and C.D. contributed equally to this work.
Corresponding authors
Correspondence toChris Denning or Christine L Mummery.
Rights and permissions
About this article
Cite this article
Braam, S., Denning, C., Matsa, E. et al. Feeder-free culture of human embryonic stem cells in conditioned medium for efficient genetic modification.Nat Protoc 3, 1435–1443 (2008). https://doi.org/10.1038/nprot.2008.140
- Published: 14 August 2008
- Issue date: September 2008
- DOI: https://doi.org/10.1038/nprot.2008.140