Assessing drug target suitability using TargetMine (original) (raw)
Software Tool Article
[version 1; peer review: 2 approved]
https://orcid.org/0000-0003-4936-8060
1, Erika Yogo2, Naoko Kurihara2, [...] Tomoshige Ohno2, Chihiro Higuchi1, Masatomo Rokushima2, Kenji Mizuguchi
https://orcid.org/0000-0003-3021-7078
1
https://orcid.org/0000-0003-4936-8060
1, Erika Yogo2, [...] Naoko Kurihara2, Tomoshige Ohno2, Chihiro Higuchi1, Masatomo Rokushima2, Kenji Mizuguchi
https://orcid.org/0000-0003-3021-7078
1
1 National Institutes of Biomedical Innovation, Health and Nutrition, Ibaraki, Osaka, 5670085, Japan
2 Shionogi Pharmaceutical Research Center, Shionogi & Co., Ltd., Toyonaka, Osaka, 5610825, Japan
Yi-An Chen
Roles: Conceptualization, Data Curation, Investigation, Methodology, Software, Visualization, Writing – Original Draft Preparation
Erika Yogo
Roles: Conceptualization, Investigation, Project Administration, Visualization, Writing – Original Draft Preparation, Writing – Review & Editing
Naoko Kurihara
Roles: Data Curation, Investigation, Methodology
Tomoshige Ohno
Roles: Data Curation, Investigation
Chihiro Higuchi
Roles: Data Curation
Masatomo Rokushima
Roles: Conceptualization, Investigation, Supervision, Visualization, Writing – Original Draft Preparation, Writing – Review & Editing
Kenji Mizuguchi
Roles: Conceptualization, Funding Acquisition, Investigation, Project Administration, Resources, Supervision, Writing – Review & Editing
OPEN PEER REVIEW
REVIEWER STATUS
Abstract
In selecting drug target candidates for pharmaceutical research, the linkage to disease and the tractability of the target are two important factors that can ultimately determine the drug efficacy. Several existing resources can provide gene-disease associations, but determining whether such a list of genes are attractive drug targets often requires further information gathering and analysis. In addition, few resources provide the information required to evaluate the tractability of a target. To address these issues, we have updated TargetMine, a data warehouse for assisting target prioritization, by integrating new data sources for gene-disease associations and enhancing functionalities for target assessment. As a data mining platform that integrates a variety of data sources, including protein structures and chemical compounds, TargetMine now offers a powerful and flexible interface for constructing queries to check genetic evidence, tractability and other relevant features for the candidate genes. We demonstrate these features by using several specific examples.
Keywords
disease, drug assessment, genetic variation, tractability
Corresponding authors: Yi-An Chen, Kenji Mizuguchi Competing interests: No competing interests were disclosed.
Grant information: This work was in part supported by JSPS KAKENHI (grant number 17K07268).
The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Copyright: © 2019 Chen YA et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. How to cite: Chen YA, Yogo E, Kurihara N et al. Assessing drug target suitability using TargetMine [version 1; peer review: 2 approved]. F1000Research 2019, 8:233 (https://doi.org/10.12688/f1000research.18214.1) First published: 28 Feb 2019, 8:233 (https://doi.org/10.12688/f1000research.18214.1) Latest published: 28 May 2019, 8:233 (https://doi.org/10.12688/f1000research.18214.2)
Introduction
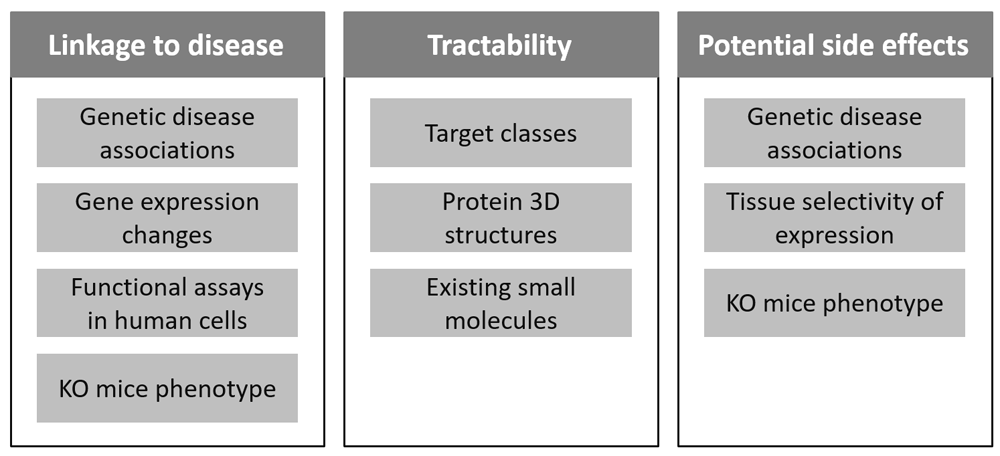
A drug discovery project typically begins with the identification of a target molecule. In evaluating potential drug targets, several factors must be taken into account: linkage to disease, tractability (the possibility of finding small molecule compounds with high affinity), potential side effects, novelty, as well as the competitiveness in the market (Figure 1). Among these factors, the linkage to disease and the tractability are particularly important in terms of the drug efficacy, and become key factors in whether or not the pharmaceutical research and development (R&D) is successful when selecting drug targets1,2. The most important part of the linkage to disease is genetic associations for the disease or relevant traits. According to analyses reported by AstraZeneca and GlaxoSmithKline, the success rate of such R&D is increased when the choice of the selected target is supported by genetic evidence. The report from AstraZeneca shows that 73% of projects with some genetic linkage of the target to the disease indication in Phase II were active or successful compared to 43% of projects without such data3, while the analysis results from GlaxoSmithKline suggest that selecting genetically supported targets could double the success rate in clinical development4. Several existing resources provide information about genetic evidences, such as DisGeNET5, Open Targets6, and Pharos7. However, a simple list of genes with genetic linkage to the disease is often insufficient for evaluating the disease rationale fully, and additional information and analysis such as pathway enrichment analysis will be needed to assess other aspects of target suitability (e.g. drug mechanisms and safety). In addition, few resources provide tractability information, with the recent update of Open Targets being an exception.
Figure 1. Key factors to be considered in drug target selection.
Linkage to disease, tractability and adverse event risk are among the major factors to assess the suitability of novel target candidates. Much of the evidences regarding these factors are available in public domain resources.
To address these issues, we have updated TargetMine8, a data warehouse for assisting target prioritization, and improved its functionalities for target assessment, particularly in small molecule drug discovery. TargetMine8 utilizes the InterMine framework9 and facilitates flexible query construction spanning a wide range of integrated data sources including those relevant for evaluating linkage to disease and tractability. More specifically, we have integrated new data sources for genetic disease associations including ClinVar, dbSNP, and 1000 Genome Project, incorporated more details of the genome wide association studies from the GWAS catalog, and improved the data model overall to enable more efficient data mining. The new version provides a user-friendly and yet powerful interface to explore the disease rationale for human genes and helps prioritize the candidate genes in terms of both the genetic evidence and target tractability.
Methods
Implementation
TargetMine8 is based on the InterMine framework, an open-source data warehouse system designed for biological data integration9. In this update, we added a few customized data sources by defining new data models and implementing new data parsers. Details of how we designed the data models are described in the following sub-sections.
GWAS catalog
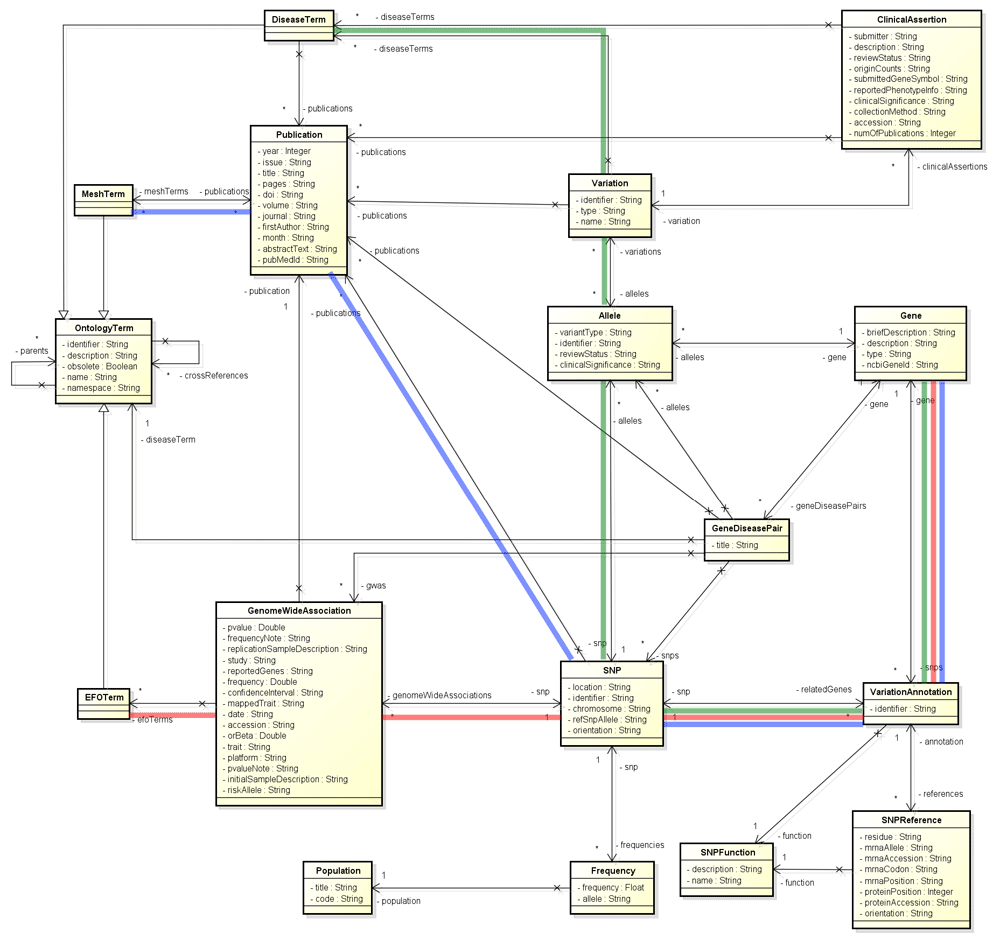
The GWAS catalog, founded by NHGRI, is a curated archive of the published genome wide association studies10. We had tried to associate genes to related diseases using the GWAS catalog in the former release of TargetMine11. To annotate disease terms to a trait or study, we first chose the disease ontology (DO)12,13 and then manually assigned the terms with the assistance of some text matching approaches. However, this process required some knowledge and involved a lot of manual examinations. Thus, it became difficult to keep updating regularly. Fortunately, the curation team started to use experiment factor ontology (EFO)14 to describe the curated GWAS traits in the recent implementation15. EFO covers several domain-specific ontologies that facilitate easier data integration. In our new implemented model, we replace DO terms with EFO terms and also incorporate some more information from each study (Figure 2). SNP annotations and details of EFO terms are retrieved from the dbSNP database and EFO, respectively.
Figure 2. The new implemented data model.
The colored lines indicate how the genes and diseases/phenotypes are associated in the post processing step.
ClinVar
ClinVar is a public archive of the relation between human variations and phenotypes16,17. As defined by ClinVar, a “Variation” could be a single variant, a compound heterozygote, or a complex haplotype. If a haplotype consists of multiple alleles, each allele is assigned with an independent identifier. On the other hand, the same allele could be the member of a different haplotype, thus the relation between the “Variation” and “Allele” is a many-to-many association. An “Allele” is supposed to describe a specific change of a variation, e.g. G>A. However, the SNP entries in dbSNP sometimes merge different combinations of variations (alleles) together if the variations occur at the same genomic position. Thus, an “SNP” entity may contain multiple “Allele” entries in the data model (Figure 2). Here, we only retrieve the SNP identifier, and the rest of the annotations are integrated from the dbSNP database. The structural variations which reference the dbVar records are not included in the current version. In addition, those alleles which were not assigned with any dbSNP or dbVar identifiers were treated as SNP entities and were stored in TargetMine8 using the information provided by ClinVar. Most of the data were processed from tab delimited files, while some information that were not available in the tab delimited files were processed from XML files. MedGen terms, which are used to integrate the human medical genetic information at NCBI (https://www.ncbi.nlm.nih.gov/medgen/), were adopted to describe diseases and phenotypes.
dbSNP
dbSNP is a database which archives short human genetic variations. We first performed a whole data dump to a relational database, and then made queries to extract the necessary information into a flat table. These data include genomic position (based on genome assembly GRCh38), reference mRNA, nucleotide variation, reference protein, and amino acid variation, if available. SNP to gene is a many-to-many relationship, thus we introduce an intermediate class named “VariationAnnotation” to associate them together (Figure 2). Although the InterMine framework is capable of incorporating whole SNP entries in dbSNP, the integration takes a few days to finish. Considering the frequency that we update TargetMine8 (once a month), it is not very practical to spend a few days doing the integration. As a tradeoff, we decided to store only a subset of SNPs. Only those SNPs which are related with GWAS associations or clinical assertions, or those where there is an associated publication, are selected for storage in TargetMine8.
Post-processing the integrated data
Our implementation allows us to associate the genetic phenotype (disease) and the gene via the GWAS or ClinVar dataset, or moreover the relation that is implied from the disease related MeSH (Medical Subject Headings, https://www.ncbi.nlm.nih.gov/mesh) terms assigned to the correlated publications of the SNPs. In order to make a shortcut and to summarize the available information, we perform post-processing and store the results using a new class named “GeneDiseasePair”. At the moment, there are three types of shortcuts. Gene to SNP to GWAS to EFO terms for GWAS catalog data (the red lines in Figure 2). Gene to SNP to clinical assertions to disease (MedGen) terms (the green lines in Figure 2). And Gene to SNP to publication to MeSH terms (the blue lines in Figure 2). The “GeneDiseasePair” class also includes correlated information including ontology terms, studies, SNPs and publications. These improvements in the data model facilitate quick access from a gene to the associated diseases, annotated by different data sources.
Operation
TargetMine8 is a Java-based web application that runs on Apache Tomcat. The user interface communicates with the integrated data stored in PostgreSQL, a relational database.
Use cases
Querying linkage to disease with TargetMine
To demonstrate the effectiveness of the new version of TargetMine8 in evaluating linkage to disease, we conducted a feasibility study, taking human PCSK9, proprotein convertase subtilisin/kexin type 9, as a typical case. The PCSK9 gene encodes a protein that promotes degradation of low-density lipoprotein (LDL) receptors in hepatocytes, thereby elevating or maintaining LDL cholesterol levels in the blood. Mutations in this gene are shown to be associated with familial hypercholesterolemia23, and monoclonal antibodies to PCSK9 have been launched on the market as drugs for hypercholesterolemia with and without genetic predispositions24,25.
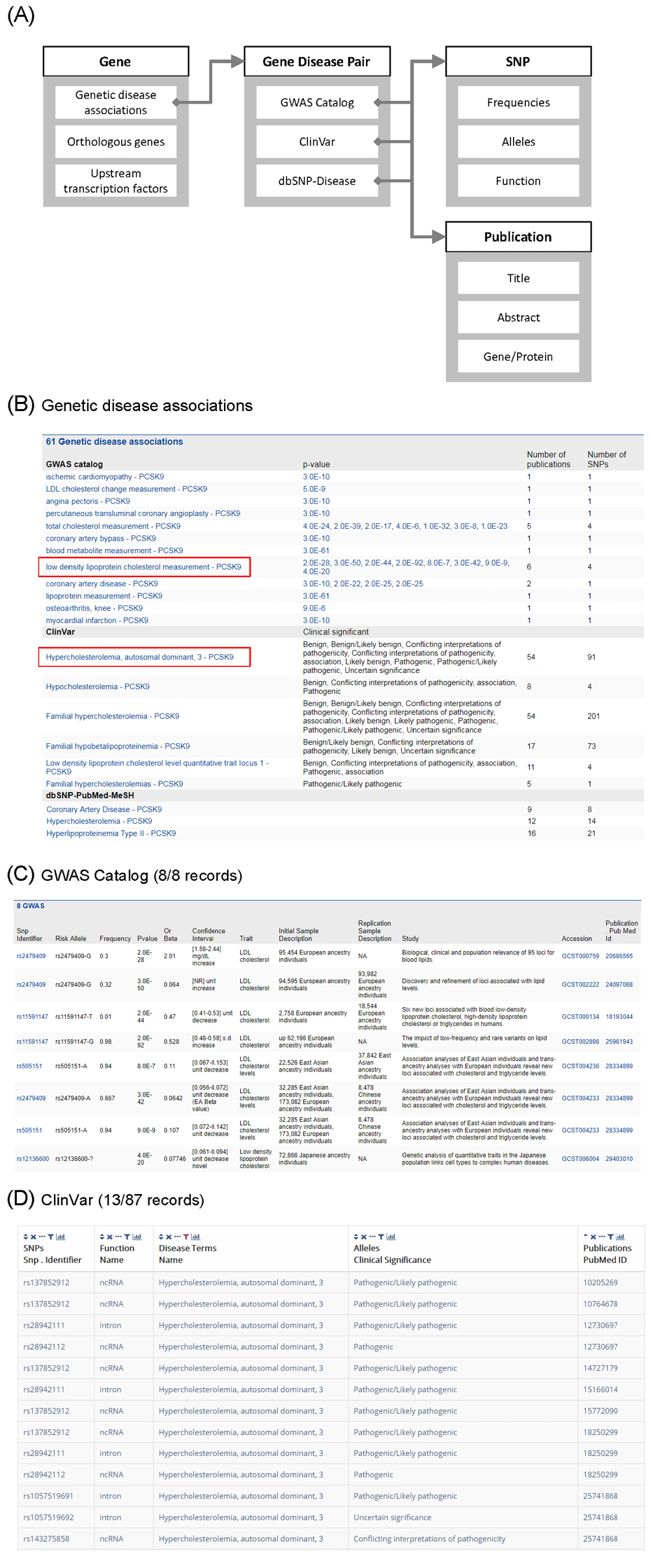
Figure 3A demonstrates a schematic representation of the searching protocol for genetic disease associations with TargetMine8. We first went to a gene report page by searching for the PCSK9 gene from the top page of TargetMine8 (not shown). From the gene report page, we got information of genetic disease associations (Figure 3B) as well as many other basic or advanced characteristics such as orthologous genes and upstream transcription factors. The results table of genetic disease associations for PCSK9 enabled us to confirm that a number of SNPs relevant to this gene have been reported to be associated with plasma LDL cholesterol levels, hypercholesterolemia, or coronary artery disease. By clicking the record of association between “low density lipoprotein cholesterol measurement” and PCSK9 in the GWAS catalog section (Figure 3B), we moved to a “gene disease pair” page and checked the details of the GWAS record, including the information on samples, statistical significance and publications (Figure 3C). Clicking on the SNP identifier (e.g., rs2479409) redirected us to an SNP report page containing the individual SNP basic information (allele, function, literature) and allele frequencies of different human populations (from 1000 Genome Project26 and others, not shown in the figure). Similarly, we examined the associations between “Hypercholesterolemia, autosomal dominant, 3” and PCSK9 from the ClinVar section in the table (Figure 3B) and got the details of the ClinVar record such as clinical assertions and publications (Figure 3D). The publications here reported mutations in PCSK9 as a cause of autosomal dominant hypercholesterolemia23 (not shown), as mentioned above.
Figure 3. Searching information about linkage to disease with TargetMine.
(A) Outline of the procedure for searching. (B) A screenshot of the summary of Genetic disease associations of PCSK9. (C) GWAS records of a pair of PCSK9 and low density lipoprotein cholesterol measurement. (D) ClinVar records of a pair of PCSK9 and hypercholesterolemia, autosomal dominant, 3.
Querying target tractability for small molecule drugs with TargetMine
We performed another feasibility study to examine whether TargetMine8 provides informative evidence to assess target tractability for small molecules. In this case we also used PCSK9 as an example because no potent small molecule inhibitors for this protein have been reported so far in spite of the intensive research activities of many laboratories27, indicating that PCSK9 is not a highly tractable target.
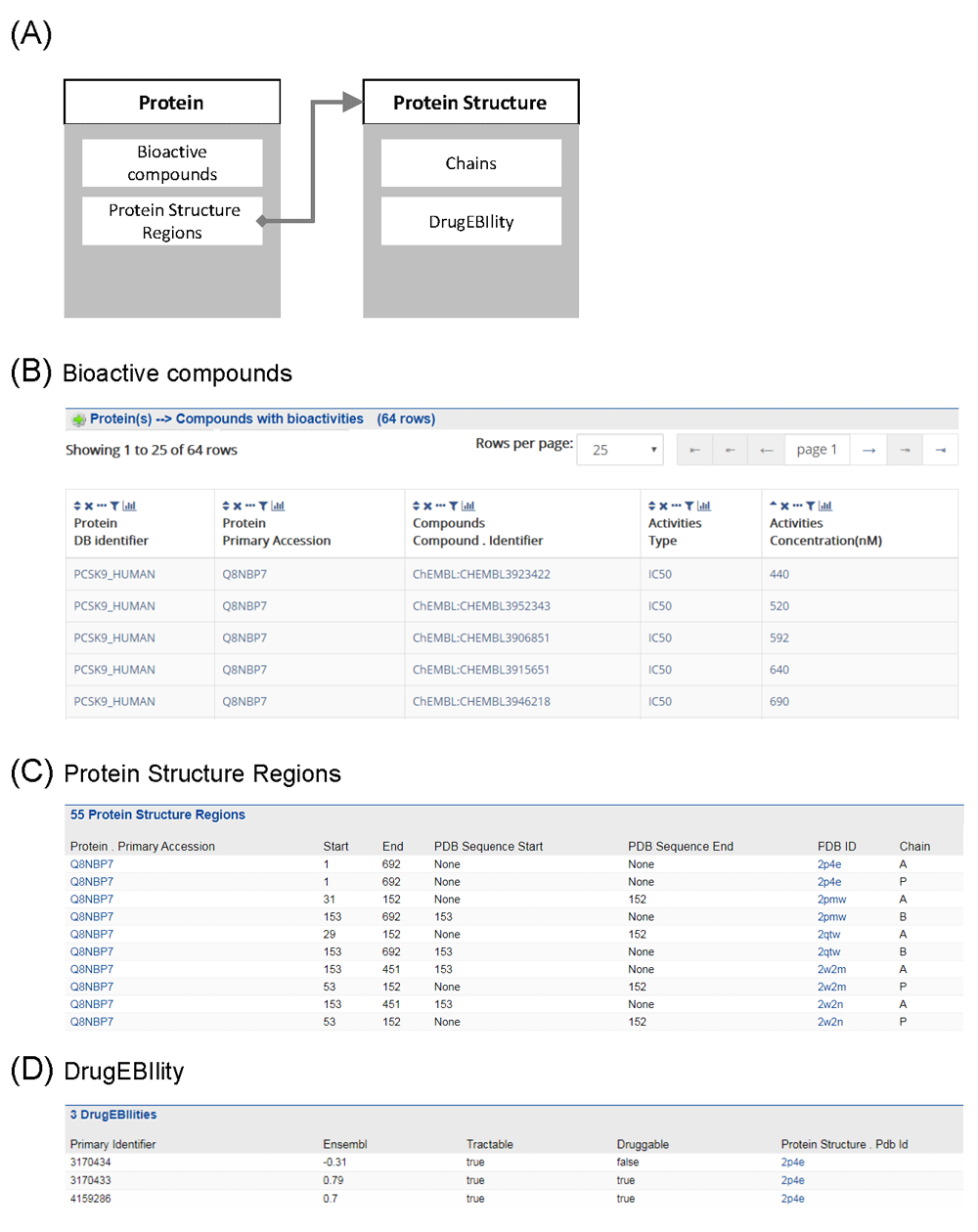
Figure 4A shows a schematic diagram of the procedure of querying tractability with TargetMine8. We first went to the protein report page of PCSK9 and found the bioactive compounds targeting this protein. As we expected, it was revealed that no potent compounds could be found in the ChEMBL database, and the lowest IC50 value was 440 nM (CHEMBL3923422) (Figure 4B). On the PCSK9 protein report page, we also checked the experimentally determined 3D structures, referred to as “protein structure regions” in TargetMine8, and identified several Protein Data Bank (PDB) entries for this protein (Figure 4C). Then, we moved to the “Protein Structure” page of a specified PDB ID (2p4e in this case) and found that in the “DrugEBIlity” table (from the DrugEBIlity database), some domains of the PCSK9 protein had positive Ensembl scores (Figure 4D), which are not ligand-based, but structure-based tractability scores. This result indicates that PCSK9 protein may contain some sites/pockets that can bind small molecules, although Ensemble scores of DrugEBIlity may need to be further validated.
Figure 4. Searching information about target tractability for small molecule drug with TargetMine.
(A) Outline of the procedure for searching. (B) Protein structure regions and their Ensembl scores calculated by DrugEBIlity. (C) Compounds with bioactivity for PCSK9.
Collectively, we were able to confirm that the new version of TargetMine8 can quickly provide lines of evidences to assess linkage to disease and target tractability of PCSK9, and that the gathered data correctly reflected the real world situation; namely, it has been a challenge to obtain potent small molecule inhibitors for PCSK9, whereas antibody drugs for this protein have been successfully developed and marketed recently.
Gathering and prioritizing candidate drug target genes
To assess the utility of the new update of TargetMine8 for prioritizing candidate targets, we conducted a case study where we employed a list of genes associated with hypercholesterolemia in literature. We tentatively defined three key properties of a novel drug target suitable for small molecules as follows: (1) being associated with hypercholesterolemia via SNPs (GWAS catalog, ClinVar, or dbSNP-Pubmed; see Materials and Methods), (2) having greater than or equal to 50% of protein 3D structures with positive Ensemble scores (DrugEBIlity), and (3) having no reported (ChEMBL) potent small molecule inhibitors (IC50 or EC50 ≤ 100 nM).
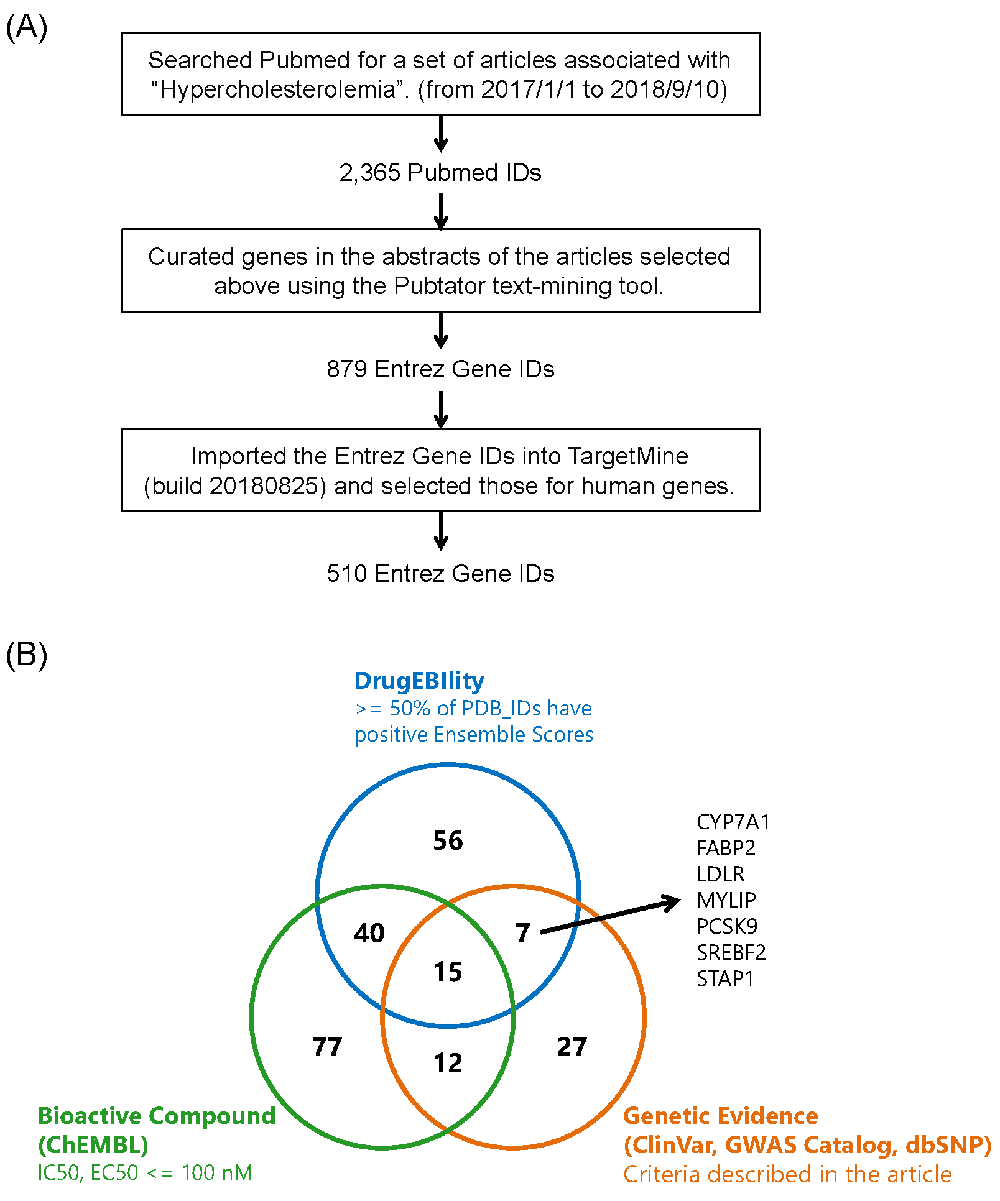
We first searched PubMed using the term “hypercholesterolemia” (from 2017/1/1 to 2018/9/10) and curated the resultant literature with the “Pubtator” text-mining tool28, resulting in 510 human genes (Figure 5A). We then selected the genes meeting the requirements defined above using the “Query Builder” in TargetMine8. Figure 5B shows an overview of the actual query, which aimed to extract the genes with gene evidences obtained from the GWAS catalog, where “Mapped Trait” contained “LDL cholesterol”, “total cholesterol”, or “low density lipoprotein cholesterol”, from ClinVar where “Reported phenotype Info” contains the term “Hypercholesterolemia”, and from dbSNP where “Mesh Terms Name” of related articles contains “Hypercholesterolemia”. Thus, the new implementation enabled us to filter objects on complex conditions with a user-friendly, intuitive graphical interface.
Figure 5. Gathering and prioritizing candidate drug target genes for hypercholesterolemia.
(A) Gathering hypercholesterolemia-related genes from article information in PubMed. (B) Prioritizing hypercholesterolemia-related genes with TargetMine to identify novel targets for small molecule drugs. Top prioritized genes were defined as those that met all of the following three requirements: 1) more than or equal to 50% of protein 3D structures (PDB IDs) having positive Ensembl scores, 2) no potent bioactive compounds (EC50 or IC50 ≤ 100 nM in ChEMBL) and 3) having genetic associations with hypercholesterolemia (for more details, see the Use Cases section).
Genes that satisfied all three requisites above are presented in Figure 5C (CYP7A1, FABP2, LDLR, MYLIP, PCSK9, SREBF2 and STAP1). Among the seven genes we found MYLIP and STAP1. MYLIP is an E3-ubiquitin ligase that degrades LDL receptors in the liver, which are therefore considered to be a potential therapeutic target for dyslipidemia29. Similarly, the STAP1 gene has been recently annotated as a fourth locus associated with autosomal-dominant hypercholesterolemia, and might be a novel target for therapeutic development of hypercholesterolemia30. This result suggests that the new version of TargetMine8 allows us to effectively prioritize target candidate genes in terms of linkage to disease, tractability and competitors. On the other hand, the list includes intractable targets such as PCSK9 and LDLR, indicating the need for improvement of the data and/or the thresholds with which tractable proteins are selected (in this study, ≥50% of protein 3D structures have positive Ensemble scores in DrugEBIlity database).
Conclusions
These use cases demonstrate that the updated version of TargetMine8 can be applied in pharmaceutical R&D, from the aspect of understanding the linkage to disease, examining the tractability of targets and prioritizing candidates. The recent update of the Open Targets platform31 also starts to cover “DrugEBIlity” data and protein structural information, suggesting that an integrated resource containing gene-disease associations and tractability information is indispensable for the pharmaceutical R&D. In addition, taking advantage of the features of the InterMine framework, TargetMine8 also facilitates more flexible and more complex queries for advanced users.
Data availability
The TargetMine data warehouse is publicly available at https://targetmine.mizuguchilab.org.
Software availability
Source code available from: https://github.com/chenyian-nibio/targetmine
Archived source code at time of publication: https://doi.org/10.5281/zenodo.25735658.
License: MIT License.
Grant information
This work was in part supported by JSPS KAKENHI (grant number 17K07268).
The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Acknowledgments
We would like to thank Andrew Myers’ help for improving the quality of writing.
Faculty Opinions recommended
References
- 1. Brown KK, Hann MM, Lakdawala AS, et al.: Approaches to target tractability assessment - a practical perspective. Medchemcomm. 2018; 9(4): 606–613. PubMed Abstract | Publisher Full Text | Free Full Text
- 2. Bunnage ME: Getting pharmaceutical R&D back on target. Nat Chem Biol. 2011; 7(6): 335–339. PubMed Abstract | Publisher Full Text
- 3. Cook D, Brown D, Alexander R, et al.: Lessons learned from the fate of AstraZeneca's drug pipeline: a five-dimensional framework. Nat Rev Drug Discov. 2014; 13(6): 419–431. PubMed Abstract | Publisher Full Text
- 4. Nelson MR, Tipney H, Painter JL, et al.: The support of human genetic evidence for approved drug indications. Nat Genet. 2015; 47(8): 856–860. PubMed Abstract | Publisher Full Text
- 5. Piñero J, Bravo À, Queralt-Rosinach N, et al.: DisGeNET: a comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Res. 2017; 45(D1): D833–D839. PubMed Abstract | Publisher Full Text | Free Full Text
- 6. Koscielny G, An P, Carvalho-Silva D, et al.: Open Targets: a platform for therapeutic target identification and validation. Nucleic Acids Res. 2017; 45(D1): D985–D994. PubMed Abstract | Publisher Full Text | Free Full Text
- 7. Nguyen DT, Mathias S, Bologa C, et al.: Pharos: Collating protein information to shed light on the druggable genome. Nucleic Acids Res. 2017; 45(D1): D995–D1002. PubMed Abstract | Publisher Full Text | Free Full Text
- 8. Sullivan J, Kalderimis A, Butano D, et al.: chenyian-nibio/targetmine: TargetMine v1.8.5.0 (Version v1.8.5.0). Zenodo. 2019. http://www.doi.org/10.5281/zenodo.2573565
- 9. Kalderimis A, Lyne R, Butano D, et al.: InterMine: extensive web services for modern biology. Nucleic Acids Res. 2014; 42((Web Server issue): W468–472. PubMed Abstract | Publisher Full Text | Free Full Text
- 10. Welter D, MacArthur J, Morales J, et al.: The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucleic Acids Res. 2014; 42(Database issue): D1001–1006. PubMed Abstract | Publisher Full Text | Free Full Text
- 11. Chen YA, Tripathi LP, Mizuguchi K: An integrative data analysis platform for gene set analysis and knowledge discovery in a data warehouse framework. Database (Oxford). 2016; 2016: pii: baw009. PubMed Abstract | Publisher Full Text | Free Full Text
- 12. Kibbe WA, Arze C, Felix V, et al.: Disease Ontology 2015 update: an expanded and updated database of human diseases for linking biomedical knowledge through disease data. Nucleic Acids Res. 2015; 43(Database issue): D1071–1078. PubMed Abstract | Publisher Full Text | Free Full Text
- 13. Schriml LM, Arze C, Nadendla S, et al.: Disease Ontology: a backbone for disease semantic integration. Nucleic Acids Res. 2012; 40(Database issue): D940–946. PubMed Abstract | Publisher Full Text | Free Full Text
- 14. Malone J, Holloway E, Adamusiak T, et al.: Modeling sample variables with an Experimental Factor Ontology. Bioinformatics. 2010; 26(8): 1112–1118. PubMed Abstract | Publisher Full Text | Free Full Text
- 15. MacArthur J, Bowler E, Cerezo M, et al.: The new NHGRI-EBI Catalog of published genome-wide association studies (GWAS Catalog). Nucleic Acids Res. 2017; 45(D1): D896–D901. PubMed Abstract | Publisher Full Text | Free Full Text
- 16. Landrum MJ, Lee JM, Benson M, et al.: ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 2016; 44(D1): D862–868. PubMed Abstract | Publisher Full Text | Free Full Text
- 17. Landrum MJ, Lee JM, Riley GR, et al.: ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 2014; 42(Database issue): D980–985. PubMed Abstract | Publisher Full Text | Free Full Text
- 18. Higasa K, Miyake N, Yoshimura J, et al.: Human genetic variation database, a reference database of genetic variations in the Japanese population. J Hum Genet. 2016; 61(6): 547–553. PubMed Abstract | Publisher Full Text | Free Full Text
- 19. Nagasaki M, Yasuda J, Katsuoka F, et al.: Rare variant discovery by deep whole-genome sequencing of 1,070 Japanese individuals. Nat Commun. 2015; 6: 8018. PubMed Abstract | Publisher Full Text | Free Full Text
- 20. NHLBI GO Exome Sequencing Project (ESP), S., WA. Vol. 2017.
- 21. 1000 Genomes Project Consortium, Auton A, Brooks LD, et al.: A global reference for human genetic variation. Nature. 2015; 526(7571): 68–74. PubMed Abstract | Publisher Full Text | Free Full Text
- 22. Sudmant PH, Rausch T, Gardner EJ, et al.: An integrated map of structural variation in 2,504 human genomes. Nature. 2015; 526(7571): 75–81. PubMed Abstract | Publisher Full Text | Free Full Text
- 23. Abifadel M, Varret M, Rabès JP, et al.: Mutations in PCSK9 cause autosomal dominant hypercholesterolemia. Nat Genet. 2003; 34(2): 154–156. PubMed Abstract | Publisher Full Text
- 24. Kiyosue A, Honarpour N, Kurtz C, et al.: A Phase 3 Study of Evolocumab (AMG 145) in Statin-Treated Japanese Patients at High Cardiovascular Risk. Am J Cardiol. 2016; 117(1): 40–47. PubMed Abstract | Publisher Full Text
- 25. Teramoto T, Kobayashi M, Tasaki H, et al.: Efficacy and Safety of Alirocumab in Japanese Patients With Heterozygous Familial Hypercholesterolemia or at High Cardiovascular Risk With Hypercholesterolemia Not Adequately Controlled With Statins - ODYSSEY JAPAN Randomized Controlled Trial. Circ J. 2016; 80(9): 1980–1987. PubMed Abstract | Publisher Full Text
- 26. Clarke L, Fairley S, Zheng-Bradley X, et al.: The international Genome sample resource (IGSR): A worldwide collection of genome variation incorporating the 1000 Genomes Project data. Nucleic Acids Res. 2017; 45(D1): D854–D859. PubMed Abstract | Publisher Full Text | Free Full Text
- 27. Liu ZP, Wang Y: PCSK9 Inhibitors: Novel Therapeutic Strategies for Lowering LDLCholesterol. Mini Rev Med Chem. 2019; 19(2): 165–176. PubMed Abstract | Publisher Full Text
- 28. Wei CH, Kao HY, Lu Z: PubTator: a web-based text mining tool for assisting biocuration. Nucleic Acids Res. 2013; 41(Web Server issue): W518–522. PubMed Abstract | Publisher Full Text | Free Full Text
- 29. Zhang CP, Tian Y, Zhang M, et al.: IDOL, inducible degrader of low-density lipoprotein receptor, serves as a potential therapeutic target for dyslipidemia. Med Hypotheses. 2016; 86: 138–142. PubMed Abstract | Publisher Full Text
- 30. Day IN: FH4=STAP1. Another gene for familial hypercholesterolemia? Relevance to cascade testing and drug development? Circ Res. 2014; 115(6): 534–536. PubMed Abstract | Publisher Full Text
- 31. Carvalho-Silva D, Pierleoni A, Pignatelli M, et al.: Open Targets Platform: new developments and updates two years on. Nucleic Acids Res. 2019; 47(D1): D1056–D1065. PubMed Abstract | Publisher Full Text | Free Full Text
Comments on this article Comments (0)
Version 2
VERSION 2 PUBLISHED 28 Feb 2019
This work was in part supported by JSPS KAKENHI (grant number 17K07268).
The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
© 2019 Chen YA et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Open Peer Review
Current Reviewer Status: ?
Key to Reviewer Statuses VIEW HIDE
ApprovedThe paper is scientifically sound in its current form and only minor, if any, improvements are suggested
Approved with reservations A number of small changes, sometimes more significant revisions are required to address specific details and improve the papers academic merit.
Not approvedFundamental flaws in the paper seriously undermine the findings and conclusions
Version 1
VERSION 1
PUBLISHED 28 Feb 2019
Reviewer Report 28 Mar 2019
Anne Hersey, European Bioinformatics Institute (EMBL-EBI), Cambridge, UK
Approved
VIEWS 0
- Is the rationale for developing the new software tool clearly explained?
Yes - Is the description of the software tool technically sound?
Yes - Are sufficient details of the code, methods and analysis (if applicable) provided to allow replication of the software development and its use by others?
Yes - Is sufficient information provided to allow interpretation of the expected output datasets and any results generated using the tool?
Yes - Are the conclusions about the tool and its performance adequately supported by the findings presented in the article?
Yes
Competing Interests: No competing interests were disclosed.
Reviewer Expertise: Chemoinformatics, drug discovery, bioactivity databases
Close
Reviewer Report 27 Mar 2019
Rachel Lyne, Department of Genetics, University of Cambridge, Cambridge, UK
Approved
VIEWS 0
- Is the rationale for developing the new software tool clearly explained?
Yes - Is the description of the software tool technically sound?
Yes - Are sufficient details of the code, methods and analysis (if applicable) provided to allow replication of the software development and its use by others?
Partly - Is sufficient information provided to allow interpretation of the expected output datasets and any results generated using the tool?
Partly - Are the conclusions about the tool and its performance adequately supported by the findings presented in the article?
Yes
Competing Interests: No competing interests were disclosed.
Reviewer Expertise: Bioinformatics, Databases, Data integration, data analysis
Close
Comments on this article Comments (0)
Version 2
VERSION 2 PUBLISHED 28 Feb 2019
Open Peer Review
Reviewer Status
Alongside their report, reviewers assign a status to the article:
Approved
The paper is scientifically sound in its current form and only minor, if any, improvements are suggested
Approved with reservations
A number of small changes, sometimes more significant revisions are required to address specific details and improve the papers academic merit.
Not approved
Fundamental flaws in the paper seriously undermine the findings and conclusions
Reviewer Reports
| Invited Reviewers | ||
|---|---|---|
| 1 | 2 | |
| Version 2 (revision) 28 May 19 | ||
| Version 1 28 Feb 19 | read | read |
- Rachel Lyne, University of Cambridge, Cambridge, UK
- Anne Hersey, European Bioinformatics Institute (EMBL-EBI), Cambridge, UK
Comments on this article
Sign up for content alerts
Browse by related subjects
Alongside their report, reviewers assign a status to the article:
Approved - the paper is scientifically sound in its current form and only minor, if any, improvements are suggested
Approved with reservations - A number of small changes, sometimes more significant revisions are required to address specific details and improve the papers academic merit.
Not approved - fundamental flaws in the paper seriously undermine the findings and conclusions
Adjust parameters to alter display
View on desktop for interactive features ![]()
Includes Interactive Elements
View on desktop for interactive features ![]()