Stochastic gene expression out-of-steady-state in the cyanobacterial circadian clock (original) (raw)
- Letter
- Published: 20 December 2007
Nature volume 450, pages 1249–1252 (2007) Cite this article
- 2910 Accesses
- 107 Citations
- 3 Altmetric
- Metrics details
Abstract
Recent advances in measuring gene expression at the single-cell level have highlighted the stochastic nature of messenger RNA and protein synthesis1,2,3. Stochastic gene expression creates a source of variability in the abundance of cellular components, even among isogenic cells exposed to an identical environment. Recent integrated experimental and modelling studies4,5,6,7,8,9,10,11,12,13 have shed light on the molecular sources of this variability. However, many of these studies focus on systems that have reached a steady state and therefore do not address a large class of dynamic phenomena including oscillatory gene expression. Here we develop a general protocol for analysing and predicting stochastic gene expression in systems that never reach steady states. We use this framework to analyse experimentally stochastic expression of genes driven by the Synechococcus elongatus circadian clock. We find that, although the average expression at two points in the circadian cycle separated by 12 hours is identical, the variability at these two time points can be different. We show that this is a general feature of out-of-steady-state systems. We demonstrate how intrinsic noise sources, owing to random births and deaths of mRNAs and proteins, or extrinsic noise sources, which introduce fluctuations in rate constants, affect the cell-to-cell variability. To distinguish experimentally between these sources, we measured how the correlation between expression fluctuations of two identical genes is modulated during the circadian cycle. This quantitative framework is generally applicable to any out-of-steady-state system and will be necessary for understanding the fidelity of dynamic cellular systems.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Additional access options:
Figure 1: Monitoring circadian oscillations in single S. elongatus PCC7942 cells using fluorescence microscopy.

The alternative text for this image may have been generated using AI.
Figure 2: Circadian oscillations in single cells.

The alternative text for this image may have been generated using AI.
Figure 3: A dynamic analysis of stochastic gene expression reveals noise loops.
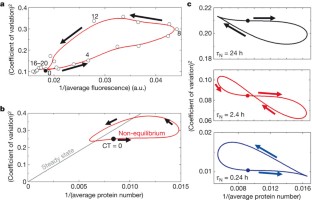
The alternative text for this image may have been generated using AI.
Figure 4: Cell to cell variability in single- and double-copy-number constructs.

The alternative text for this image may have been generated using AI.
Similar content being viewed by others
References
- Kaern, M., Elston, T. C., Blake, W. J. & Collins, J. J. Stochasticity in gene expression: from theories to phenotypes. Nature Rev. Genet. 6, 451–464 (2005)
Article CAS PubMed Google Scholar - Kaufmann, B. B. & van Oudenaarden, A. Stochastic gene expression: from single molecules to the proteome. Curr. Opin. Genet. Dev. 17, 107–112 (2007)
Article CAS PubMed Google Scholar - Maheshri, N. & O’Shea, E. K. Living with noisy genes: how cells function reliably with inherent variability in gene expression. Annu. Rev. Biophys. Biomol. Struct. 36, 413–434 (2007)
Article CAS PubMed Google Scholar - Ozbudak, E. M., Thattai, M., Kurtser, I., Grossman, A. D. & van Oudenaarden, A. Regulation of noise in the expression of a single gene. Nature Genet. 31, 69–73 (2002)
Article CAS PubMed Google Scholar - Elowitz, M. B., Levine, A. J., Siggia, E. D. & Swain, P. S. Stochastic gene expression in a single cell. Science 297, 1183–1186 (2002)
Article ADS CAS PubMed Google Scholar - Blake, W. J., Kaern, M., Cantor, C. R. & Collins, J. J. Noise in eukaryotic gene expression. Nature 422, 633–637 (2003)
Article ADS CAS PubMed Google Scholar - Raser, J. M. & O’Shea, E. K. Control of stochasticity in eukaryotic gene expression. Science 304, 1811–1814 (2004)
Article ADS CAS PubMed PubMed Central Google Scholar - Paulsson, J. Summing up the noise in gene networks. Nature 427, 415–418 (2004)
Article ADS CAS PubMed Google Scholar - Cai, L., Friedman, N. & Xie, X. S. Stochastic protein expression in individual cells at the single molecule level. Nature 440, 358–362 (2006)
Article ADS CAS PubMed Google Scholar - Newman, J. R. et al. Single-cell proteomic analysis of S. cerevisiae reveals the architecture of biological noise. Nature 441, 840–846 (2006)
Article ADS CAS PubMed Google Scholar - Bar-Even, A. et al. Noise in protein expression scales with natural protein abundance. Nature Genet. 38, 636–643 (2006)
Article CAS PubMed Google Scholar - Sigal, A. et al. Variability and memory of protein levels in human cells. Nature 444, 643–646 (2006)
Article ADS CAS PubMed Google Scholar - Geva-Zatorsky, N. et al. Oscillations and variability in the p53 system. Mol. Syst. Biol. 2, 0033 (2006)
Article PubMed Google Scholar - Kondo, T. et al. Circadian rhythms in prokaryotes: luciferase as a reporter of circadian gene expression in cyanobacteria. Proc. Natl Acad. Sci. USA 90, 5672–5676 (1993)
Article ADS CAS PubMed PubMed Central Google Scholar - Liu, Y., Golden, S. S., Kondo, T., Ishiura, M. & Johnson, C. H. Bacterial luciferase as a reporter of circadian gene expression in cyanobacteria. J. Bacteriol. 177, 2080–2086 (1995)
Article CAS PubMed PubMed Central Google Scholar - Mihalcescu, I., Hsing, W. & Leibler, S. Resilient circadian oscillator revealed in individual cyanobacteria. Nature 430, 81–85 (2004)
Article ADS CAS PubMed Google Scholar - Andersen, J. B. et al. New unstable variants of green fluorescent protein for studies of transient gene expression in bacteria. Appl. Environ. Microbiol. 64, 2240–2246 (1998)
CAS PubMed PubMed Central Google Scholar - Andersson, C. R. et al. Application of bioluminescence to the study of circadian rhythms in cyanobacteria. Methods Enzymol. 305, 527–542 (2000)
Article CAS PubMed Google Scholar - Xu, Y., Mori, T. & Johnson, C. H. Cyanobacterial circadian clockwork: roles of KaiA, KaiB and the kaiBC promoter in regulating KaiC. EMBO J. 22, 2117–2126 (2003)
Article CAS PubMed PubMed Central Google Scholar - Thattai, M. & van Oudenaarden, A. Intrinsic noise in gene regulatory networks. Proc. Natl Acad. Sci. USA 98, 8614–8619 (2001)
Article ADS CAS PubMed PubMed Central Google Scholar - Swain, P. S., Elowitz, M. B. & Siggia, E. D. Intrinsic and extrinsic contributions to stochasticity in gene expression. Proc. Natl Acad. Sci. USA 99, 12795–12800 (2002)
Article ADS CAS PubMed PubMed Central Google Scholar - Kepler, T. B. & Elston, T. C. Stochasticity in transcriptional regulation: origins, consequences, and mathematical representations. Biophys. J. 81, 3116–3136 (2001)
Article ADS CAS PubMed PubMed Central Google Scholar - Becskei, A., Kaufmann, B. B. & van Oudenaarden, A. Contributions of low molecule number and chromosomal positioning to stochastic gene expression. Nature Genet. 37, 937–944 (2005)
Article CAS PubMed Google Scholar - Volfson, D. et al. Origins of extrinsic variability in eukaryotic gene expression. Nature 439, 861–864 (2006)
Article ADS CAS PubMed Google Scholar - Rosenfeld, N., Young, J. W., Alon, U., Swain, P. S. & Elowitz, M. B. Gene regulation at the single-cell level. Science 307, 1962–1965 (2005)
Article ADS CAS PubMed Google Scholar - Pedraza, J. M. & van Oudenaarden, A. Noise propagation in gene networks. Science 307, 1965–1969 (2005)
Article ADS CAS PubMed Google Scholar - Johnson, C. H. Global orchestration of gene expression by the biological clock of cyanobacteria. Genome Biol. 5, 217 (2004)
Article PubMed PubMed Central Google Scholar - Smith, R. M. & Williams, S. B. Circadian rhythms in gene transcription imparted by chromosome compaction in the cyanobacterium Synechococcus elongatus. Proc. Natl Acad. Sci. USA 103, 8564–8569 (2006)
Article ADS CAS PubMed PubMed Central Google Scholar - Vilar, J. M., Kueh, H. Y., Barkai, N. & Leibler, S. Mechanisms of noise-resistance in genetic oscillators. Proc. Natl Acad. Sci. USA 99, 5988–5992 (2002)
Article ADS CAS PubMed PubMed Central Google Scholar - Gonze, D., Halloy, J. & Goldbeter, A. Robustness of circadian rhythms with respect to molecular noise. Proc. Natl Acad. Sci. USA 99, 673–678 (2002)
Article ADS CAS PubMed PubMed Central Google Scholar
Acknowledgements
We thank S. S. Golden and J. L. Ditty for assistance with the initial phase of this work and their gifts of plasmids and strains. We acknowledge I. Lipchin and M. J. T. O’Kelly for assistance with data collection, cloning and bioluminescence measurements. We acknowledge A. Tolonen, S. W. Chisholm, M. Thattai, H. Lim, J. C. Gore and A. Raj for discussions and suggestions. This work was performed in part at the MIT Laser Biomedical Research Center. This work was supported by NSF and NIH grants.
Author Contributions J.R.C. and P.L. performed the experiments. J.M.P. developed the model. J.R.C., .J.M.P. and A.v.O. designed the experiments, interpreted the results and wrote the paper.
Author information
Author notes
- Jeffrey R. Chabot
Present address: Present address: Systems Biology Group, Pfizer Research Technology Center, Cambridge, Massachusetts 02139, USA.,
Authors and Affiliations
- Department of Physics and George Harrison Spectroscopy Laboratory, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA,
Jeffrey R. Chabot, Juan M. Pedraza, Prashant Luitel & Alexander van Oudenaarden
Authors
- Jeffrey R. Chabot
- Juan M. Pedraza
- Prashant Luitel
- Alexander van Oudenaarden
Corresponding author
Correspondence toAlexander van Oudenaarden.
Supplementary information
Supplementary Information (download PDF )
This file contains Supplementary experimental and modeling Methods and Materials with Supplementary Figures S1-S2. (PDF 585 kb)
Supplementary Movie 1 (download AVI )
This file contains Supplementary Movie 1 demonstrating circadian oscillations in single S. elongatus PCC7942 cells. This file was modified on 31 January 2008 because of the technical difficulties with the file format. (AVI 3601 kb)
Rights and permissions
About this article
Cite this article
Chabot, J., Pedraza, J., Luitel, P. et al. Stochastic gene expression out-of-steady-state in the cyanobacterial circadian clock.Nature 450, 1249–1252 (2007). https://doi.org/10.1038/nature06395
- Received: 13 August 2007
- Accepted: 16 October 2007
- Issue date: 20 December 2007
- DOI: https://doi.org/10.1038/nature06395